User:Gabriel Berriz/Notebook/ICBP45/Overview
EXPERIMENTAL DESIGN/CONFOUNDERS
I will organize the following description of the experimental design according to the dimensions of the resulting data arrays. These arrays have 6 dimensions, and they are divided according to "subassay", so this section will be organized according to the following headings (of which all but the first one are array dimensions):
- subassay
- (cell_line, repno) (this a "composite dimension", whose allowable levels are pairs of values, as described in more detail below)
- ligand_name
- ligand_concentration
- time
- signal
- stat
See ICBP45 plate layout.
0. subassay [unitless]
- 2 "subassays" per assay:
- GF ("growth factors"), 4 96-well plates/subassay
- GF1
- GF2
- GF3
- GF4
- CK ("cytokines"), 2 96-well plates/subassay
- CK1
- CK2
...but, due to the details of the experimental design for the CK subassay (see below), it is useful to think of these 2 plates as 4 "(left or right) half- (or 48-well) plates)": - CK1-L
- CK1-R
- CK2-L
- CK2-R
- GF ("growth factors"), 4 96-well plates/subassay
- For both GF and CK subassays:
- 4 microscopy fields/well
- 3 channels/field (1 filter wavelength/channel)
- The signals (discussed further below) are paired (two are used per well)
- All the wells in each plate for the GF subassay get the same pair of signal probes, as follows:
- GF1 ('pAkt-m-488', 'pErk-r-647')
- GF2 ('pErk-m-488', 'pAkt-r-647')
- GF3 ('pJNK-m-488', 'pP38-r-647')
- GF4 ('pP38-m-488', 'pJNK-r-647')
- all the wells in each (left or right) *half-plate* for the CK subassay get the same pair of probes:
- CK1-L ('NF-κB-m-488', 'STAT1-r-647')
- CK1-R ('pErk-m-488', 'STAT3-r-647')
- CK2-L ('STAT1-r-488', 'NF-κB-m-647')
- CK2-R ('STAT3-r-488', 'pErk-m-647')
- All the wells in each plate for the GF subassay get the same pair of signal probes, as follows:
1. (cell_line, repno) [unitless]
This is a composite dimension, whose allowed levels are pairs of identifiers, where the first element of the pair is the name of a cell line, and the second one is best though of as an internal replica number indicator.
The first column in the table below shows the levels for this dimensions; the next two columns shows synonyms that were used in earlier internal releases of the data; the last column, when not empty, either indicates the few cases where the same cell line was used in multiple assays, or the one assay (20110427_MDAMB436) for which there is only one subassay (GF).
| 1. | ('HCC1187', ('0',)) | 20100924_HCC1187 | HCC1187 | |
| 2. | ('HCC1806', ('0',)) | 20100925_HCC1806 | HCC1806 | |
| 3. | ('CAMA1', ('0',)) | 20100928_CAMA1 | CAMA1 | |
| 4. | ('HCC1954', ('0',)) | 20101004_HCC1954 | HCC1954 | |
| 5. | ('AU565', ('0',)) | 20101005_AU565 | AU565 | |
| 6. | ('HCC1569', ('0',)) | 20101006_HCC1569 | HCC1569 | |
| 7. | ('BT20', ('0',)) | 20101007_BT20 | BT20 | |
| 8. | ('HCC38', ('0',)) | 20101008_HCC38 | HCC38 | |
| 9. | ('MCF7', ('0',)) | 20101012_MCF7 | MCF7__a | * |
| 10. | ('HCC70', ('0',)) | 20101018_HCC70 | HCC70 | |
| 11. | ('RA', ('0',)) | 20101021_RA_Rob | RA | |
| 12. | ('N', ('0',)) | 20101022_N_Rob | N | |
| 13. | ('HCC1419', ('0',)) | 20101025_HCC1419 | HCC1419 | |
| 14. | ('HCC1937', ('0',)) | 20101122_HCC1937 | HCC1937 | |
| 15. | ('SKBR3', ('0',)) | 20101202_SKBR3 | SKBR3__a | * |
| 16. | ('MCF10A', ('0',)) | 20101206_MCF10A | MCF10A | |
| 17. | ('SKBR3', ('1',)) | 20101210_SKBR3 | SKBR3__b | * |
| 18. | ('HCC1395', ('0',)) | 20101213_HCC1395 | HCC1395 | |
| 19. | ('ZR751', ('0',)) | 20101215_ZR751 | ZR751 | |
| 20. | ('HCC202', ('0',)) | 20101216_HCC202 | HCC202 | |
| 21. | ('HCC1428', ('0',)) | 20101221_HCC1428 | HCC1428 | |
| 22. | ('MDAMB231', ('0',)) | 20101222_MDAMB231 | MDAMB231__a | * |
| 23. | ('ZR7530', ('0',)) | 20110128_ZR7530 | ZR7530 | |
| 24. | ('SKBR3', ('2',)) | 20110201_SKBR3 | SKBR3__c | * |
| 25. | ('MCF7', ('1',)) | 20110210_MCF7 | MCF7__b | * |
| 26. | ('MDAMB175', ('0',)) | 20110308_MDAMB175 | MDAMB175 | |
| 27. | ('HCC1500', ('0',)) | 20110310_HCC1500 | HCC1500 | |
| 28. | ('MDAMB453', ('0',)) | 20110311_MDAMB453 | MDAMB453 | |
| 29. | ('MDAMB231', ('1',)) | 20110317_MDAMB231 | MDAMB231__b | * |
| 30. | ('Hs578T', ('0',)) | 20110318_Hs578T | Hs578T | |
| 31. | ('T47D', ('0',)) | 20110321_T47D | T47D | |
| 32. | ('BT549', ('0',)) | 20110322_BT549 | BT549 | |
| 33. | ('MDAMB361', ('0',)) | 20110324_MDAMB361 | MDAMB361 | |
| 34. | ('MDAMB157', ('0',)) | 20110325_MDAMB157 | MDAMB157 | |
| 35. | ('UACC893', ('0',)) | 20110330_UACC893 | UACC893 | |
| 36. | ('MCF10F', ('0',)) | 20110414_MCF10F | MCF10F | |
| 37. | ('MCF12A', ('0',)) | 20110415_MCF12A | MCF12A | |
| 38. | ('BT474', ('0',)) | 20110418_BT474 | BT474 | |
| 39. | ('UACC812', ('0',)) | 20110420_UACC812 | UACC812 | |
| 40. | ('184B5', ('0',)) | 20110421_184B5 | 184B5 | |
| 41. | ('MDAMB415', ('0',)) | 20110421_MDAMB415 | MDAMB415 | |
| 42. | ('MDAMB436', ('0',)) | 20110427_MDAMB436 | MDAMB436 | (GF only) |
| 43. | ('BT-483', ('0',)) | 20110502_BT-483 | BT-483 | |
| 44. | ('MDAMB134', ('0',)) | 20110517_MDAMB134 | MDAMB134 |
Summary of replicated assays
| ('MCF7', ('0',)) | 20101012_MCF7 | MCF7__a |
| ('MCF7', ('1',)) | 20110210_MCF7 | MCF7__b |
| ('MDAMB231', ('0',)) | 20101222_MDAMB231 | MDAMB231__a |
| ('MDAMB231', ('1',)) | 20110317_MDAMB231 | MDAMB231__b |
| ('SKBR3', ('0',)) | 20101202_SKBR3 | SKBR3__a |
| ('SKBR3', ('1',)) | 20101210_SKBR3 | SKBR3__b |
| ('SKBR3', ('2',)) | 20110201_SKBR3 | SKBR3__c |
- cell lines:
- 184B5
- AU565
- BT20
- BT474
- BT-483
- BT549
- CAMA1
- HCC1187
- HCC1395
- HCC1419
- HCC1428
- HCC1500
- HCC1569
- HCC1806
- HCC1937
- HCC1954
- HCC202
- HCC38
- HCC70
- Hs578T
- MCF10A
- MCF10F
- MCF12A
- MCF7
- MDAMB134
- MDAMB157
- MDAMB175
- MDAMB231
- MDAMB361
- MDAMB415
- MDAMB436
- MDAMB453
- N_Rob (non-standard)
- RA_Rob (non-standard)
- SKBR3
- T47D
- UACC812
- UACC893
- ZR751
- ZR7530
2. ligand_name [unitless]
- levels:
- GF:
- VEGFF
- NGF
- EGF
- INS
- EPR
- IGF-1
- BTC
- IGF-2
- HRG
- SCF
- CTRL
- HGF
- FGF1
- PDGFBB
- FGF2
- EFNA1
- CK:
- LPS
- IL-1α
- IL-6
- CTRL
- IFN-α
- IFN-γ
- TNF-α
- IL-2
- GF:
- the two rosters above are disjoint except for the pseudo-ligand "CTRL", which represents the "null ligand" ("DMSO")
- possible synonyms:
- IL-1a = IL-1α
- IFN-a = IFN-α
- IFN-g = IFN-γ
- TNF-a = TNF-α
- CTRL-GF-1 = CTRL
- CTRL-GF-100 = CTRL
3. ligand_concentration [ng/ml]
- levels:
- 0
- 1
- 100
- data for ligand_concentration=0 is derived from the data for the wells with ligand_name=CTRL (procedure described below)
4. time [min]
- levels:
- 0
- 10
- 30
- 90
- data for time=0 is derived from the data for the wells with ligand_name=CTRL (procedure described below)
5. signal [unitless]
- levels:
- GF:
- pAkt-m-488
- pErk-r-647
- pErk-m-488
- pAkt-r-647
- pJNK-m-488
- pP38-r-647
- pP38-m-488
- pJNK-r-647
- CK:
- NF-κB-m-488
- STAT1-r-647
- pErk-m-488
- STAT3-r-647
- STAT1-r-488
- NF-κB-m-647
- STAT3-r-488
- pErk-m-647
- GF:
- pErk-m-488 is the only signal shared by both GF and CK
- possible synonyms:
- NF-kB-m-488 = NF-κB-m-488
- NF-kB-m-647 = NF-κB-m-647
- pErk-CK-m-488 = pErk-m-488
- pErk-CK-m-647 = pErk-m-647
- CTRL-CK-L-1 = CTRL
- CTRL-CK-L-100 = CTRL
- CTRL-CK-R-1 = CTRL
- CTRL-CK-R-100 = CTRL
- each signal identifier is a 3-part composite:
<target of primary antibody>-<species of primary antibody>-<wavelength of fluorophore on secondary antibody>- nonetheless, at the model level, we are treating them as "opaque" labels, i.e., as irreducible levels
- '-' is a poor choice of separator for this composite key, due to cases like NF-κB-m-488 (e.g. this example should be parsed as NF-κB, m, 488; e.g., in Python, the simplest way to get around this problem is:
target, species, wavelength = signal.rsplit('-', 2)
- with only the exception of NF-κB-488/647, the statistic that serves as the starting point for subsequent analysis is the mean intensity per cell ('Whole_w530 (Mean)' and 'Whole_w685 (Mean)'). For NF-κB-488/647 the statistic used is the ratio of the mean nuclear intensity to the mean cytoplasmic intensity ('Nucleus_w530 (Integrated)'/'Cyto_w530 (Mean)', ('Nucleus_w685 (Integrated)'/'Cyto_w685 (Mean)')).
The following subsections do not correspond to dimensions proper; there are only additional remarks on the components of the signal identifiers.
5.1 target of primary antibody
- possible values:
- GF:
- pAkt
- pErk
- pJNK
- pP38
- CK:
- NF-κB
- STAT1
- pErk
- STAT3
- GF:
5.2 species of primary antibody
- possible values:
- m = mouse
- r = rabbit
5.3 wavelength of fluorophore on secondary antibody
- possible values:
- 488 (corresponds to 530 channel)
- 647 (corresponds to 685 channel)
6. stat [unitless]
- levels:
- mean
- stddev
ORGANIZATION OF PRIMARY DATA
- The primary microscopy files for these assays (the TIFF files plus other ancillary files generated by the microscope's software) are stored under
/research.files/ImStor/ICBP45/CNI - ImageWoRx scans/
- I created an alternative directory tree that uses symlinks to point to the TIFF files. The structure of this alternative tree can be read off the path components (after <ROOT>) for the directories containing the primary files:
<ROOT> / <ASSAY> / <PLATE> / <WELL> / <FIELD> / <SYMLINK_TO_TIFF>
For example,
- I created an alternative directory tree that uses symlinks to point to the TIFF files. The structure of this alternative tree can be read off the path components (after <ROOT>) for the directories containing the primary files:
<ROOT> ├── 20100924_HCC1187 │ ├── CK1 │ │ ├── A01 │ │ │ ├── 1 │ │ │ │ ├── CK1_A01_1_w460.tif -> <ROOT_0>/20100924_HCC1187/CK1/TIFF/CK1_A01_1_w460.tif │ │ │ │ ├── CK1_A01_1_w530.tif -> <ROOT_0>/20100924_HCC1187/CK1/TIFF/CK1_A01_1_w530.tif │ │ │ │ └── CK1_A01_1_w685.tif -> <ROOT_0>/20100924_HCC1187/CK1/TIFF/CK1_A01_1_w685.tif . . . . . . . . . . . .
- where <ROOT> is
<ROOT_0>/linkfarm
and <ROOT_0> is
/research.files/ImStor/ICBP45/CNI - ImageWoRx scans
- where <ROOT> is
- In addition to simplifying the logic required for iterating over the image files, the alternative symlink-terminated tree offers the opportunity to fix various errors and inconsistencies in the naming of original files. For example, plate CK1 for the 20101216_HCC202 assay was inadvertently rotated 180° when it was scanned by the microscope, so the resulting files are systematically misnamed. The symlinks to these files reflect the necessary corrections:
<ROOT> . . . . . . . . . ├── 20101216_HCC202 │ ├── CK1 │ │ ├── A01 │ │ │ ├── 1 │ │ │ │ ├── CK1_A01_1_w460.tif -> <ROOT_0>/20101216_HCC202/CK1/TIFF/CK1_H12_3_w460.tif │ │ │ │ ├── CK1_A01_1_w530.tif -> <ROOT_0>/20101216_HCC202/CK1/TIFF/CK1_H12_3_w530.tif │ │ │ │ └── CK1_A01_1_w685.tif -> <ROOT_0>/20101216_HCC202/CK1/TIFF/CK1_H12_3_w685.tif . . . . . . . . . . . . . .
- The symlink-based scheme was also used to implement the censoring of subassays of substandard quality, as was the case for 20110427_MDAMB436/CK;
- In this case, the symlinks were simply moved to a subtree under .ARCHIVE:
- The symlink-based scheme was also used to implement the censoring of subassays of substandard quality, as was the case for 20110427_MDAMB436/CK;
% tree -a -L 1 <ROOT>/linkfarm/20110427_MDAMB436
<ROOT>/linkfarm/20110427_MDAMB436
├── .ARCHIVE
├── GF1
├── GF2
├── GF3
└── GF4
% tree -a -L 2 <ROOT>/linkfarm/20110427_MDAMB436/.ARCHIVE
<ROOT>/linkfarm/20110427_MDAMB436/.ARCHIVE
└── 111001S
├── CK1
└── CK2
% tree -a <ROOT>/linkfarm/20110427_MDAMB436/.ARCHIVE
<ROOT>/linkfarm/20110427_MDAMB436/.ARCHIVE
└── 111001S
├── CK1
│ ├── A01
│ │ ├── 1
│ │ │ ├── CK1_A01_1_w460.tif -> <ROOT>/20110427_MDAMB436/CK1/TIFF/plate5_A01_1_w460.tif
│ │ │ ├── CK1_A01_1_w530.tif -> <ROOT>/20110427_MDAMB436/CK1/TIFF/plate5_A01_1_w530.tif
│ │ │ └── CK1_A01_1_w685.tif -> <ROOT>/20110427_MDAMB436/CK1/TIFF/plate5_A01_1_w685.tif
│ │ ├── 1.sdc
│ │ │ ├── Data.h5
│ │ │ └── ExpDesign.xml
│ │ ├── 2
│ │ │ └── ...
│ │ ├── 2.sdc
│ │ │ └── ...
│ │ ├── 3
│ │ │ └── ...
│ │ ├── 3.sdc
│ │ │ └── ...
│ │ ├── 4
│ │ │ └── ...
│ │ └── 4.sdc
│ │ └── ...
│ ├── A02
│ │ └── ...
. . .
. . .
. . .
│ ├── H11
│ │ └── ...
│ └── H12
│ └── ...
└── CK2
├── A01
│ └── ...
├── A02
│ └── ...
. .
. .
. .
├── H11
│ └── ...
└── H12
└── ...
DATA PROCESSING
This section describes the processing performed on the raw data (TIFF files), and stored in the file [Dropbox]/Breast Cancer Ligand Reponse Screen/Mario Data/20120106_h5 files/icbp45.h5.
- A command-line-oriented form of ImageRail's (IR) segmentation and characterization functionality was run for all the TIFF files, with segmentation parameters as recorded in segmentation_params.tsv. Note: a single set of segmentation parameters was used for each assay. E.g., for 20100924_HCC1187, whose record in segmentation_params.tsv is
20100924_HCC1187 7500 1500 1000
the ImageRail runs were like this:
(first some groundwork)
% IR_BASE=$(pwd)
% CLASSPATH="${IR_BASE}/*:${IR_BASE}/jai/*"
% LIBPATH="${IR_BASE}:${IR_BASE}/jai"
% SEGPARAM0=7500
% SEGPARAM1=1500
% SEGPARAM2=1000
% INPUTPATH=scans/linkfarm/20100924_HCC1187/GF4/E06/1
% OUTPUTPATHSTEM=newdir/tmp
% OUTPUTPATH="${OUTPUTPATHSTEM}.sdc"
% ls -ltr $INPUTPATH
total 5
lrwxrwx--- 1 gfb2 ImStor_ICBP45 96 Sep 30 13:18 GF4_E06_1_w530.tif -> /research.files/ImStor/ICBP45/CNI - ImageWoRx scans/20100924_HCC1187/GF4/TIFF/GF4_E06_1_w530.tif
lrwxrwx--- 1 gfb2 ImStor_ICBP45 96 Sep 30 13:18 GF4_E06_1_w460.tif -> /research.files/ImStor/ICBP45/CNI - ImageWoRx scans/20100924_HCC1187/GF4/TIFF/GF4_E06_1_w460.tif
lrwxrwx--- 1 gfb2 ImStor_ICBP45 96 Sep 30 13:18 GF4_E06_1_w685.tif -> /research.files/ImStor/ICBP45/CNI - ImageWoRx scans/20100924_HCC1187/GF4/TIFF/GF4_E06_1_w685.tif
% rm -rf $( dirname $OUTPUTPATH )
% mkdir $( dirname $OUTPUTPATH )
Now, the actual run:
% java -Xmx1000M -cp "${CLASSPATH}" -D"java.library.path=${LIBPATH}" run.Segment "${INPUTPATH}" "${OUTPUTPATHSTEM}" 0 0 0 0 0 $SEGPARAM0 $SEGPARAM1 $SEGPARAM2 Centroid
*****Processing Input Directory: 1
*** Creating new ImageRail_IO
Jun 27, 2012 4:29:50 PM ncsa.hdf.hdf5lib.H5 <clinit>
INFO: HDF5 library: jhdf5 resolved to: libjhdf5.so; successfully loaded from java.library.path
[Fatal Error] ExpDesign.xml:1:1: Premature end of file.
Initializing the Hashtable
___Found 0 Samples in this project___
...Successfuly indexed 0 Samples
________________
w460
w530
w685
Step 1: Computing Euclidean Maps
Step 2: Finding Ultimate Eroded Points
Num Nuclei: 1666
-->> Performing Feature Computations
1666x26
------------ Caching cell data Matrix and Coordinates to HDF file: ------------
(Note that the "[Fatal Error]" above is actually harmless; it should really be a warning, at most.) A look at the results (NB: the output of h5dump has been abridged, as indicated by ellipses):
% ls -ltr $OUTPUTPATH
total 652
-rw-rw-r-- 1 gfb2 gfb2 261 Jun 27 16:29 ExpDesign.xml
-rw-rw-r-- 1 gfb2 gfb2 200109 Jun 27 16:29 Data.h5
% h5dump ${OUTPUTPATH}/Data.h5
HDF5 "newdir/tmp.sdc/Data.h5" {
GROUP "/" {
GROUP "Children" {
GROUP "0" {
GROUP "Children" {
GROUP "0" {
GROUP "Children" {
}
GROUP "Data" {
DATASET "coords_centroids" {
DATATYPE H5T_STD_I32LE
DATASPACE SIMPLE { ( 1666 ) / ( 1666 ) }
DATA {
(0): 2050, 66565, 78851, 84995, 91141, 97284, 107522,
(7): 125956, 213002, 276484, 313346, 379910, 391172,
...
(1655): 205818, 229371, 319485, 332797, 344060, 402426,
(1661): 575485, 869372, 954365, 1019894, 1039351
}
ATTRIBUTE "dataType" {
DATATYPE H5T_STRING {
STRSIZE 14;
STRPAD H5T_STR_NULLTERM;
CSET H5T_CSET_ASCII;
CTYPE H5T_C_S1;
}
DATASPACE SIMPLE { ( 1 ) / ( 1 ) }
DATA {
(0): "H5T_NATIVE_INT"
}
}
ATTRIBUTE "dim0" {
DATATYPE H5T_STRING {
STRSIZE 5;
STRPAD H5T_STR_NULLTERM;
CSET H5T_CSET_ASCII;
CTYPE H5T_C_S1;
}
DATASPACE SIMPLE { ( 1 ) / ( 1 ) }
DATA {
(0): "cells"
}
}
}
DATASET "feature_values" {
DATATYPE H5T_IEEE_F32LE
DATASPACE SIMPLE { ( 1666, 26 ) / ( 1666, 26 ) }
DATA {
(0,0): 2, 2, 96128, 3224.11, 234629, 2439.49, 14184,
(0,7): 143.615, 197246, 12368.5, 195163, 5674.15, 8855,
...
(1665,20): 7.17585e+06, 21999.2, 4.49477e+06, 6932.38,
(1665,24): 213422, 230.146
}
ATTRIBUTE "dataType" {
DATATYPE H5T_STRING {
STRSIZE 16;
STRPAD H5T_STR_NULLTERM;
CSET H5T_CSET_ASCII;
CTYPE H5T_C_S1;
}
DATASPACE SIMPLE { ( 1 ) / ( 1 ) }
DATA {
(0): "H5T_NATIVE_FLOAT"
}
}
ATTRIBUTE "dim0" {
DATATYPE H5T_STRING {
STRSIZE 5;
STRPAD H5T_STR_NULLTERM;
CSET H5T_CSET_ASCII;
CTYPE H5T_C_S1;
}
DATASPACE SIMPLE { ( 1 ) / ( 1 ) }
DATA {
(0): "cells"
}
}
ATTRIBUTE "dim1" {
DATATYPE H5T_STRING {
STRSIZE 14;
STRPAD H5T_STR_NULLTERM;
CSET H5T_CSET_ASCII;
CTYPE H5T_C_S1;
}
DATASPACE SIMPLE { ( 1 ) / ( 1 ) }
DATA {
(0): "feature_values"
}
}
}
}
GROUP "Meta" {
DATASET "Field_ID" {
DATATYPE H5T_STRING {
STRSIZE 6;
STRPAD H5T_STR_SPACEPAD;
CSET H5T_CSET_ASCII;
CTYPE H5T_C_S1;
}
DATASPACE SIMPLE { ( 1 ) / ( 1 ) }
DATA {
(0): "p0w0f0"
}
}
DATASET "Height_Width_Channels" {
DATATYPE H5T_STD_I32LE
DATASPACE SIMPLE { ( 3, 1 ) / ( 3, 1 ) }
DATA {
(0,0): 1024,
(1,0): 1024,
(2,0): 3
}
ATTRIBUTE "dataType" {
DATATYPE H5T_STRING {
STRSIZE 18;
STRPAD H5T_STR_NULLTERM;
CSET H5T_CSET_ASCII;
CTYPE H5T_C_S1;
}
DATASPACE SIMPLE { ( 1 ) / ( 1 ) }
DATA {
(0): "H5T_NATIVE_INTEGER"
}
}
}
DATASET "compartment_names" {
DATATYPE H5T_STRING {
STRSIZE 8;
STRPAD H5T_STR_SPACEPAD;
CSET H5T_CSET_ASCII;
CTYPE H5T_C_S1;
}
DATASPACE SIMPLE { ( 1 ) / ( 1 ) }
DATA {
(0): "Centroid"
}
}
DATASET "feature_names" {
DATATYPE H5T_STRING {
STRSIZE 25;
STRPAD H5T_STR_SPACEPAD;
CSET H5T_CSET_ASCII;
CTYPE H5T_C_S1;
}
DATASPACE SIMPLE { ( 26 ) / ( 26 ) }
DATA {
(0): "Coordinate_X ",
(1): "Coordinate_Y ",
(2): "Cyto_w460 (Integrated) ",
(3): "Cyto_w460 (Mean) ",
(4): "Cyto_w530 (Integrated) ",
(5): "Cyto_w530 (Mean) ",
(6): "Cyto_w685 (Integrated) ",
(7): "Cyto_w685 (Mean) ",
(8): "Nucleus_w460 (Integrated)",
(9): "Nucleus_w460 (Mean) ",
(10): "Nucleus_w530 (Integrated)",
(11): "Nucleus_w530 (Mean) ",
(12): "Nucleus_w685 (Integrated)",
(13): "Nucleus_w685 (Mean) ",
(14): "Ratio_nuc/cyt_w460 ",
(15): "Ratio_nuc/cyt_w530 ",
(16): "Ratio_nuc/cyt_w685 ",
(17): "Size_cyto ",
(18): "Size_nucleus ",
(19): "Size_whole ",
(20): "Whole_w460 (Integrated) ",
(21): "Whole_w460 (Mean) ",
(22): "Whole_w530 (Integrated) ",
(23): "Whole_w530 (Mean) ",
(24): "Whole_w685 (Integrated) ",
(25): "Whole_w685 (Mean) "
}
}
}
}
}
GROUP "Data" {
}
GROUP "Meta" {
DATASET "Plate_Well" {
DATATYPE H5T_STD_I32LE
DATASPACE SIMPLE { ( 2, 1 ) / ( 2, 1 ) }
DATA {
(0,0): 0,
(1,0): 0
}
ATTRIBUTE "dataType" {
DATATYPE H5T_STRING {
STRSIZE 18;
STRPAD H5T_STR_NULLTERM;
CSET H5T_CSET_ASCII;
CTYPE H5T_C_S1;
}
DATASPACE SIMPLE { ( 1 ) / ( 1 ) }
DATA {
(0): "H5T_NATIVE_INTEGER"
}
}
}
DATASET "Sample_ID" {
DATATYPE H5T_STRING {
STRSIZE 21;
STRPAD H5T_STR_SPACEPAD;
CSET H5T_CSET_ASCII;
CTYPE H5T_C_S1;
}
DATASPACE SIMPLE { ( 1 ) / ( 1 ) }
DATA {
(0): "p0w0_t20120119_165332"
}
}
DATASET "Sample_TYPE" {
DATATYPE H5T_STRING {
STRSIZE 12;
STRPAD H5T_STR_SPACEPAD;
CSET H5T_CSET_ASCII;
CTYPE H5T_C_S1;
}
DATASPACE SIMPLE { ( 1 ) / ( 1 ) }
DATA {
(0): "ImageRail_v1"
}
}
}
}
}
GROUP "Data" {
}
GROUP "Meta" {
DATASET "ExpDesign.xml" {
DATATYPE H5T_STD_I8LE
DATASPACE SIMPLE { ( 261 ) / ( 261 ) }
DATA {
(0): 60, 63, 120, 109, 108, 32, 118, 101, 114, 115, 105, 111, 110,
(13): 61, 34, 49, 46, 48, 34, 63, 62, 10, 60, 115, 100, 99, 117, 98,
(28): 101, 32, 120, 115, 105, 58, 115, 99, 104, 101, 109, 97, 76,
(41): 111, 99, 97, 116, 105, 111, 110, 61, 39, 104, 116, 116, 112,
(54): 58, 47, 47, 112, 105, 112, 101, 108, 105, 110, 101, 46, 109,
(67): 101, 100, 46, 104, 97, 114, 118, 97, 114, 100, 46, 101, 100,
(80): 117, 47, 105, 109, 97, 103, 101, 114, 97, 105, 108, 45, 109,
(93): 101, 116, 97, 100, 97, 116, 97, 45, 49, 46, 48, 32, 105, 109,
(107): 97, 103, 101, 114, 97, 105, 108, 45, 109, 101, 116, 97, 100,
(120): 97, 116, 97, 45, 49, 46, 48, 46, 120, 115, 100, 39, 32, 120,
(134): 109, 108, 110, 115, 58, 120, 115, 105, 61, 39, 104, 116, 116,
(147): 112, 58, 47, 47, 119, 119, 119, 46, 119, 51, 46, 111, 114,
(160): 103, 47, 50, 48, 48, 49, 47, 88, 77, 76, 83, 99, 104, 101,
(174): 109, 97, 45, 105, 110, 115, 116, 97, 110, 99, 101, 39, 32,
(187): 120, 109, 108, 110, 115, 61, 39, 104, 116, 116, 112, 58, 47,
(200): 47, 112, 105, 112, 101, 108, 105, 110, 101, 46, 109, 101,
(212): 100, 46, 104, 97, 114, 118, 97, 114, 100, 46, 101, 100, 117,
(225): 47, 105, 109, 97, 103, 101, 114, 97, 105, 108, 45, 109, 101,
(238): 116, 97, 100, 97, 116, 97, 45, 49, 46, 48, 39, 62, 10, 60,
(252): 47, 115, 100, 99, 117, 98, 101, 62, 10
}
}
DATASET "PlateCount_PlateSize" {
DATATYPE H5T_STD_I32LE
DATASPACE SIMPLE { ( 2, 1 ) / ( 2, 1 ) }
DATA {
(0,0): 1,
(1,0): 96
}
ATTRIBUTE "dataType" {
DATATYPE H5T_STRING {
STRSIZE 18;
STRPAD H5T_STR_NULLTERM;
CSET H5T_CSET_ASCII;
CTYPE H5T_C_S1;
}
DATASPACE SIMPLE { ( 1 ) / ( 1 ) }
DATA {
(0): "H5T_NATIVE_INTEGER"
}
}
}
}
GROUP "Raw" {
}
}
}- NB: segmentation failed for 7 fields (out of ~105); see Unsegmentable fields; also, the mean intensities for some field/channels is negative, probably due to artefactually elevated background estimates; see Anomalous fields.
- After segmentation, IR computes several features, such as mean pixel intensity per cell per channel, etc. See the h5dump output above, near 'DATASET "feature_values"', for the exact names of these features.
- Potential problems: I know of at least three sources of systematic error in the values produced by IR.
- As an expedient, a single set of threshholds (estimated by Mario by using IR interactively) is used for all the images of each cell line; it is to be expected that for at least some images these parameters are significantly less than optimal.
- IR's segmentation recipe uses uniform thresholds for the entire image, so it will perform suboptimally when the illumination across the image is not uniform; the standard way to solve this problem is to perform some form of background subtraction to equalize the illumination across the image, prior to applying thresholds for segmentation, but as of now, IR does not perform this preprocessing step.
- IR's segmentation algorithm is very sensitive to microscopy artefacts such as bubbles and debris; in fact all the segmentation failures noted under 1.1 appear to be due such artefacts; furthermore, IR does not attempt to detect (and at least report, if not correct) obvious errors, such as cells without cytoplasm.
- I have not attempted to measure/estimate the magnitude of these possible sources of error.
- Each segmentation run generates one directory, having extension .sdc (sdc = Semantic Data Cube), and containing a pair of files, called ExpDesign.xml and Data.h5.
- The 102,528 .sdc directories thus generated were organized in the file structure indicated below:
/files/ImStor/ICBP45/CNI - ImageWoRx scans/linkfarm/20100924_HCC1187/CK1/A01/1.sdc
/files/ImStor/ICBP45/CNI - ImageWoRx scans/linkfarm/20100924_HCC1187/CK1/A01/2.sdc
/files/ImStor/ICBP45/CNI - ImageWoRx scans/linkfarm/20100924_HCC1187/CK1/A01/3.sdc
/files/ImStor/ICBP45/CNI - ImageWoRx scans/linkfarm/20100924_HCC1187/CK1/A01/4.sdc
/files/ImStor/ICBP45/CNI - ImageWoRx scans/linkfarm/20100924_HCC1187/CK1/A02/1.sdc
...
/files/ImStor/ICBP45/CNI - ImageWoRx scans/linkfarm/20110517_MDAMB134/GF4/H11/4.sdc
/files/ImStor/ICBP45/CNI - ImageWoRx scans/linkfarm/20110517_MDAMB134/GF4/H12/1.sdc
/files/ImStor/ICBP45/CNI - ImageWoRx scans/linkfarm/20110517_MDAMB134/GF4/H12/2.sdc
/files/ImStor/ICBP45/CNI - ImageWoRx scans/linkfarm/20110517_MDAMB134/GF4/H12/3.sdc
/files/ImStor/ICBP45/CNI - ImageWoRx scans/linkfarm/20110517_MDAMB134/GF4/H12/4.sdc
- Informally, "confounders" are metadata that do not belong in the projected model for the data, but that we want to keep track of, mostly for possible forensic analyses, e.g. to identify "batch effects".
- This information is collected in the file [Dropbox]/Breast Cancer Ligand Reponse Screen/Mario Data/20111122_h5 files/icbp45_metadata; its first few line look like this:
cell_line,ligand_name,ligand_concentration,time,signal, ,assay,plate,well,channel,antibodyHCC1187,VEGFF,1,10,pAkt-m-488, ,20100924_HCC1187,GF1,A01,530,pAkt|mouse|488HCC1187,VEGFF,1,10,pErk-r-647, ,20100924_HCC1187,GF1,A01,685,pErk|rabbit|647HCC1187,VEGFF,100,10,pAkt-m-488, ,20100924_HCC1187,GF1,A02,530,pAkt|mouse|488HCC1187,VEGFF,100,10,pErk-r-647, ,20100924_HCC1187,GF1,A02,685,pErk|rabbit|647HCC1187,VEGFF,1,30,pAkt-m-488, ,20100924_HCC1187,GF1,A03,530,pAkt|mouse|488HCC1187,VEGFF,1,30,pErk-r-647, ,20100924_HCC1187,GF1,A03,685,pErk|rabbit|647HCC1187,VEGFF,100,30,pAkt-m-488, ,20100924_HCC1187,GF1,A04,530,pAkt|mouse|488HCC1187,VEGFF,100,30,pErk-r-647, ,20100924_HCC1187,GF1,A04,685,pErk|rabbit|647HCC1187,VEGFF,1,90,pAkt-m-488, ,20100924_HCC1187,GF1,A05,530,pAkt|mouse|488 - This confounder metadata is also stored in two arrays, one for each subassay, under the group "confounders" of icbp45.h5. These arrays have the same number of dimensions as do their counterparts data arrays under the "from_IR" group (see below), and the names and levels for all but the last one of these dimensions are the same for confounder and data arrays. In fact, if we interpret all of these arrays as (n-1)-dimensional arrays of 1-d vectors (rather than n-dimensional arrays of scalars), then the confounder and data arrays can be said to have exactly the same shape, although the cells of the confounder arrays contain vectors of integers while the data arrays contain vectors of doubles. (I'll describe the confounder vectors shortly, and describe the data vectors further down.) In other words associated with each cell (data vector) in the data arrays there is a corresponding vector of confounder values, and the matching confounder and data vectors have the same set of coordinates.
- The dimensions of confounder arrays are described in the associated "labels" subgroups, one per subassay. Each "labels" subgroup is a YAML-serialized data structure having the following general form:
[[dimension_0_name,[dimension_0_level_0, dimension_0_level_1, ..., dimension_0_level_n_0]],[dimension_1_name,[dimension_1_level_0, dimension_1_level_1, ..., dimension_1_level_n_1]],...[dimension_d_name,[dimension_d_level_0, dimension_d_level_1, ..., dimension_d_level_n_d]]]
Each element of the tuple is a key-value pair, where the key is the name of a dimension, and the value is a tuple of the levels for that dimension. The ordering of the pairs in this tuple correspond to the ordering of the array's dimensions, and similarly, the ordering of the levels in the value of each pair corresponds to their ordering along the array's dimension. By using this information (both the explicit names as well as their ordering) it is possible to translate between the standard scheme for addressing arrays (as tuples of integer indices), and an addressing scheme based on dimension and level names. This second addressing scheme not only obviates the need to remember the ordering of the dimensions and levels on each array, but in addition it is insensitive to any reordering of the dimensions or the levels. - The components of the confounders vector (which are, in fact, the levels of the last dimension of the confounder array, as described in the "labels" subgroup) are these:
- assay
- plate
- well
- channel (wavelength)
- antibody; this component has three subcomponents (separated by "|"):
- target of primary antibody
- species of primary antibody
- wavelength of fluorescent label on secondary antibody
- The possible values for these various components are encoded as integers (8-bit, unsigned), for the sake of storage efficiency. The subgroup "confounders/keymap" contains a YAML-serialized representation of a tuple whose elements are dictionaries, mapping the confounder values to their local integer id. For example, the second element of this YAML-serialized tuple is
{CK1: 112, CK2: 116, GF1: 2, GF2: 103, GF3: 106, GF4: 109}.
- The dimensions of confounder arrays are described in the associated "labels" subgroups, one per subassay. Each "labels" subgroup is a YAML-serialized data structure having the following general form:
- I computed mean and standard deviations, averaging over all cells for a given well, of the statistic described above (('Whole_w530 (Mean)' and 'Whole_w685 (Mean)'), for all signals, except for signals NF-κB-488/647, for which the statistics I used for this were 'Nucleus_w530 (Integrated)'/'Cyto_w530 (Mean)', ('Nucleus_w685 (Integrated)'/'Cyto_w685 (Mean)') (or NCR, for "nuclear-cytoplasmic ratio"). (The values obtained from this computation are stored in the from_IR arrays in the icbp45.h5 file.)
- In all cases, I computed the mean and standard deviation by invoking the mean and std methods of the appropriate numpy.ndarray objects.
- NB: the numpy.std functions (which is what gets used when one invokes an numpy.ndarray object's std method) computes the square root of the data's second central moment (aka as the "biased sample variance"), given by [math]\displaystyle{ (\sum_{i=1}^n \left(y_i - \overline{y} \right)^2)/n }[/math]. Note that the denominator in this expression is [math]\displaystyle{ n }[/math], not [math]\displaystyle{ n - 1 }[/math].
- When computing the NCR-based statistics, I censored all the datapoints (i.e. cells) for which the cytoplasmic value ('Nucleus_w530 (Integrated)' or 'Nucleus_w685 (Integrated)') was 0. This was done separately for each channel, meaning that the same cell could be censored in one channel but not in the other. [The censoring of these values takes place in the _cull_zeros(d, i) internal function of icbp45_makecube.get_extractor.]
- The "labels" subgroup for the data arrays serve the same function as the "labels" subgroup for the confounder arrays, as described above. They differ only in the name, length, and levels of their last entries. (Namely, they differ in the name and values of the last dimension.)
- In all cases, I computed the mean and standard deviation by invoking the mean and std methods of the appropriate numpy.ndarray objects.
- I averaged and "propagated" the data from ligand_name=CTRL wells to the array cells corresponding to time=0 or/and ligand_concentration=0.
- For each 6-dimensional data ndarray (one for GF and one for CK, for each assay), I first computed the mean and standard deviation (again, using numpy.mean and numpy.std) of the 6 available values (for each GF plate or each CK half-plate) corresponding to ligand_name=CTRL, namely those for ligand_concentration [math]\displaystyle{ \in \{1, 100\} }[/math] and time [math]\displaystyle{ \in \{10, 30, 90\} }[/math]. Note that this mean is in fact a "mean of means of means", or more specifically, a "(6-well) mean of (well) means of (cell) means (of pixel intensities)". Likewise, the standard deviation is a "standard deviation of means of means".
- I added a "hyperslab" to the data original ndarray corresponding ligand_concentration=0, and set the value of all the cells in this hyperslab to the 6-well mean and standard deviation computed above;
- I repeated this procedure on the newly expanded ndarray, this time adding the new "hyperslab" at position corresponding to time=0;
- I stored these expanded ndarrays in the icbp45.h5 file, under the group name from_IR_w_zeros.
- Snapshot of current state of icbp45.h5.
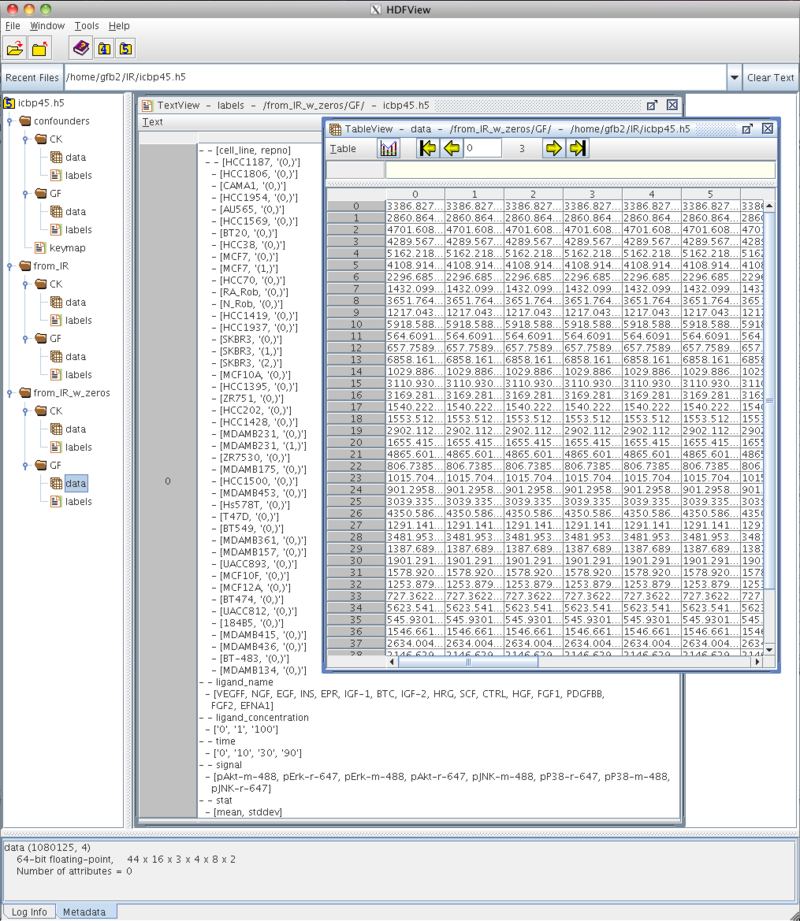
icbp45.h5 - Output of h5dump for current state of icbp45.h5:
HDF5 "icbp45.h5" {
GROUP "/" {
GROUP "confounders" {
GROUP "CK" {
DATASET "data" {
DATATYPE H5T_STD_U8LE
DATASPACE SIMPLE { ( 43, 8, 2, 3, 8, 5 ) / ( 43, 8, 2, 3, 8, 5 ) }
DATA {
(0,0,0,0,0,0): 1, 112, 3, 4, 113,
(0,0,0,0,1,0): 1, 112, 3, 6, 114,
...
(42,7,1,2,6,0): 163, 116, 102, 4, 119,
(42,7,1,2,7,0): 163, 116, 102, 6, 120
}
}
DATASET "labels" {
DATATYPE H5T_STRING {
STRSIZE 1352;
STRPAD H5T_STR_NULLPAD;
CSET H5T_CSET_ASCII;
CTYPE H5T_C_S1;
}
DATASPACE SCALAR
DATA {
(0): "- - [cell_line, repno]
- - [HCC1187, '(0,)']
- [HCC1806, '(0,)']
- [CAMA1, '(0,)']
- [HCC1954, '(0,)']
- [AU565, '(0,)']
- [HCC1569, '(0,)']
- [BT20, '(0,)']
- [HCC38, '(0,)']
- [MCF7, '(0,)']
- [MCF7, '(1,)']
- [HCC70, '(0,)']
- [RA_Rob, '(0,)']
- [N_Rob, '(0,)']
- [HCC1419, '(0,)']
- [HCC1937, '(0,)']
- [SKBR3, '(0,)']
- [SKBR3, '(1,)']
- [SKBR3, '(2,)']
- [MCF10A, '(0,)']
- [HCC1395, '(0,)']
- [ZR751, '(0,)']
- [HCC202, '(0,)']
- [HCC1428, '(0,)']
- [MDAMB231, '(0,)']
- [MDAMB231, '(1,)']
- [ZR7530, '(0,)']
- [MDAMB175, '(0,)']
- [HCC1500, '(0,)']
- [MDAMB453, '(0,)']
- [Hs578T, '(0,)']
- [T47D, '(0,)']
- [BT549, '(0,)']
- [MDAMB361, '(0,)']
- [MDAMB157, '(0,)']
- [UACC893, '(0,)']
- [MCF10F, '(0,)']
- [MCF12A, '(0,)']
- [BT474, '(0,)']
- [UACC812, '(0,)']
- [184B5, '(0,)']
- [MDAMB415, '(0,)']
- [BT-483, '(0,)']
- [MDAMB134, '(0,)']
- - ligand_name
- [LPS, IL-1\37777777716\37777777661, IL-6, CTRL, IFN-\37777777716\37777777661, IFN-\37777777716\37777777663, TNF-\37777777716\37777777661, IL-2]
- - ligand_concentration
- ['1', '100']
- - time
- ['10', '30', '90']
- - signal
- [NF-\37777777716\37777777672B-m-488, STAT1-r-647, pErk-m-488, STAT3-r-647, STAT1-r-488, NF-\37777777716\37777777672B-m-647,
STAT3-r-488, pErk-m-647]
- - confounder
- [assay, plate, well, channel, antibody]
"
}
}
}
GROUP "GF" {
DATASET "data" {
DATATYPE H5T_STD_U8LE
DATASPACE SIMPLE { ( 44, 16, 2, 3, 8, 5 ) / ( 44, 16, 2, 3, 8, 5 ) }
DATA {
(0,0,0,0,0,0): 1, 2, 3, 4, 5,
(0,0,0,0,1,0): 1, 2, 3, 6, 7,
...
(43,15,1,2,6,0): 163, 109, 102, 4, 110,
(43,15,1,2,7,0): 163, 109, 102, 6, 111
}
}
DATASET "labels" {
DATATYPE H5T_STRING {
STRSIZE 1412;
STRPAD H5T_STR_NULLPAD;
CSET H5T_CSET_ASCII;
CTYPE H5T_C_S1;
}
DATASPACE SCALAR
DATA {
(0): "- - [cell_line, repno]
- - [HCC1187, '(0,)']
- [HCC1806, '(0,)']
- [CAMA1, '(0,)']
- [HCC1954, '(0,)']
- [AU565, '(0,)']
- [HCC1569, '(0,)']
- [BT20, '(0,)']
- [HCC38, '(0,)']
- [MCF7, '(0,)']
- [MCF7, '(1,)']
- [HCC70, '(0,)']
- [RA_Rob, '(0,)']
- [N_Rob, '(0,)']
- [HCC1419, '(0,)']
- [HCC1937, '(0,)']
- [SKBR3, '(0,)']
- [SKBR3, '(1,)']
- [SKBR3, '(2,)']
- [MCF10A, '(0,)']
- [HCC1395, '(0,)']
- [ZR751, '(0,)']
- [HCC202, '(0,)']
- [HCC1428, '(0,)']
- [MDAMB231, '(0,)']
- [MDAMB231, '(1,)']
- [ZR7530, '(0,)']
- [MDAMB175, '(0,)']
- [HCC1500, '(0,)']
- [MDAMB453, '(0,)']
- [Hs578T, '(0,)']
- [T47D, '(0,)']
- [BT549, '(0,)']
- [MDAMB361, '(0,)']
- [MDAMB157, '(0,)']
- [UACC893, '(0,)']
- [MCF10F, '(0,)']
- [MCF12A, '(0,)']
- [BT474, '(0,)']
- [UACC812, '(0,)']
- [184B5, '(0,)']
- [MDAMB415, '(0,)']
- [MDAMB436, '(0,)']
- [BT-483, '(0,)']
- [MDAMB134, '(0,)']
- - ligand_name
- [VEGFF, NGF, EGF, INS, EPR, IGF-1, BTC, IGF-2, HRG, SCF, CTRL, HGF, FGF1, PDGFBB,
FGF2, EFNA1]
- - ligand_concentration
- ['1', '100']
- - time
- ['10', '30', '90']
- - signal
- [pAkt-m-488, pErk-r-647, pErk-m-488, pAkt-r-647, pJNK-m-488, pP38-r-647, pP38-m-488,
pJNK-r-647]
- - confounder
- [assay, plate, well, channel, antibody]
"
}
}
}
DATASET "keymap" {
DATATYPE H5T_STRING {
STRSIZE 2305;
STRPAD H5T_STR_NULLPAD;
CSET H5T_CSET_ASCII;
CTYPE H5T_C_S1;
}
DATASPACE SCALAR
DATA {
(0): "- {20100924_HCC1187: 1, 20100925_HCC1806: 121, 20100928_CAMA1: 122, 20101004_HCC1954: 123,
20101005_AU565: 124, 20101006_HCC1569: 125, 20101007_BT20: 126, 20101008_HCC38: 127,
20101012_MCF7: 128, 20101018_HCC70: 129, 20101021_RA_Rob: 130, 20101022_N_Rob: 131,
20101025_HCC1419: 132, 20101122_HCC1937: 133, 20101202_SKBR3: 134, 20101206_MCF10A: 135,
20101210_SKBR3: 136, 20101213_HCC1395: 137, 20101215_ZR751: 138, 20101216_HCC202: 139,
20101221_HCC1428: 140, 20101222_MDAMB231: 141, 20110128_ZR7530: 142, 20110201_SKBR3: 143,
20110210_MCF7: 144, 20110308_MDAMB175: 145, 20110310_HCC1500: 146, 20110311_MDAMB453: 147,
20110317_MDAMB231: 148, 20110318_Hs578T: 149, 20110321_T47D: 150, 20110322_BT549: 151,
20110324_MDAMB361: 152, 20110325_MDAMB157: 153, 20110330_UACC893: 154, 20110414_MCF10F: 155,
20110415_MCF12A: 156, 20110418_BT474: 157, 20110420_UACC812: 158, 20110421_184B5: 159,
20110421_MDAMB415: 160, 20110427_MDAMB436: 161, 20110502_BT-483: 162, 20110517_MDAMB134: 163}
- {CK1: 112, CK2: 116, GF1: 2, GF2: 103, GF3: 106, GF4: 109}
- {A01: 3, A02: 8, A03: 9, A04: 10, A05: 11, A06: 12, A07: 13, A08: 14, A09: 15, A10: 16,
A11: 17, A12: 18, B01: 19, B02: 20, B03: 21, B04: 22, B05: 23, B06: 24, B07: 25,
B08: 26, B09: 27, B10: 28, B11: 29, B12: 30, C01: 31, C02: 32, C03: 33, C04: 34,
C05: 35, C06: 36, C07: 37, C08: 38, C09: 39, C10: 40, C11: 41, C12: 42, D01: 43,
D02: 44, D03: 45, D04: 46, D05: 47, D06: 48, D07: 49, D08: 50, D09: 51, D10: 52,
D11: 53, D12: 54, E01: 55, E02: 56, E03: 57, E04: 58, E05: 59, E06: 60, E07: 61,
E08: 62, E09: 63, E10: 64, E11: 65, E12: 66, F01: 67, F02: 68, F03: 69, F04: 70,
F05: 71, F06: 72, F07: 73, F08: 74, F09: 75, F10: 76, F11: 77, F12: 78, G01: 79,
G02: 80, G03: 81, G04: 82, G05: 83, G06: 84, G07: 85, G08: 86, G09: 87, G10: 88,
G11: 89, G12: 90, H01: 91, H02: 92, H03: 93, H04: 94, H05: 95, H06: 96, H07: 97,
H08: 98, H09: 99, H10: 100, H11: 101, H12: 102}
- {'530': 4, '685': 6}
- {NF-\37777777716\37777777672B|mouse|488: 113, NF-\37777777716\37777777672B|mouse|647: 118, STAT1|rabbit|488: 117, STAT1|rabbit|647: 114,
STAT3|rabbit|488: 119, STAT3|rabbit|647: 115, pAkt|mouse|488: 5, pAkt|rabbit|647: 105,
pErk|mouse|488: 104, pErk|mouse|647: 120, pErk|rabbit|647: 7, pJNK|mouse|488: 107,
pJNK|rabbit|647: 111, pP38|mouse|488: 110, pP38|rabbit|647: 108}
"
}
}
}
GROUP "from_IR" {
GROUP "CK" {
DATASET "data" {
DATATYPE H5T_IEEE_F64LE
DATASPACE SIMPLE { ( 43, 8, 2, 3, 8, 2 ) / ( 43, 8, 2, 3, 8, 2 ) }
DATA {
(0,0,0,0,0,0): 1.64638, 1.31093,
(0,0,0,0,1,0): 174.688, 84.2019,
...
(42,7,1,2,6,0): 734.847, 327.546,
(42,7,1,2,7,0): 1034.71, 710.928
}
}
DATASET "labels" {
DATATYPE H5T_STRING {
STRSIZE 1321;
STRPAD H5T_STR_NULLPAD;
CSET H5T_CSET_ASCII;
CTYPE H5T_C_S1;
}
DATASPACE SCALAR
DATA {
(0): "- - [cell_line, repno]
- - [HCC1187, '(0,)']
- [HCC1806, '(0,)']
- [CAMA1, '(0,)']
- [HCC1954, '(0,)']
- [AU565, '(0,)']
- [HCC1569, '(0,)']
- [BT20, '(0,)']
- [HCC38, '(0,)']
- [MCF7, '(0,)']
- [MCF7, '(1,)']
- [HCC70, '(0,)']
- [RA_Rob, '(0,)']
- [N_Rob, '(0,)']
- [HCC1419, '(0,)']
- [HCC1937, '(0,)']
- [SKBR3, '(0,)']
- [SKBR3, '(1,)']
- [SKBR3, '(2,)']
- [MCF10A, '(0,)']
- [HCC1395, '(0,)']
- [ZR751, '(0,)']
- [HCC202, '(0,)']
- [HCC1428, '(0,)']
- [MDAMB231, '(0,)']
- [MDAMB231, '(1,)']
- [ZR7530, '(0,)']
- [MDAMB175, '(0,)']
- [HCC1500, '(0,)']
- [MDAMB453, '(0,)']
- [Hs578T, '(0,)']
- [T47D, '(0,)']
- [BT549, '(0,)']
- [MDAMB361, '(0,)']
- [MDAMB157, '(0,)']
- [UACC893, '(0,)']
- [MCF10F, '(0,)']
- [MCF12A, '(0,)']
- [BT474, '(0,)']
- [UACC812, '(0,)']
- [184B5, '(0,)']
- [MDAMB415, '(0,)']
- [BT-483, '(0,)']
- [MDAMB134, '(0,)']
- - ligand_name
- [LPS, IL-1\37777777716\37777777661, IL-6, CTRL, IFN-\37777777716\37777777661, IFN-\37777777716\37777777663, TNF-\37777777716\37777777661, IL-2]
- - ligand_concentration
- ['1', '100']
- - time
- ['10', '30', '90']
- - signal
- [NF-\37777777716\37777777672B-m-488, STAT1-r-647, pErk-m-488, STAT3-r-647, STAT1-r-488, NF-\37777777716\37777777672B-m-647,
STAT3-r-488, pErk-m-647]
- - stat
- [mean, stddev]
"
}
}
}
GROUP "GF" {
DATASET "data" {
DATATYPE H5T_IEEE_F64LE
DATASPACE SIMPLE { ( 44, 16, 2, 3, 8, 2 ) / ( 44, 16, 2, 3, 8, 2 ) }
DATA {
(0,0,0,0,0,0): 3327.29, 1702.67,
(0,0,0,0,1,0): 231.557, 224.42,
...
(43,15,1,2,6,0): 2001.45, 693.117,
(43,15,1,2,7,0): 167.17, 59.7259
}
}
DATASET "labels" {
DATATYPE H5T_STRING {
STRSIZE 1381;
STRPAD H5T_STR_NULLPAD;
CSET H5T_CSET_ASCII;
CTYPE H5T_C_S1;
}
DATASPACE SCALAR
DATA {
(0): "- - [cell_line, repno]
- - [HCC1187, '(0,)']
- [HCC1806, '(0,)']
- [CAMA1, '(0,)']
- [HCC1954, '(0,)']
- [AU565, '(0,)']
- [HCC1569, '(0,)']
- [BT20, '(0,)']
- [HCC38, '(0,)']
- [MCF7, '(0,)']
- [MCF7, '(1,)']
- [HCC70, '(0,)']
- [RA_Rob, '(0,)']
- [N_Rob, '(0,)']
- [HCC1419, '(0,)']
- [HCC1937, '(0,)']
- [SKBR3, '(0,)']
- [SKBR3, '(1,)']
- [SKBR3, '(2,)']
- [MCF10A, '(0,)']
- [HCC1395, '(0,)']
- [ZR751, '(0,)']
- [HCC202, '(0,)']
- [HCC1428, '(0,)']
- [MDAMB231, '(0,)']
- [MDAMB231, '(1,)']
- [ZR7530, '(0,)']
- [MDAMB175, '(0,)']
- [HCC1500, '(0,)']
- [MDAMB453, '(0,)']
- [Hs578T, '(0,)']
- [T47D, '(0,)']
- [BT549, '(0,)']
- [MDAMB361, '(0,)']
- [MDAMB157, '(0,)']
- [UACC893, '(0,)']
- [MCF10F, '(0,)']
- [MCF12A, '(0,)']
- [BT474, '(0,)']
- [UACC812, '(0,)']
- [184B5, '(0,)']
- [MDAMB415, '(0,)']
- [MDAMB436, '(0,)']
- [BT-483, '(0,)']
- [MDAMB134, '(0,)']
- - ligand_name
- [VEGFF, NGF, EGF, INS, EPR, IGF-1, BTC, IGF-2, HRG, SCF, CTRL, HGF, FGF1, PDGFBB,
FGF2, EFNA1]
- - ligand_concentration
- ['1', '100']
- - time
- ['10', '30', '90']
- - signal
- [pAkt-m-488, pErk-r-647, pErk-m-488, pAkt-r-647, pJNK-m-488, pP38-r-647, pP38-m-488,
pJNK-r-647]
- - stat
- [mean, stddev]
"
}
}
}
}
GROUP "from_IR_w_zeros" {
GROUP "CK" {
DATASET "data" {
DATATYPE H5T_IEEE_F64LE
DATASPACE SIMPLE { ( 43, 8, 3, 4, 8, 2 ) / ( 43, 8, 3, 4, 8, 2 ) }
DATA {
(0,0,0,0,0,0): 2.2956, 0.103067,
(0,0,0,0,1,0): 223.036, 33.296,
...
(42,7,2,3,6,0): 734.847, 327.546,
(42,7,2,3,7,0): 1034.71, 710.928
}
}
DATASET "labels" {
DATATYPE H5T_STRING {
STRSIZE 1331;
STRPAD H5T_STR_NULLPAD;
CSET H5T_CSET_ASCII;
CTYPE H5T_C_S1;
}
DATASPACE SCALAR
DATA {
(0): "- - [cell_line, repno]
- - [HCC1187, '(0,)']
- [HCC1806, '(0,)']
- [CAMA1, '(0,)']
- [HCC1954, '(0,)']
- [AU565, '(0,)']
- [HCC1569, '(0,)']
- [BT20, '(0,)']
- [HCC38, '(0,)']
- [MCF7, '(0,)']
- [MCF7, '(1,)']
- [HCC70, '(0,)']
- [RA_Rob, '(0,)']
- [N_Rob, '(0,)']
- [HCC1419, '(0,)']
- [HCC1937, '(0,)']
- [SKBR3, '(0,)']
- [SKBR3, '(1,)']
- [SKBR3, '(2,)']
- [MCF10A, '(0,)']
- [HCC1395, '(0,)']
- [ZR751, '(0,)']
- [HCC202, '(0,)']
- [HCC1428, '(0,)']
- [MDAMB231, '(0,)']
- [MDAMB231, '(1,)']
- [ZR7530, '(0,)']
- [MDAMB175, '(0,)']
- [HCC1500, '(0,)']
- [MDAMB453, '(0,)']
- [Hs578T, '(0,)']
- [T47D, '(0,)']
- [BT549, '(0,)']
- [MDAMB361, '(0,)']
- [MDAMB157, '(0,)']
- [UACC893, '(0,)']
- [MCF10F, '(0,)']
- [MCF12A, '(0,)']
- [BT474, '(0,)']
- [UACC812, '(0,)']
- [184B5, '(0,)']
- [MDAMB415, '(0,)']
- [BT-483, '(0,)']
- [MDAMB134, '(0,)']
- - ligand_name
- [LPS, IL-1\37777777716\37777777661, IL-6, CTRL, IFN-\37777777716\37777777661, IFN-\37777777716\37777777663, TNF-\37777777716\37777777661, IL-2]
- - ligand_concentration
- ['0', '1', '100']
- - time
- ['0', '10', '30', '90']
- - signal
- [NF-\37777777716\37777777672B-m-488, STAT1-r-647, pErk-m-488, STAT3-r-647, STAT1-r-488, NF-\37777777716\37777777672B-m-647,
STAT3-r-488, pErk-m-647]
- - stat
- [mean, stddev]
"
}
}
}
GROUP "GF" {
DATASET "data" {
DATATYPE H5T_IEEE_F64LE
DATASPACE SIMPLE { ( 44, 16, 3, 4, 8, 2 ) / ( 44, 16, 3, 4, 8, 2 ) }
DATA {
(0,0,0,0,0,0): 3386.83, 139.386,
(0,0,0,0,1,0): 237.524, 11.408,
...
(43,15,2,3,6,0): 2001.45, 693.117,
(43,15,2,3,7,0): 167.17, 59.7259
}
}
DATASET "labels" {
DATATYPE H5T_STRING {
STRSIZE 1391;
STRPAD H5T_STR_NULLPAD;
CSET H5T_CSET_ASCII;
CTYPE H5T_C_S1;
}
DATASPACE SCALAR
DATA {
(0): "- - [cell_line, repno]
- - [HCC1187, '(0,)']
- [HCC1806, '(0,)']
- [CAMA1, '(0,)']
- [HCC1954, '(0,)']
- [AU565, '(0,)']
- [HCC1569, '(0,)']
- [BT20, '(0,)']
- [HCC38, '(0,)']
- [MCF7, '(0,)']
- [MCF7, '(1,)']
- [HCC70, '(0,)']
- [RA_Rob, '(0,)']
- [N_Rob, '(0,)']
- [HCC1419, '(0,)']
- [HCC1937, '(0,)']
- [SKBR3, '(0,)']
- [SKBR3, '(1,)']
- [SKBR3, '(2,)']
- [MCF10A, '(0,)']
- [HCC1395, '(0,)']
- [ZR751, '(0,)']
- [HCC202, '(0,)']
- [HCC1428, '(0,)']
- [MDAMB231, '(0,)']
- [MDAMB231, '(1,)']
- [ZR7530, '(0,)']
- [MDAMB175, '(0,)']
- [HCC1500, '(0,)']
- [MDAMB453, '(0,)']
- [Hs578T, '(0,)']
- [T47D, '(0,)']
- [BT549, '(0,)']
- [MDAMB361, '(0,)']
- [MDAMB157, '(0,)']
- [UACC893, '(0,)']
- [MCF10F, '(0,)']
- [MCF12A, '(0,)']
- [BT474, '(0,)']
- [UACC812, '(0,)']
- [184B5, '(0,)']
- [MDAMB415, '(0,)']
- [MDAMB436, '(0,)']
- [BT-483, '(0,)']
- [MDAMB134, '(0,)']
- - ligand_name
- [VEGFF, NGF, EGF, INS, EPR, IGF-1, BTC, IGF-2, HRG, SCF, CTRL, HGF, FGF1, PDGFBB,
FGF2, EFNA1]
- - ligand_concentration
- ['0', '1', '100']
- - time
- ['0', '10', '30', '90']
- - signal
- [pAkt-m-488, pErk-r-647, pErk-m-488, pAkt-r-647, pJNK-m-488, pP38-r-647, pP38-m-488,
pJNK-r-647]
- - stat
- [mean, stddev]
"
}
}
}
}
}
}