Biomod/2013/UT-Austin/Introduction
<html> <style type="text/css"> // </style>
<a href="http://openwetware.org/wiki/Biomod/2013/UT-Austin"><img src="http://openwetware.org/images/2/2f/G4262.png" width ="50%" height="50%"/>
<a href="http://openwetware.org/wiki/Biomod/2013/UT-Austin/Introduction"><img src="http://openwetware.org/images/0/0d/Intro.png" width="20%" alt="" /></a>
<a href="http://openwetware.org/wiki/Biomod/2013/UT-Austin/Design"><img src="http://openwetware.org/images/0/0a/Design.png" width="10%" alt="" /></a>
<a href="http://openwetware.org/wiki/Biomod/2013/UT-Austin/Methods"><img src="http://openwetware.org/images/0/07/MEthods.png" width="15%" alt="" /></a>
<a href="http://openwetware.org/wiki/Biomod/2013/UT-Austin/Results"><img src="http://openwetware.org/images/1/17/Results.png" width="12%" alt="" /></a>
<a href="http://openwetware.org/wiki/Biomod/2013/UT-Austin/Team Info"><img src="http://openwetware.org/images/3/32/Team.png" width="15%" alt="" /></a>
<a href="http://openwetware.org/wiki/Biomod/2013/UT-Austin/References"><img src="http://openwetware.org/images/0/0d/References.png" width="20%" alt="" /></a>
</html>
Introduction
Modern medicine is defined by its relationship with large centralized medical laboratories. Advances in biomedical science has led to the development of many powerful, yet expertise and resource intensive, diagnostic techniques. Due to these expertise and resource requirements, the deployment of these modern techniques was realized through the development of large centralized medical laboratories. These labs have greatly expanded a physician’s ability to diagnose, and in turn, treat patients. However, these labs are not without their flaws. Testing through a central laboratory increases the time between attainment of the patient’s fluid or tissue sample to delivery of test results to the physician. Additionally, the current state of dependence upon these labs has hindered the deployment of modern care to resource poor areas.
Point-of-Care Diagnostics and Biosensors
Point-of-care (POC) diagnostics are diagnostic technologies that move the analysis of patient samples from the lab bench to the bedside and in some cases the home. More specifically, POC diagnostics are small and simple to operate devices that can process complex clinical samples for various biomarkers. This both decreases the time to diagnosis and removes some of the barriers to deploying modern care into the field.
Frequently, the miniaturization and simplification of sample analysis is achieved through the use of biosensors. An illustrative example of this is the home pregnancy test. Modern pregnancy tests use antibodies to detect elevated levels of hormones found during pregnancy. Specifically, most pregnancy test use anti-human chorionic gonadotropin (hCG) conjugated with colloidal metals to visualize elevated hCG in the urine of pregnant women.
Proteins vs. Nucleic Acids As Biosensors
While modern pregnancy tests are relatively rugged, i.e. no refrigeration or exhaustive storage criteria exist, the development of protein-based biosensors is frequently hindered by the inherent instability of protein components. Various techniques for stabilization of useful proteins, e.g. antibodies and enzymes, are in development. However, the tendency of proteins to irreversibly denature remains highly problematic.
In this context, functional nucleic acids offer the potential to replace proteins in the development of rugged biosensors. Nucleic acids (NAs) can generally be reversibly denatured and, when desiccated and isolated from nucleases, degrade on a millennial timescale. NAs are amenable to a variety of conjugation chemistries that allow for attachment of a range of functional groups and dyes. Additionally NAs, like proteins, can specifically recognize biomarkers through the formation of complex tertiary structure (e.g. aptamers) and, in the case of genomic material, direct hybridization. The predictable nature of these hybridization interactions has enabled the development of complex DNA circuits for both biomarker signal amplification and transduction.
DNA Circuitry
Toehold-Mediated Strand Displacement
Toehold-mediated strand displacement is a specific hybridization interaction that is frequently exploited for the construction of complex DNA circuits. The process, illustrated in Figure 2, begins with an ssDNA oligo, green, binding a short (frequently 4-10 base pairs in length) single stranded domain of a DNA duplex, red and blue. This single stranded domain is deemed a toehold, as it allows the green oligo to weakly associate with the duplex through hybridization to this small single stranded domain. The green oligo then undergoes a process referred to as branch migration. Branch migration is the random walk process in which one domain displaces another of identical sequence through a series of reversible single nucleotide dissociation and hybridization steps. While each hybridization event in branch migration is reversible, the greater total complementarity of the green strand to the blue strand makes displacement of the red strand thermodynamically favorable.
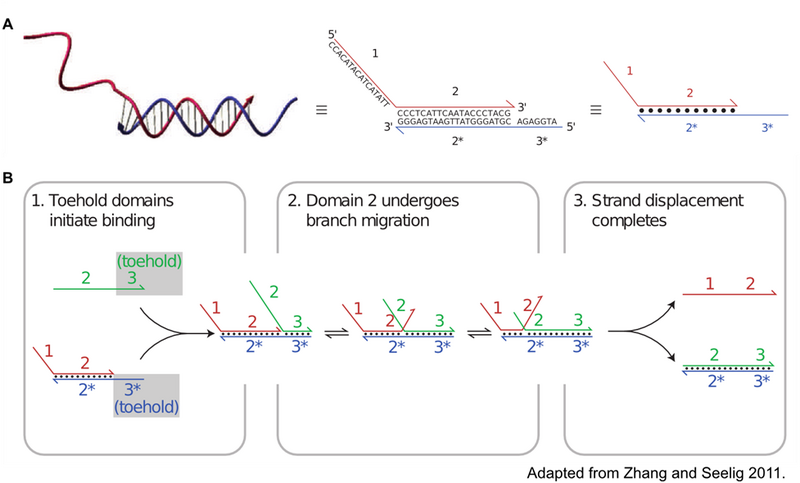
Catalyzed Hairpin Assembly
Catalyzed hairpin assembly (CHA) is an enzyme-free nucleic acid signal amplification circuit that enables specific detection of oligonucleotides. The signal amplification reaction involves three oligos, depicted in Figure 3: the catalyst (C1), the first hairpin (H1), and the second hairpin (H2). The reaction begins with C1 opening the H1 hairpin by toehold-mediated strand displacement through the 1 toehold of H1. The now linear H1 now has an exposed 3* domain. This acts as a toehold for H2, in turn, allowing H2 to fully hybridize to H1. C1, which is only weekly associated with the H1:H2 duplex, will eventually dissociate, thereby completing one cycle of the reaction and freeing C1 to generate additional H1:H2 duplexes. In this fashion, the presence, or signal, of a few catalyst molecules is amplified through the generation of numerous H1:H2.
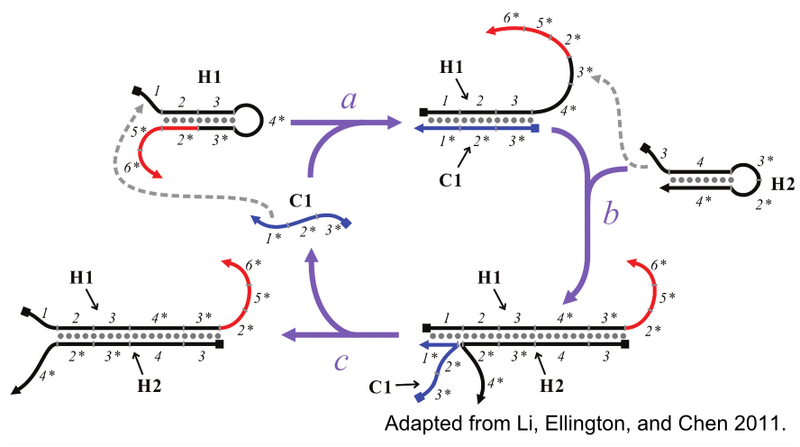
In addition to amplifying signal, the formation of H1:H2 fully exposes the 2*, 5* and 6* domains, in red, of H1. These domains can be modified and used in concert with reporter oligonucleotides to produce various forms of signals for the presence of H1:H2. Figure 4 depicts CHA reporter schemes that include fluorescent, colorimetric, and electrochemical readout. All reporter schemes make use of the 2* domain that is exposed on formation of H1:H2.
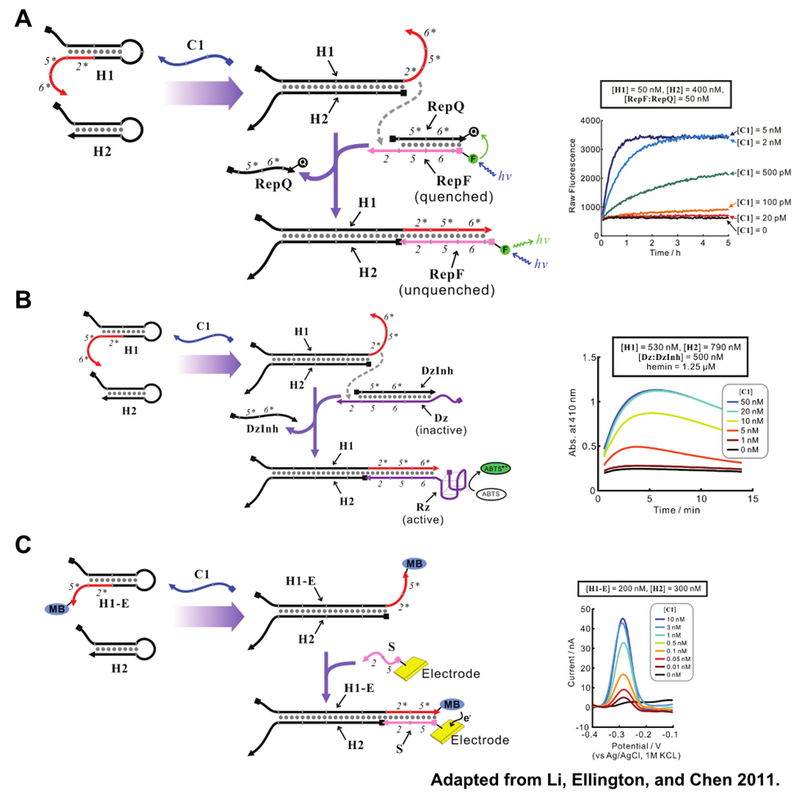
The potential impact of CHA as an analytical tool inspired our team to investigate if CHA could not only detect oligonucleotides but also characterize them. With slight design modifications, discussed in the Design section, CHA can detect small point defects in oligos. The ability to characterize oligos with an enzyme free circuit would potentially allow for the development of rugged POC diagnostic devices for detection of genetic abnormalities, virulent strains of bacteria, and the activity of toxins that act upon nucleic acids. Therefore, our team decided to develop modified CHA circuits for the detection of cancer-linked MDM2 single nucleotide polymorphisms and ricin generated apurinic sites.
Single Nucleotide Polymorphisms
A single nucleotide polymorphism (SNP) is a DNA sequence variation at a single position (i.e. an A, T, C, or G) within a population. SNPs account for a large portion of human genetic diversity. For example, twenty-five percent of the population may possess an A-T pair at a certain locus, while the remaining seventy-five percent possess a G-C pair. In the case of human SNPs, certain SNP alleles have been associated with adverse responses to certain medications, susceptibility to infectious diseases, and propensity to develop cancer. In the case of bacterial SNPs, certain alleles have been associated with heightened virulence or resistance to antibiotics.
MDM2 SNP309
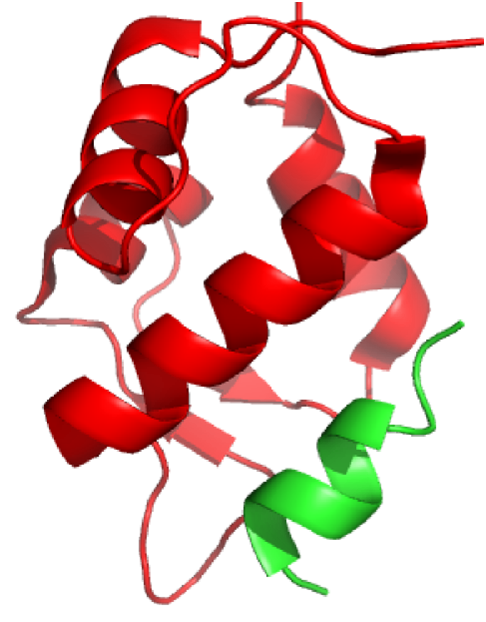
Our specific target for this project is SNP309 of the mouse double minute 2 homolog (MDM2) gene. This is a well characterized SNP that is associated with increase cancer risk. SNP309, located in the promoter of the gene, is a T/G SNP with the G allele being the cancer-linked allele. The G allele increases cancer risk by upregulating expression of MDM2. This upregulation leads to increased cancer risk due to MDM2’s ability to bind and inhibit the tumor suppressor protein, p53, depicted in Figure 5. Therefore, easy detection of SNP309 would be useful for evaluating an individual’s cancer risk.
Current SNP Detection Methods
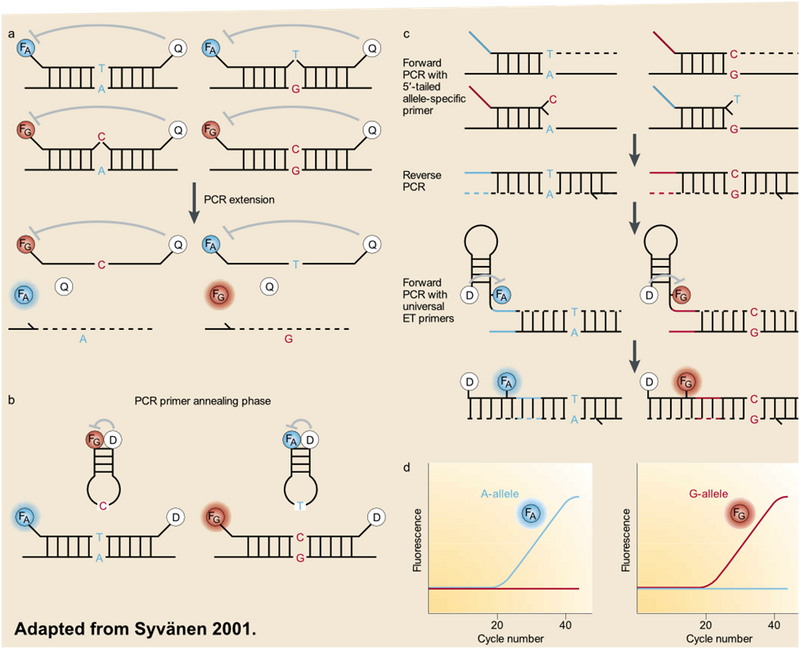
The most common method of SNP detection is real-time PCR (rtPCR). rtPCR is essentially PCR with fluorescent probes (depicted in Figure 6) to quantify the products of the reaction. Probe designs vary but all operate by exploiting the single base pair mismatch that can result from SNP allele variation.
Apurinic Sites
Similar to SNPs, apurinic (AP) sites also affect a single nucleotide. AP sites (depicted in Figure 7) occur when a purine ring is removed from the ribose group of the nucleic acid backbone. For design purposes, apurinic sites should behave in a similar fashion to single base pair mismatches associated with SNPs.
Ricin Generated AP Sites
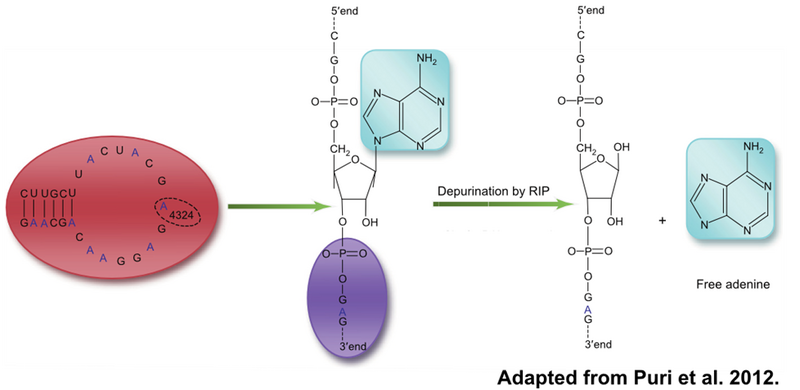
Ricin, found in the seeds of the castor oil plant, is a highly lethal toxin that acts by inactivating the ribosome. Specifically, ricin depurinates the 4324 adenine of the 28S ribosomal RNA. This depurination weakens the binding of elongation factors to the A4324 loop. Without functioning elongation factors, the ribosome cannot translocate tRNAs following peptidyl transfer in protein synthesis. The inability to produce new proteins leads to rapid cell death.
Current Ricin Detection Methods
Current ricin detection methods are relatively limited. In vivo (i.e. lethal doses delivered to mice) assays and in situ (i.e. cytotoxicity) assays exist. However, recent efforts have been focused on attaining in vitro assays for ricin activity. These assays would allow for easier and quicker detection of ricin.