User talk:Farnad Zaghi
From OpenWetWare
Jump to navigationJump to search
Proteomic E-coli Project
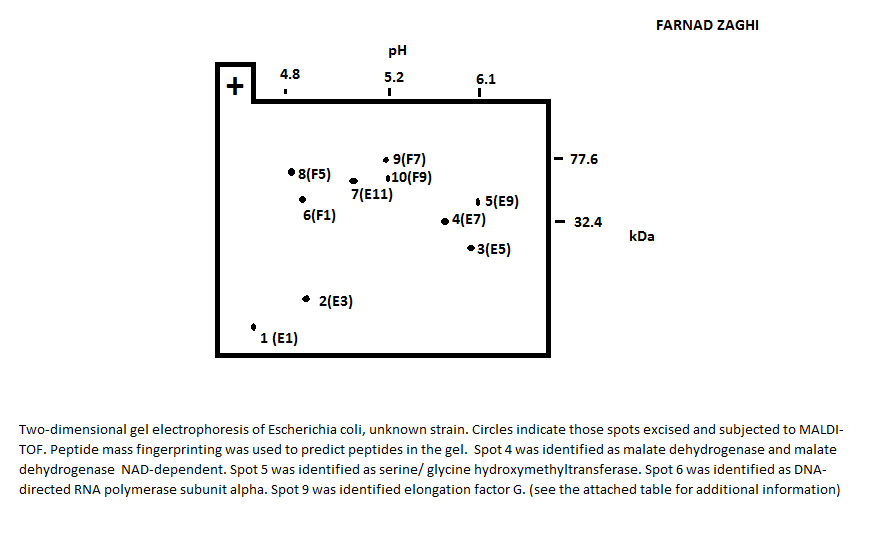
Two-dimensional gel electrophoresis of E. Coli, followed by MALDI-TOF spectra were acquired for protein identification.
Mascot Wizard Results
result
| Spot Number | NCBI accession (gI) number | Uniprot accession # | Theor-etical Mass | Theor-etical pI | RMS error (ppm) | # peptides matched | % Coverage | Mascot Protein Score | Mascot Score Highest Non-Homologous | Protein Name | Summary URL | Protein View URL | Revised Protein Summary | Additional Protein View | Additional Protein View | Comment | input_data_file | ' |
| 1 (E1) | no hit | http://128.59.63.100/mascot/cgi/master_results.pl?file=../data/20111207/F015434.dat&sessionID=FZaghi_81508037746369 | Invirtogel_E1_0001 | |||||||||||||||
| 2 (E3) | no hit | http://128.59.63.100/mascot/cgi/master_results.pl?file=../data/20111207/F015433.dat&sessionID=FZaghi_81508037746369 | Invirtogel_E3_0001 | |||||||||||||||
| 3 (E5) | no hit | http://128.59.63.100/mascot/cgi/master_results.pl?file=../data/20111207/F015432.dat&sessionID=FZaghi_81508037746369 | Invirtogel_E5_0001 | |||||||||||||||
| 4 (E7) | 297521112|ref | NP_06939498 | 19514 | 5.17 | 30 | 10 | 69% | 120 | 87 | malate dehydrogenase [Escherichia coli OP50] | http://128.59.63.100/mascot/cgi/master_results.pl?file=../data/20111207/F015435.dat&sessionID=FZaghi_81508037746369 | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111207/F015435.dat&hit=1 | malate dehydrogenase and malate dehydrogenase NAD-dependent for deifferent strant of E coli | Invirtogel_E7_0001 | ||||
| 15803770|ref | NP_289804 | 32488 | 5.61 | 31 | 11 | 44% | 112 | 87 | malate dehydrogenase [Escherichia coli O157:H7 EDL933] | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111207/F015435.dat&hit=2 | ||||||||
| 91212656|ref | YP_542642 | 35207 | 6.03 | 31 | 11 | 41% | 109 | 87 | malate dehydrogenase [Escherichia coli UTI89] | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111207/F015435.dat&hit=9 | ||||||||
| 170018520|ref | YP_001723474 | 32478 | 5.61 | 31 | 11 | 44% | 112 | 87 | malate dehydrogenase [Escherichia coli ATCC 8739] | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111207/F015435.dat&hit=6 | ||||||||
| 209920706|ref | YP_002294790 | 32504 | 5.61 | 31 | 11 | 44% | 112 | 87 | malate dehydrogenase [Escherichia coli SE11 | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111207/F015435.dat&hit=8 | ||||||||
| 218555800|ref | YP_002388713 | 32461 | 5.61 | 30 | 10 | 37% | 95 | 87 | malate dehydrogenase [Escherichia coli IAI1] | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111207/F015435.dat&hit=19 | ||||||||
| 218706850|ref | YP_002414369 | 32487 | 6.21 | 30 | 10 | 37% | 95 | 87 | "malate dehydrogenase [Escherichia coli UMN026] | |||||||||
| " | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111207/F015435.dat&hit=20 | |||||||||||||||||
| 110643470|ref | YP_671200 | 32502 | 5.61 | 30 | 10 | 37% | 95 | 87 | malate dehydrogenase [Escherichia coli 536] | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111207/F015435.dat&hit=21 | ||||||||
| 194429110|ref | ZP_03061640 | 32488 | 5.61 | 31 | 11 | 44% | 112 | 87 | malate dehydrogenase, NAD-dependent [Escherichia coli B171] | http://128.59.63.100/mascot/cgi/master_results.pl?file=../data/20111207/F015435.dat&sessionID=FZaghi_81508037746369 | ||||||||
| 191168165|ref | ZP_03029961 | 32477 | 5.61 | 30 | 10 | 37% | 95 | 87 | malate dehydrogenase, NAD-dependent [Escherichia coli B7A] | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111207/F015435.dat&hit=17 | ||||||||
| 300898002|ref | ZP_07116376 | 35206 | 6.85 | 30 | 10 | 35% | 92 | 87 | malate dehydrogenase, NAD-dependent [Escherichia coli MS 198-1] | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111207/F015435.dat&hit=22 | ||||||||
| 301022169|ref | ZP_07186088 | 35207 | 6.36 | 30 | 10 | 35% | 92 | 87 | malate dehydrogenase, NAD-dependent [Escherichia coli MS 69-1] | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111207/F015435.dat&hit=23 | ||||||||
| 300937348|ref | ZP_07152185 | 35223 | 6.36 | 30 | 10 | 35% | 92 | 87 | malate dehydrogenase, NAD-dependent [Escherichia coli MS 21-1] | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111207/F015435.dat&hit=24 | ||||||||
| 300817513|ref | ZP_07097729 | 35196 | 6.36 | 30 | 10 | 35% | 92 | 87 | malate dehydrogenase, NAD-dependent [Escherichia coli MS 107-1] | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111207/F015435.dat&hit=25 | ||||||||
| 300979785|ref | ZP_07174711 | 35221 | 6.36 | 30 | 10 | 35% | 92 | 87 | malate dehydrogenase, NAD-dependent [Escherichia coli MS 200-1] | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111207/F015435.dat&hit=26 | ||||||||
| 5 (E9) | 16130476|ref | NP_417046 | 45459 | 6.03 | 24 | 16 | 27% | 169 | NONE | serine hydroxymethyltransferase [Escherichia coli str. K-12 substr. MG1655] | http://128.59.63.100/mascot/cgi/master_results.pl?file=../data/20111207/F015415.dat&sessionID=FZaghi_450724603025029 | http://128.59.63.100/mascot/cgi/protein_view.pl?file=..%2Fdata%2F20111207%2FF015415.dat;hit=2 | serine/ glycine hydroxymethyltransferase for different strains of E coli | Invirtogel_E9_0001 | ||||
| 15803076|ref | NP_289107 | 45487 | 6.03 | 24 | 16 | 27% | 169 | NONE | serine hydroxymethyltransferase [Escherichia coli O157:H7 EDL933] | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111207/F015415.dat&hit=3 | ||||||||
| 218706054|ref | YP_002413573 | 45445 | 6.03 | 24 | 16 | 27% | 169 | NONE | serine hydroxymethyltransferase [Escherichia coli O157:H7 EDL933] | http://128.59.63.100/mascot/cgi/protein_view.pl?file=..%2Fdata%2F20111208%2FF015472.dat;hit=7 | ||||||||
| 26248915|ref | NP_754955 | 45746 | 6.13 | 24 | 16 | 27% | 167 | NONE | serine hydroxymethyltransferase [Escherichia coli CFT073 | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111208/F015472.dat&hit=8 | ||||||||
| 291283776|ref | YP_003500594 | 45501 | 6.13 | 24 | 16 | 27% | 167 | NONE | Serine hydroxymethyltransferase [Escherichia coli O55:H7 str. CB9615] | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111208/F015472.dat&hit=10 | ||||||||
| 297521279|ref | ZP_06939665 | 16829 | 7.74 | 44 | 11 | 51% | 128 | serine hydroxymethyltransferase [Escherichia coli OP50] | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111208/F015472.dat&hit=16 | |||||||||
| 300898338|ref | ZP_07116686 | 45732 | 6.13 | 24 | 16 | 27% | 167 | NONE | glycine hydroxymethyltransferase [Escherichia coli MS 198-1]. | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111208/F015472.dat&hit=9 | ||||||||
| 301024834|ref | ZP_07188471 | 45776 | 6.13 | 24 | 15 | 26% | 155 | NONE | glycine hydroxymethyltransferase [Escherichia coli MS 69-1] | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111208/F015472.dat&hit=11 | ||||||||
| 7 (E11) | 16131841|ref | NP_418439 | 47777 | 5.16 | 23 | 29 | 61% | 293 | 210 | isocitrate lyase [Escherichia coli str. K-12 substr. MG1655] | http://128.59.63.100/mascot/cgi/master_results.pl?file=../data/20111207/F015436.dat&sessionID=FZaghi_81508037746369 | http://128.59.63.100/mascot/cgi/protein_view.pl?file=..%2Fdata%2F20111207%2FF015436.dat;hit=1 | 227886989|ref for E coli 83972, gi|218561077|ref for E coli S88, gi|91213527|ref for E coli UTI89, gi|15804601|ref for E coli O157:H7 EDL933 , gi|291285425|ref for E coli O55:H7 str. CB9615, gi|26250784|ref for E coli CFT073, and gi|300820055|ref for E coli MS 107-1. | |||||
| 193062786|ref | ZP_03043879 | 51533 | 5.06 | 18 | 13 | 33% | 85 | 293 | 6-phosphogluconate dehydrogenase, decarboxylating [Escherichia coli E22] | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111207/F015436.dat&hit=63 | It has been reported as a Mixture with aboave Protien ID. | |||||||
| 16129099|ref | NP_415654 | 46070 | 5.15 | 18 | 22 | 60% | 210 | 293 | "e14 prophage; isocitrate dehydrogenase, specific for NADP+ [Escherichia coli str. K-12 substr. MG1655] | |||||||||
| [Escherichia coli str. K-12 substr. MG1655]. | ||||||||||||||||||
| " | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111207/F015436.dat&hit=24 | Invirtogel_E11_0001 | ||||||||||||||||
| 300922666|ref | ZP_07138763 | 46011 | 5.35 | 17 | 20 | 55% | 179 | 293 | isocitrate dehydrogenase, NADP-dependent [Escherichia coli MS 182-1] | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111207/F015436.dat&hit=63 | ||||||||
| 6 (F1) | 15803822|ref | NP_289856 | 36717 | 4.98 | 31 | 14 | 46% | 199 | NONE | DNA-directed RNA polymerase subunit alpha [Escherichia coli O157:H7 EDL933] | http://128.59.63.100/mascot/cgi/master_results.pl?file=../data/20111207/F015431.dat&sessionID=FZaghi_81508037746369 | http://128.59.63.100/mascot/cgi/protein_view.pl?file=..%2Fdata%2F20111207%2FF015431.dat;hit=1 | Invirtogel_F1_0001 | |||||
| 227883427|ref | ZP_04001232 | 36691 | 4.98 | 31 | 14 | 46% | 199 | NONE | DNA-directed RNA polymerase subunit alpha [Escherichia coli 83972] | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111207/F015431.dat&hit=6 | ||||||||
| 8 (F5) | 15804735|ref | NP_290776 | 57447 | 4.81 | 15 | 22 | 44% | 243 | NONE | chaperonin GroEL [Escherichia coli O157:H7 EDL933] | http://128.59.63.100/mascot/cgi/master_results.pl?file=../data/20111207/F015437.dat&sessionID=FZaghi_81508037746369 | http://128.59.63.100/mascot/cgi/protein_view.pl?file=..%2Fdata%2F20111211%2FF015491.dat;hit=4 | homologous protein find in different strains of E coli | Invirtogel_F5_0001 | ||||
| 15834378|ref | NP_313151 | 57464 | 4.81 | 15 | 22 | 44% | 243 | NONE | chaperonin GroEL [Escherichia coli O157:H7 str. Sakai] | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111211/F015491.dat&hit=6 | ||||||||
| 110644502|ref | YP_672232 | 57464 | 4.85 | 15 | 22 | 44% | 243 | NONE | chaperonin GroEL [Escherichia coli 536] | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111211/F015491.dat&hit=7 | ||||||||
| 260870948|ref | YP_003237350 | 57709 | 4.85 | 15 | 22 | 44% | 243 | NONE | chaperonin Cpn60 [Escherichia coli O111:H- str. 11128] | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111211/F015491.dat&hit=11 | ||||||||
| 297520397|ref | ZP_06938783 | 35245 | 4.85 | 21 | 11 | 36% | 112 | NONE | chaperonin GroEL [Escherichia coli OP50] | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111211/F015491.dat&hit=51 | ||||||||
| 9 (F7) | 300926981|ref | ZP_07142741 | 77677 | 5.24 | 29 | 37 | 62% | 313 | 100 | translation elongation factor G [Escherichia coli MS 182-1] | http://128.59.63.100/mascot/cgi/master_results.pl?file=../data/20111207/F015430.dat&sessionID=FZaghi_304826335457584 | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111211/F015490.dat&hit=23 | homologous protein find in different strains of E coli | Invirtogel_F7_0001 | ||||
| 218706934|ref | YP_002414453 | 77762 | 5.21 | 28 | 37 | 61% | 311 | 100 | elongation factor G [Escherichia coli UMN026] | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111211/F015490.dat&hit=23 | ||||||||
| 15803853|ref | NP_289887 | 77704 | 5.24 | 28 | 38 | 63% | 324 | 100 | elongation factor G [Escherichia coli O157:H7 EDL933] | http://128.59.63.100/mascot/cgi/protein_view.pl?file=..%2Fdata%2F20111207%2FF015430.dat;hit=1 | ||||||||
| 297517420|ref | ZP_06935806 | 58232 | 5.16 | 28 | 27 | 55% | 211 | 100 | elongation factor G [Escherichia coli OP50] | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111211/F015490.dat&hit=23 | ||||||||
| 297518827|ref | ZP_06937213 | 20222 | 5.33 | 29 | 11 | 78% | 101 | 100 | elongation factor G [Escherichia coli OP50] | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111211/F015490.dat&hit=57 | ||||||||
| 15803116|ref | NP_289147 | 96041 | 5.37 | 34 | 22 | 29% | 100 | 324 | protein disaggregation chaperone [Escherichia coli O157:H7 EDL933] | http://128.59.63.100/mascot/cgi/protein_view.pl?file=../data/20111211/F015490.dat&hit=60 | reported as a mixture with above protein ID | |||||||
| 10 (F9) | no hit | http://128.59.63.100/mascot/cgi/master_results.pl?file=../data/20111211/F015492.dat&sessionID=FZaghi_206677355780492 | Invirtogel_F9_0001 | |||||||||||||||
Mascot Daemon Results
| Spot Number | accession | seq_database | result number | task_UID | hit_mass | hit_score | title_text | result_comment | input_data_file | result_url |
| 1 (E1) | 289448397|ref | NCBInr_081610.fasta | 5 | 2 | 35020 | 83 | PE-PGRS family protein [Mycobacterium tuberculosis CPHL_A] | NOT Accepted. The protien ID does not belong to any of E coli strains. | C:\Documents and Settings\XPMUser\Desktop\Protiomic\farnadZaghi_c\Invirtogel_E1_0001.dat | http://128.59.63.100/mascot/cgi/master_results.pl?file=../data/20111212/F015500.dat |
| 157154780|ref | YP_001461306 | 5 | 4 | 21143 | 64 | putative fimbrial protein [Escherichia coli E24377A] | NOT Accepted. There are only 4 weak peaks matching with spectrum and no matching for the strong peaks. | C:\Documents and Settings\XPMUser\Desktop\Protiomic\farnadZaghi_c\Invirtogel_E1_0001.dat | http://128.59.63.100/mascot/cgi/master_results.pl?file=../data/20111212/F015500.dat | |
| 2 (E3) | 213029173|ref | NCBInr_081610.fasta | 6 | 2 | 10695 | 47 | membrane component of 2-aminoethylphosphonate transporter [Salmonella enterica subsp. enterica serovar Typhi str. 404ty] | NOT Accepted. The protien ID does not belong to any of E coli strains. | C:\Documents and Settings\XPMUser\Desktop\Protiomic\farnadZaghi_c\Invirtogel_E3_0001.dat | http://128.59.63.100/mascot/cgi/master_results.pl?file=../data/20111212/F015501.dat |
| 3 (E5) | 46143512|ref | NCBInr_081610.fasta | 7 | 2 | 109967 | 44 | COG2931: RTX toxins and related Ca2+-binding proteins [Actinobacillus pleuropneumoniae serovar 1 str. 4074] | NOT Accepted. The protien ID does not belong to any of E coli strains. | C:\Documents and Settings\XPMUser\Desktop\Protiomic\farnadZaghi_c\Invirtogel_E5_0001.dat | http://128.59.63.100/mascot/cgi/master_results.pl?file=../data/20111212/F015502.dat |
| 4 (E7) | 194334347|ref | NCBInr_081610.fasta | 8 | 2 | 27113 | 58 | Uroporphyrinogen-III synthase [Prosthecochloris aestuarii DSM 271] | NOT Accepted. The protien ID does not belong to any of E coli strains. | C:\Documents and Settings\XPMUser\Desktop\Protiomic\farnadZaghi_c\Invirtogel_E7_0001.dat | http://128.59.63.100/mascot/cgi/master_results.pl?file=../data/20111212/F015503.dat |
Keywords: E.coli protein expression, protein fragments, 2D gel electrophoresis, MALDI, limited proteolysis
(abv) matrix-assisted laser desorption/ionization mass spectrometry (MALDI-MS)
Personal/Lab Info
User:Farnad_Zaghi,User:Julius_B._Lucks, User:Jason_R._Kelly and User:Reshma_P._Shetty.