User:Jessie Wang: Assignments
Thinking M13
1. M13 Redesign Considerations
| Protein | Function | Re-engineering |
|---|---|---|
| I | Assembly | Mutate the C terminus of the protein and see how that affects interactions with P4 and channel opening |
| II | Replication of DNA + strand | Mutate parts of the DNA - strand to see if P2 still nicks it. It would be interesting to find a particular motif to which it binds because P2 can then be used to nick other pieces of DNA. |
| III | Phage tail protein (5 copies) | Remove 3 copies of the gene and see how that affects M13 |
| IV | Assembly | Alter the N terminus to make it interact more strongly with the C terminus of P1 and P11 and observe effects |
| V | binds ssDNA | Alter the gene so that it produces more/less of the protein to generate a curve of ssDNA vs dsDNA concentration. |
| VI | Phage tail protein (5 copies) | Make p6 protrude more and see if changes the behavior of M13 |
| VII | Phage head protein (5 copies) | Withdraw P7 even more to observe changes, if any |
| VIII | Phage coat proteins (2700 copies) | Remove its ability to replace p5 |
| IX | Phage head protein (5 copies) | Add more copies of p9 |
| X | DNA replication | Increase production of p10 and see how that would also affect p2 |
| XI | Assembly | Mutate the C terminus of the protein and see how that affects interactions with P4 and channel opening |
2. M13's Lovely Relatives
M13's closest evolutionary relatives are the fl and fd phages. While they are all filamentous, ssDNA phages with an F pilus. There are a few differences when it comes to surface proteins, but in general, a lot of their genes are homologous.
3. BBa_M1307 = biological part?
Biological parts are biological tools, building bricks, things like particular genes and certain DNA sequences that can be assembled to make a particular biological machine within a biological organism. BBa_M1307 is a biological part because it has been engineered so that it can be utilized as a building block. The addition of a Kanamycin-resistant gene and an origin of replication allows the M13 genome to be transformed into a bacteria. Once inside a bacteria, the genome can be easily reproduced which means that we can easily re-engineer the genome to do what we want it to do and then have the bacteria reproduce the results.
Of Agarose Gels and Plaques
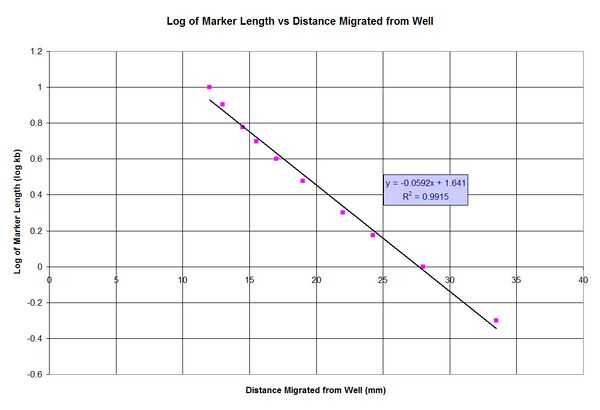
1. Using the Agarose Gel to Create Graphs
From the graph:
y = -0.0592x + 1.641
y = log(size)
x = distance from well
For the cut vector, x = 12 mm
log(size) = -0.0592(12) + 1.641
log(size) = 0.9306
size = 10^0.9306
size = 8.523 kb
The given length is 8.669 kb, which is a little more than what we calculated. This is probably just due to the errors in curve fitting, since our distance measurements were not absolutely exact.
2. Plaquing
Titer = 10^12 units/ml
Dilution = 10^-8
Volume = 10 ul = .01 ml
Plaques = Titer x Dilution x Volume
Plaques = 10^12 units/ml x 10^-8 x .01 ml
Plaques = 100 units
Should expect the same number for DH5, given that DH5 is M13 compatible.
3. Student Writing Survey
Done and sent!
4. "Synthetic biology is about engineering while genetic engineering is about biology."
Synthetic biology and genetic engineering differ in the fundamental concept behind the two. As The New Atlantis article states, genetic engineering is all about moving genes around in a way that we'll help us study something or fix something; it is about revising what we already have. Synthetic biology, on the other hand, is about creating something new with genes as the building blocks or making what we have better; it is about starting from scratch or editing with a whole library of tools. Synthetic biology incorporates a broader set of purposes with more of an engineering focus, goals like re-engineering a genome so that it is more efficient, engineering genes for a new purpose, or creating custom-made, artificial genes. Synthetic biologists envision organisms as machines with genes as parts that can be assembled. Synthetic biology is all about "building" whereas genetic engineering is about "learning," "using," or "moving". And that is the fundamental difference between engineering and science.
Of Bacterial Transformations
1. Transformation Calculation
10^9 units/ug x 1 ug/1000 ng x 1/1000 = 1000 colonies
50 colonies x 1 ug/10^9 units x 100 x 1/2 ul = 2.5 x 10^-6 ug/ul
2. Transformation Outcomes
- Outcome 1: No colonies on all the plates -- Either the transformations or the ligations didn't work.
- Outcome 2: Thousands of colonies on all the plates -- The plates were contaminated or the plates were did not have Kanamycin.
- Outcome 3: approximately the same number of colonies on the backbone+ligase+kill cut as the backbone+insert+ligase+kill cut -- The kill cut digestion didn't work.
3. M13K07 Plasmid Map
Check other printout
| Diagnostic digest 1 | plasmid with insert | plasmid no insert |
|---|---|---|
| Enzyme(s) used | ||
| Buffer used | ||
| Temperature | ||
| Predicted fragments | ||
| Diagnostic digest 2 | ||
| Enzyme(s) used | ||
| Buffer used | ||
| Temperature | ||
| Predicted fragments |
4. Plaque Assay
Titer = Plaques / (Dilution * Volumes)
- M13K07:
- Titer = 65 / (10^-6 * 10 ul * (1 ml / 10^3 ul))
- Titer = 6.5 x 10^9 plaque forming units/ml
- E4:
- Titer = 704 / (10^-6 * 10 ul * (1 ml / 10^3 ul))
- Titer = 7.04 x 10^10 plaque forming units/ml
E4 had bigger plaques. This may be due to the added glutamic acids on the N-terminus of p8. Perhaps this causes the virus to slow the growth of the bacteria even more. (The virus is more robust.) That would then make the plaques bigger.
5. Refactoring Bacteriophage T7
Read.
Examining Candidate Clones
1. Ligation Results
| Transformations | Colonies | Should Expect | What might have happened |
|---|---|---|---|
| 5 ng original M13K07 plasmid | 872 | lots | Seems like it went fine |
| bkb, no ligase | 0 | none | Seems like it went fine |
| bkb + ligase | 5 | a few | Seems like it went fine. A few were probably not completely digested by the killcut |
| bkb + insert + ligase | 2 | more | The insert probably wasn't too good for the cell, and so it couldn't take it up very readily. Either that or the ligation might not have been so good. |
Transformation efficiency:
872 colonies / 5 ng DNA * 1000 ng / 1 ug
= 174400 colonies / ug DNA
Based on the transformation results, we can assume that nothing went wrong with the competent cells because most of the plates appeared as they should, especially the positive control. We can also assume that the kill cut worked because there were very few colonies on the bkb + ligase plate. However, what we don't know is whether the ligation worked well or whether the insert was bad for the cell since we only got two colonies for the bkb + insert + ligase plate. Also, from looking at the agarose gel, it's a little hard to tell whether it worked. The second digestion for one of the plasmids seems right, but the first digestion doesn't. The other plasmid has one band where it's supposed to be, but you can't see the second band. So it's most likely that it didn't quite work...
