User:Erik Maradiaga/Notebook/Biology 210 at AU
Lab Entry #7 3/3/15: 16S Sequence Analysis
Purpose: The purpose of this experiment was to use the 16S DNA Sequence from two of our bacterial colonies to be able to classify them. Before we set up the PCR in order to amplify this DNA sequence, my lab partner and I gram stained four of our bacterial colonies in order to see if they were gram positive or gram negative. As a result, we were able to classify that 3 of our bacterial colonies were gram positive and 1 was gram negative. This is significant to note because our white colony with tetracycline was gram positive, and our yellow colony without tetracycline was gram negative and these were the two bacteria that was used to amplify the DNA sequence. It can be concluded then that the white colony that was gram positive is Bacillus cereus strain, and the yellow colony that was gram negative is Pseudomonas sp.
Material and Methods: Before we began identifying the bacteria by using the DNA sequence, my partner and I used Gel electrophoresis in order to identify which colony to use for the 16S Sequence for to amplify the gene.Then, by using the MB code that was labeled for our tube, we were able to go to a website call Gene Wiz in order to view the DNA sequence corresponding to the MB tube. Afterward, we were able to copy the DNA sequence for each of our colonies and paste them on a different website called Blast to see if we have a match for a bacteria. Afterward, this information was used to compared results from the Gram Stain procedure and see if the bacteria that was identified correspond to the other results.
Data and Observation: White with tetra colony #3
NNNNNNNNNNNNNNCNANANTGCAGTCGAGCGAATGGATTAAGAGCTTGCT CTTATGAAGTTAGCGGCGGACGGGTGAGT AACACGTGGGTAACCTGCCCATAAGACTGGGATAACTCCGGGAAACCGGGGCTAATACCGGATAACATTTTGAACTGCAT GGTTCGAAATTGAAAGGCGGCTTCGGCTGTCACTTATGGATGGACCCGCGTCGCATTAGCTAGTTGGTGAGGTAACGGCT CACCAAGGCAACGATGCGTAGCCGACCTGAGAGGGTGATCGGCCACACTGGGACTGAGACACGGCCCAGACTCCTACGGG AGGCAGCAGTAGGGAATCTTCCGCAATGGACGAAAGTCTGACGGAGCAACGCCGCGTGAGTGATGAAGGCTTTCGGGTCG TAAAACTCTGTTGTTAGGGAAGAACAAGTGCTAGTTGAATAAGCTGGCACCTTGACGGTACCTAACCAGAAAGCCACGGC TAACTACGTGCCAGCAGCCGCGGTAATACGTAGGTGGCAAGCGTTATCCGGAATTATTGGGCGTAAAGCGCGCGCAGGTG GTTTCTTAAGTCTGATGTGAAAGCCCACGGCTCANCCGTGGAGGGTCATTGGAAACTGGGAGACTTGANTGCAGAAGAGG AAAGTGGAATTCCATGTGTAGCGGTGAAATGCGTAGAGATATGGAGGAACACCAGTGGCGAANGCGACTTTCTGGTCTGT AACTGACACTGAGGCGCGAAAGCGTGGGGAGCAAACAGGATTAGATACCCTGGTAGTCCACGCCGTAAACNATGAGTGCT AANTGTTAGAGGGTTTCCGCCCTTTANTGCTGAAGTTAACGCATTAAGCACTCCGCCTGGGGAGTACGGCCGCAAGGCTG AAACTCAAGGAATTGACNGGGGCCCGCACAAGCNGTGNANCATGTGGTTTAATTCNAANCAACGCGANNACCTTACCAGG TCNTGACNTCCTCTGANACCCTANAGATNNGGCTTNNNCNTCGGGANCAGAGTGACAGTGNNCATGNTGTCNTCANCTCN NGTCNTGAGNNGTTGGTNANNNCNNCAACNAGCNGCAACCCNNNATCTTNNTTGCCNNNNTNANNTNN
Yellow without tetra colony #1
NNNNNNNNNNNNNNNNNNNNNTANNNTNCNNNCGGGGNNCGATAGNANCNTGCTNAAGATTCNNGANNGACGGGTGAGTA
ATGCCAAGGAATCTACCTAGGGGTGGGGGACACCTTTTCGAACGGAAACCTCATACCGCATACATCGTACNGGANAGGGC
AGGGGACCCTCCGGCCCTGCCCTAATAAATGAGCCCAGGTCGGATTAGCTGGTTGGTGAGGTAATGGCTCACCAAGGCNA
CGATCCGTAGCTGGTCTGAAANGATGATCNGTCACCCTGGAACTGAGACACGGTCCNNACTCCTACGGGAGGCNGANTGG
GGAATATGGGAGAATGGGTGATCGCNTNNNCCANCCCGCCCGCGTGTGTGANNAAGGTCTTCNGATCGTTTTGACNTTNA
NANANGATNNAACGCGTGAAATTCNGANCGTACTGTTTTAAAATACACCNNNNAACACCGAGCCCTNNNCCNNGTNCCAC
CANNNGGTNGTAACNTTNTNCGGATTTCTTTTGTTCANAGTTTCTGGACGTGGAGCNGNATANNTGNNGAAATATCTTAN
TNTNANATCACCCTGCCANNNATANNGNCTGTCTTGANAAATTACAAGCGNCAGGATGAAGTAGTGTANCGGAGATCCGC
NTAGATANTACTGANAACANNNATTGCGAAGGGAAGTCGCTATGTCNTAACGGGACCCCCGATAGCACAAN
Image of DNA sequence for Yellow Colony without Tetracycline
Image of DNA sequence for white colony with Tetracycline
Image of band for each colony from Gel Electrophoresis
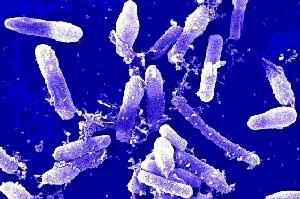
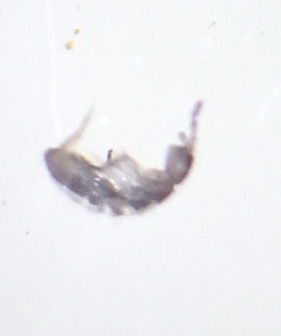
Image of Bacillus cereus that corresponds to white colony with tetracycline
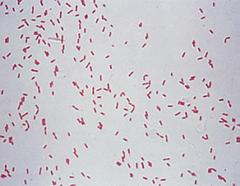
Image of Pseudomonas sp. that corresponds to the yellow colony without tetracycline
The sequence of each colony allowed us to identify the unknown bacteria in our transect. For example, the white colony that grew under tetracycline is Bacillus cereus because it matches the result from our previous experiments. For instance, Bacillus cereus is a gram-positive bacteria that has a shape of rods. This matches with our description from the gram- stain procedure because my lab partner and I said that it was purple under entry #3. Also, we found that the unknown bacteria under entry #3 had a bacillus shape, and this matches with the results that is found associated with Bacillus cereus. Under the image showing Bacillus cereus, it can be seen that it had a bacillus shape, which matches with our results. However, we could not tell if the bacteria was moving under the microscope so this interferes with our results. Likewise, the Pseudomonas that corresponds to the yellow colony without tetracycline matches with our results. For example, Pseudomonas is a gram-negative bacteria and this matches with our results that we found from the microbiology lab. The yellow colony without tetracycline was also gram-negative, so this is matches the result from the DNA sequence. Similarly, the bacteria that was viewed under the microscope from lab 3 had a rod shaped and the Pseudomonas also has a rod-shaped. Likewise, we were not able compare the motility because we were not able to view the motility under the microscope since we saw no movement.
Conclusion and Future Directions: It can be concluded that the white colony that was gram positive is Bacillus cereus and the yellow bacteria that was gram negative was Pseudomonas. This can be certain because the 16S DNA sequence matches each bacteria with the results from the microbiology lab. For future directions, I would do the same procedure but for a different colony to see if the 16S DNA sequence matches the description from the gram-stain procedure.
- Erik Maradiaga 16:57, 3 March 2015 (EST):Erik Maradiaga
Lab Entry #6 2/25/15 - 3/4/15: Updated Zebrafish Experiment
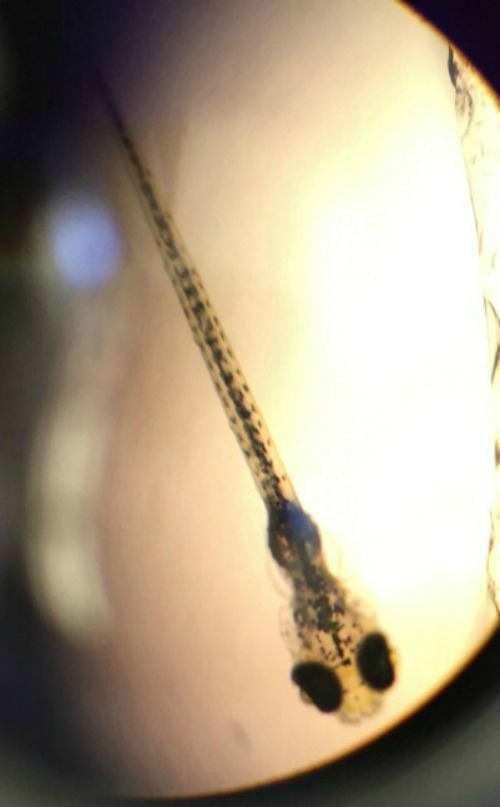
Purpose: The purpose of this experiment is to see the effects of alcohol in the development of zebrafish. Zebrafish are a great microorganism to study embryology because of many reasons. One reason is that they are transparent to see. As a result, it makes it easier to see at what stage of development they are in and record observations. Also, zebrafish can grow in a matter of days. This is significant to note because my lab partner and I only have 2 weeks to see results from our experiment. Last, zebrafish are really sensitive to their surrounding. This, of course, will make it easier to note the change that alcohol has on the development of zebrafish. Our hypothesis for this experiment is that alcohol will have an effect on the development of zebrafish compare to our control group which only contains water. Alcohol may have an effect on the neurons that control movement in the zebrafish. Likewise, because alcohol is used to disinfect, we predict that the zebrafish that are immersed under alcohol will have cleaner plates compared to the control group, which should be dirtier with fungi and bacteria.
Materials and Method: In order to begin making observations of the zebrafish, we started by preparing 3 petri dish. For the first petri dish, this was going to be our control group. We added 20 mLs of distilled water. For the second petri dish, we diluted 1.5% of alcohol into .75% of alcohol by adding 10 mLs of water and adding 10mLs of 1.5% alcohol. For the last petri dish, we added 20 mLs of 1.5% alcohol. Afterward, we began adding 20 healthy translucent zebrafish into each of our petri dish. It is important to note that we added zebrafish that arrived on 2/19/15 which means that they were less than 24 hours old. We let this sit in the lab overnight. On Monday 2/23/15, we removed the empty egg cases and any mold that we saw. Likewise, we recorded our observations and noted how much were alive or dead and the stage of development they are in. Also, we added a single drop of paramecium into each of our petri dish in order to feed the zebrafish. This will replenish the zebrafish in each of our petri dish. Last, we removed 8 mL of water and added 10 mL of freshwater back to each of our petri dish. Similarly, we added 10 mL of alcohol back to each of our petri dish to replenish it. This step was repeated for the last 2 days of observation. On the last 2 days of observations, we were testing the movement of the zebrafish when we flicked the plates. We flicked the plates and recorded the reaction that the zebrafish made when they responded to this. Also, a pipette test was used to evaluate the movement as well. Pipette was used to touch the zebrafish and record the reaction that each of the zebrafish made under this response.
Data and Observation:
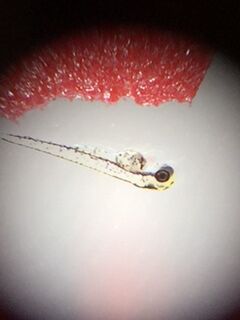
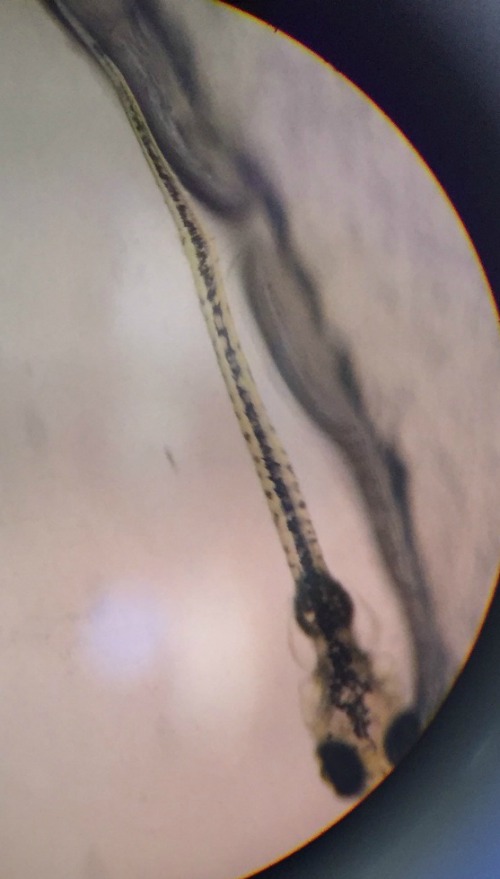
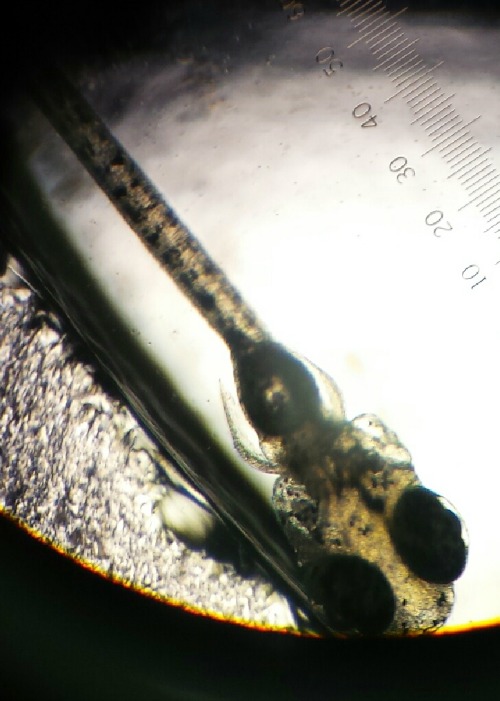
Image of Zebrafish in Pric-fin stage from 1.5% alcohol group on Feb 23.
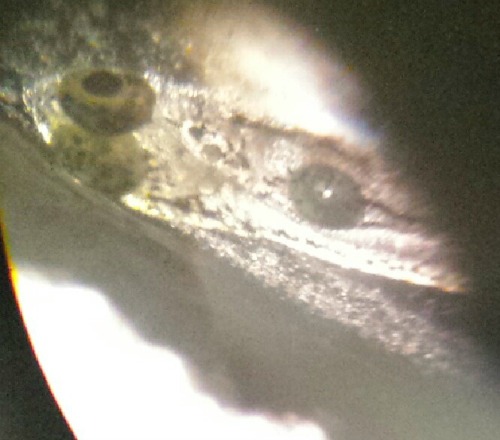
Image of Zebrafish in Prim-6 stage from control group on Feb 23.
Image of Zebrafish from Prim-6 stage from the .75% alcohol group on Feb 23.
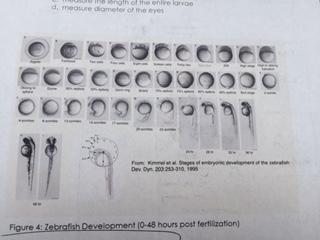
Image of all the possible developmental stages in a zebrafish.
Image of Zebrafish on March 2. Fully grown with long tail
Image of Zebrafish on March 2. Longer and has darker eyes and heart compared to the first day when we observed
Image of Zebrafish on March 2. Fully grown compared to the one from Feb 23.
Image of the control group from observations made on March 4. Notice how fully grown and darker the zebrafish is compared to the other days
Image of control group from observations made on March 4. It has darker eyes and it is fully grown. Hard to tell in the picture
Image of 1.5% Alcohol group taken on March 4. Notice the large eyes and the heart.
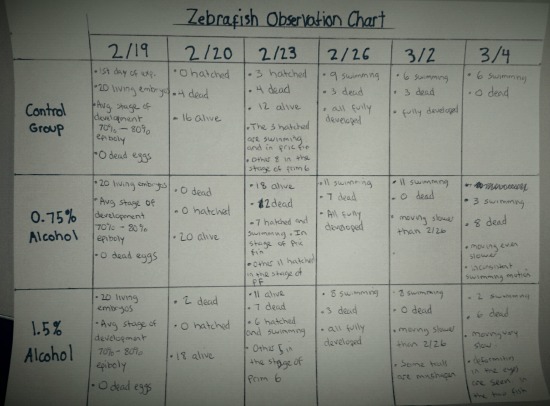
Image of Observation chart for testing of movement and Pipette test that Lab partner and I made
Image of movement chart. Was testing the effect of alcohol on movement by tap on plates and pipette test
In the first diagram it is a picture of a zebrafish that is in pric-fin stage. This diagram was taken from the petri dish that contains 1.5 % alcohol. The Pric-fin stage means that it is long and ready to swim. It also means that it has a heartbeat. You can visually the heart in the diagram by looking behind the dark eyes. Likewise, the second diagram is a picture of a zebrafish in prim-6 stage from the control group. Prim- 6 means that there are black cells growing on the side of the egg. From diagram 2, it can be seen that the eyes are growing and the zebrafish's tail on the side. The last diagram is showing the zebrafish are in prim-6 stage from the .75% alcohol. This means that this petri dish is similar to the one from the control group because most are in this stage and only a few are swimming. The zebrafish from the .75% alcohol have the most zebrafish that are swimming. In the table above, it is showing the recorded observation from Friday and Monday with specific dates stated in the material and method section.We are identifying the stages of the zebrafish by using the last table. This table describes all the stages of the embryonic development of a zebrafish, and this is being used to describe the stages of our zebrafish in our experiment. The diagram above show that there is eye movement and fin development. However in the diagram above, the mouth cannot be seen which means that it may have not yet developed. The yolk size that my lab partner and I determined is said to be 2.5 nm. All of these observations were made in the first 2 days when we were just observing how much were alive and dead and what stage they were in. The following image are zebrafish that were observe on the following days.
Conclusion and Future Directions: This data entry only includes the observations taken from Friday and Monday. We still have two more days to note results, so this experiment is still not done. So far, my lab partner and I can conclude that the zebrafish from .75% alcohol are hatching and swimming quicker compared to both of the extremes. My prediction is that the zebrafish from 1.5% alcohol and .75% will die quicker compare to the control group because of the effect of alcohol. This can be determined by taking notes in the following next 2 days of observations. After we finished making all the observations, my lab partner and i concluded that our hypothesis was indeed correct. Alcohol did had an effect on the movement of the zebrafish and how clean or dirty the plates were. Whenever my lab partner and I tapped on the plates to record the reaction of the zebrafish, we saw that the control group immediately responded and swam. However, the groups that were underneath alcohol barely responded and some only moved. This shows that alcohol may have an effect on the neurons on zebrafish because it took them a longer time to respond to this stimuli. Likewise, the pipette test had similar results so this supported our hypothesis. Also, we noticed that the control group have dirtier plates compared to the ones with alcohol. The control group had fungi on some areas along with some dirt, unlike the alcohol groups which were almost completely clean. This shows that the alcohol had an effect in destroying the bacteria and fungi, hence these groups were cleaner.
- Erik Maradiaga 11:20, 19 March 2015 (EDT):Erik Maradiaga
2.24.15 Very good entry. Image of data table identifying invertebrates isn't clear. Detailed description of Vertebrates and nice food web. SK
Lab Entry #5 2/18/15: Invertebrates and Vertebrates
Purpose: The purpose of this experiment was to identify certain invertebrates in our Berlese Funnel that we set up last week. Also, we identified certain vertebrates that might cross our transect at any given moment and from this information, we were able to construct a food web showing how each organisms is consumed in a given community. By the end of this experiment, we were able to identified these organisms by using a key and able to understand which phylum and class they belong to. Likewise, we were able to identified at least five vertebrates that might intersect at our transect and classify them from their phylum all the way to the species level. Our hypothesis is that we were going to find at least two invertebrates in our Berlese Funnel, but we were able to identified at least three.
Materials and Method: Before we began searching for invertebrates using a key, my lab partners and I had to set up a Berlese Funnel the week before. We poured 25 mL of 50:50 ethanol/water solution into a 50 mL conical tube. From there, we grabbed a funnel and tape screening material at the bottom of the funnel in order so that the leaf litter do not fall into the preservative. We carefully put the leaf litter sample on top of the funnel and placed it below a 40 watt lamp. We also added soil on top of our leaf litter in case there was invertebrates present in the soil. We let it sit the lab under the lamp for one week. Later in the experiment, we used the insect identification key in order to identified our invertebrates in our Berlese Funnel. We also used the microscope and dissecting microscope in order to have practice identifying which phylum and class our invertebrates belong to.
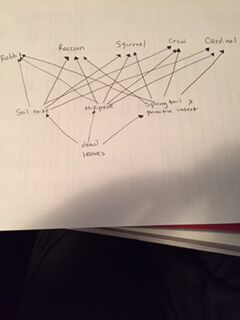
The pictures above show the invertebrates that we discover under the microscope and the food web diagram including the vertebrates. From the microscope, we were able to discover the Soil Mite. This invertebrates belongs to the phylum Arthropoda and in the class Arachida. We saw two samples underneath the microscope and they were very small. They were red and symmetrical. Another organism that we saw was the millipede. this organism belongs to the phylum Arthopoda and in the class Diplopoda. This organism had many longs, no antenna, and was very long. We only saw 1 sample underneath the microscope. The last organism we saw was the Springtail X Primitive Insect. This invertebrate belongs to the phylum Arthropoda and in the class Eutognatha. Under the microscope, it was purple/grayish, had 3 legs, and had an antenna. These were the invertebrates that we saw underneath the microscope. The vertebrates that we believe may come in contact with our transect are the following:
Rabbit: Phylum: Chordota, Class: Mammalia, Order: Lagomorpha, Family: Leporidae, Genus: Oryctolagus, Species: Oryctolagus cuniculus
Squirrel: Phylum: Chordota, Class: Mammalia, Order: Rodentia, Family: Sciuridae, Genus: Sciurus, Species: Sciurus carolinensis
Raccon: Phylum: Chordota, Class: Mammalia, Order: Carnivora, Family: Procyonidae, Genus: Procyon, Species: Lotor
Cardinal: Phylum: Chordota, Class: Aves, Order: Passeriformes, Family: Cardinalidae, Genus: Cardinalis, Species: C. cardinalis
Crow: Phylum: Chordota, Class: Bird, Order: Passeriformes, Family: Corridae, Genus: Corvus, Species: Corvus brachyrhynchos
Above are the following vertebrates that we believe may inhabit our transect. Soil and leaves would benefit these species because this is where the invertebrates may grow in. As a result, the vertebrates may prey on these invertebrates for their source of energy. The leaves and grass would represent the first trophic level because they will be consumed by the primary consumers which in this case would be the invertebrates. The secondary consumers would be these vertebrates because they are able to prey on the primary consumers for their source of energy. These organisms represent a community because they interact with each other and their abiotic and biotic factors. However, there comes a limit where they are not able to retrieve more energy or any source and this is refer to as the carrying capacity.
Conclusion and Future Directions: Overall, this experiment was a success because we were able to identified at least three invertebrates from our Brelese Funnels. We were able to identified the Soil Mite, Millipede, and the Springtail X Primitive Insect. Likewise, the vertebrates were also able to be identified and this allowed me to construct the food web showing how the source of energy travel in a community. For future directions, I would grabbed a larger funnel. This would allowed more leaves litter to be inserted in the funnel in order for the invertebrates to grow. As a result, this may bring more results to our table and be able to identified more invertebrates.
Resources:
http://a-z-animals.com/animals/rabbit/ http://bioweb.uwlax.edu/bio203/s2012/toellner_kayl/classification.htm http://www.fcps.edu/islandcreekes/ecology/common_crow.htm
- Erik Maradiaga 11:09, 19 February 2015 (EST):Erik Maradiaga
2.19.15 Excellent. Included all relevant data and discussed findings and procedures clearly. SK
Lab Entry #4 2/6/15: Plantae and Fungi
Purpose: The purpose of this experiment was to go back to our transect and identify the structures of the plants that we have. In our transect, there are a variety of different plants that are located throughout and we were interested in identifying what type of plant it is and the structure that it composed. The reason why we are doing this experiment is to not only to be able to identify the protists that we find in the soil, but also there are invertebrates that are living in the leaf litter. This would allow us to identify the protists and the invertebrates for next week lab. There is no hypothesis in this this experiment because we were just identifying the structures of five different plants from our transect using a microscope.
Materials and Method: For this experiment, we visited our transect that is located in front of Kogod and our objective was to collect leaves litter and 5 different plants. When we got to our transect, we decided to pick up moss that was located in front of our transect for one of our plants. Next, we noticed that there was a red flower in the far back to the left and we knew that this plant may be interesting to study. We also got grass from the front of the transect. Four our fourth plant, we got a cattail that was in the front to the right. Last, we got this flower that we were uncertain of but it had 3 branches, and it was located to the right of our transect. Once we collected our five different plants, we grabbed leaves litter from our transect and placed both the flowers and leaves into two different zip lock bags. In the lab, we used the dissecting microscope to look at the structures of our plants and different types of fungi. Last, we used Gel Electrophoresis to identify which sample of bacteria was more efficient in having the 16S DNA sequence.
Above are the pictures of our 5 different plants, the table on page 41 that describes the different plants, and a picture of the fungi we looked at under the dissecting microscope. The image of the grass was located in front of the transect. It had a xylem and phloem and was very soft.The red flower that we found was in the far left back of the transect and it was very rough to break. It was thick and cubicle were present. It also had xylem and phloem. Next, the moss was found in front of the transect and it was very tiny and circular. Its vascularization was Rhizoids and it did not have specialized structures. The cattail was found in the front of the transect to the right. It was very large and cylinder, and xylem and phloem were also present. it was rough and seeds would travel in the air when we grabbed a sample. Last, the flower with 3 branches was located in the front of the transect to the right. it was very rough and had 3 branches. Xylem and Phloem were present and it had cubicle as well. This means that the flower was also rough to deal with. The leaves of our transect was approximately 4 inches and very clustered throughout. The size vary in our transect as there were many large and small leaves. As for the image of the fungi, we studied the components of it. The fungi Sporangia are black spots that it contains and it also have spores. The image indeed is a fungi and they belong to the Ascomycota group.
Conclusion and Future Directions: For our experiment, we were able to identify the structures of our five different plants from the transect. We were also able to identify the method of reproduction that the plants undergoes and described its specialized structures. For future directions, I would grab a different plant from different areas of the transect in order to study. This would allow our data and future lab to have a variety of invertebrates that we will have to find in future labs. Overall, we were able to identity the protists that are in our transect as well as some of the plants and its mechanism.
- Erik Maradiaga 12:35, 6 February 2015 (EST):Erik Maradiaga
2.10.15 There is a bit of confusion here as you refer to protists in some parts of the notebook entry rather than bacteria. This whole lab was based on looking at bacteria that was cultured on agar plates from your transect, not protists. The methods were thorough. SK
Lab Entry #3 2/5/15: Microbiology and Identifying Protist' structure under Microscope
Purpose: The purpose of this experiment was to study the protists that we cultured from last week's experiment. We are going to study the motility and structure of the protists that we identified by using the gram stain method. The reason why my lab partners and I wanted did this experiment was to familiarize ourselves how each protists looked under the microscope very closely by using 100x with oil and whether our protists are gram-positive or gram-negative creatures. A hypothesis as to why the appearance or smell might change week to week in our hay infusion is that bacteria still may be growing in those niches. As a result, the appearance may change because bacteria are growing and this may affect the smell as well.
Materials and Method: Before we began this lab, my lab partner and I had to dilute and prepared the Agar plates in order to grow the bacteria. We took 100 mL of broth and inserted it into 6 different agar plates that either contain tetracycline or not. After we prepared the Agar plates so that the colonies of bacteria can grow, we let it grow for about 7 days. In the lab, we took a scrape of colonies from the agar plate that contain tetracycline and the one that did not in order to proceed with the Gram Stain procedure. We took colonies and place it into slides in order to dry the bacterial by using the Bunsen Burner. After this, we covered the bacterial from each different slide with crystal violet for 1 minute and then rinse it off. Then we added Gram's Iodine mordant for another minute and rinse it off as well. After that, we decolorized the bacteria by using 95% alcohol for 10-20 seconds. Last, we covered the slides with Safranin stain for about 30 seconds and rinse it off. After Gram Staining, we prepared for next week lab by isolating the DNA and setting up the PCR Amplification. We transfer a colony to 100 macro liter of water into a tube. Then, we allowed it to incubate for about 10 minutes in a 100 degrees celsius. We centrifuged the samples for 5 minutes at 13.400 rpm. During this step, we added 20 macro liter of primer/water mixture to 16S PCR reaction and placed the tube in the PCR machine.
Before we sawed the protists under the microscope, we noticed that there are less colonies on the plate that has tetracycline than the plates that does not. This suggests that there are few bacteria that are resistant to the antibiotic compared to the other plate. Tetracycline is an antibiotic that kills bacteria and those who are able to live under those conditions are said to be resistant. Also, we noticed that fungi grew in the plates that had tetracycline as well. Likewise, the protists that we saw under the microscope were mostly gram-positive. For example, the bacteria that grew in the agar plates without tetracycline has one gram positive and one gram negative. The first group from this particular agar plate was a orange colony. We noticed that they are cocci and they have cilia that enables them to move. The second colony from the agar plate that did not contain tetracycline was a white colony. We noticed that they were a bacillus shape, but they were not able to move and were bunch together. As to the ones that did contain tetracycline, we notice that both colonies were gram positive. One colony was a orange colony that had a cocci shape. It was able to move because of its cilia's that it has. For the other colony, it was a white colony. It had a rough surface and the bacterias had a spirillum shape.
Conclusion and Future Directions: From this experiment, we were able to gram stain the bacteria from each agar plate and were able to determine if it was a gram positive or gram negative bacteria. The diagrams above shows three pictures that are purple and one that is entirely pink. The ones that are purple are considered to be Gram positive and the one that is considered pink is considered to be gram negative. These indications allowed us to identify the shape of the bacteria under the microscope and see whether it had a method of motility. For future directions, I would take samples from of the same bacteria from a different colony and compared the results that we get. For example, if one is gram negative, then maybe the other colony that has the same bacteria should also considered to be gram negative as well.
- Erik Maradiaga 12:17, 5 February 2015 (EST): Erik Maradiaga
2.4.15 Excellent notebook entry. Plenty of detailed observations of Hay Infusion and protists identified. Well organized. SK
Lab Entry 2 1/27/15: Identifying Protists in our Hay Infusion Jar
Purpose: The purpose of this experiment was to identify protists that may be living in different niches in the Hay Infusion Jar. Some protists may be living in the surface of the water, and there might be some protists that may be living in the bottom of the jar where soil is present. The reason why my lab members and I did this experiment was in order to familiarize ourself how to use the Dichotomous key so that we would be able to classify protists that are present in our transect. Once again, there was no need for a hypothesis for this first experiment since we were viewing sections of our transect under a microscope.
Material and Methods: In order to view the protists under a microscope, we had to prepared the Hay Infusion Jar. We took 10 to 12 grams of ground vegetation into a plastic jar filled with 500 mLs of deer park water. We then added .1 gram of dried milk and shook the jar for 10 seconds. We removed the lid and let it sat in the classroom for seven days. After the Hay Infusion was prepared, we used the microscope and the Dichotomous key to identify protists in our jar. We took two slides and added a drop of sample from two different niches, the surface of the water and the bottom of the jar containing soil, into the slides by using a transfer pipette. We then added protoslo in order to slow the protists to be able to see them. From there, we viewed the slides under the microscope and identified protists by their size, shape, and method of motility.These factors allowed us to identified a certain type of protist in the different niches in our jar.
Figure 1: Microscopic View Figure 2: Aerial View of Hay Infusion Jar Figure 3: Diagram of Colpidium and Chlamydomones
From Figure 2, it can be seen that there was green shoots in the surface of the water. As my lab members and I got very close to the Hay Infusion Jar, we notice that there was a terrible smell. The water of the jar was mostly black with soil present in the bottom, and there was green algae in the surface of the water. However, we notice that the jar had the same volume of the water from last week which we assume that the water did not completely evaporated. We prepared slides from samples of the surface of the water without including the algae and from the bottom of the jar with soil present. My lab members and I notice that there were more organisms in the slide containing soil compared to the slide from the surface of the water. The organisms may differ close to versus away from the plant/soil matter because the organisms may need soil in order to be able to survive compared to the organism that is able to live in the surface of the water without the presence of soil. These organisms that live in the soil may need soil in order for food. In figure 3, we see the identification of the Colpidium and Chlamydomones protists. The Colpidium protist was taken from the bottom of the hay infusion. Under the microscope, it measurement was 50 micrometers and it had cilia in order for it to move. The protist does not photosynthesize as well. On the sample from the surface of the hay infusion, we found the protist Chlamydomones and Colpidium as well. Under the microscope, the Chlamydomones were around 5 micrometers. It was more circular and it was motile because of its flagella. It does not photosynthesize as well. The Colpidium was 60 micrometers, and it has the same functions as to the one we saw from the sample of the bottom of the hay infusion.
Conclusion and Future Directions: The protists that we identified that were living in our hay infusion jar was fascinating. My lab members and I was able to identify at least two protists that were living in different niches. However, we found that the protist Colpidium was living in both niches. We can assume that the protist may live when soil is present and when it is not present. For future directions, I may want to try to find protists that were close to the green algae on the surface of the water. There might be different protists in that particular niche and this would allow us to identified more protists in our transect.
- Erik Maradiaga 11:19, 30 January 2015 (EST):Erik Maradiaga ( Forgot to sign my entry yesterday)
1.27.15 Excellent first entry. Well organized and thorough. Just missing description of Hay Infusion set up. SK
1/19/2015 Observing our Transect at AU
Purpose: The purpose of this experiment was to introduce ourselves to our niche that we will be observing. In every ecosystems there are millions of living microorganisms that may be living in the soil, a certain type of plant, or anywhere else in the niche. The second part of this experiment was to see how evolution took place in the Volvocine Line. The reason why we are doing these two experiments is to introduce ourselves to the many different types of protist and to introduce ourselves with the concept of evolution. There was no need for a hypothesis in this first experiment.
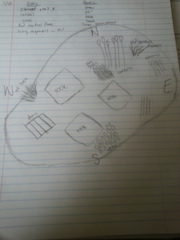
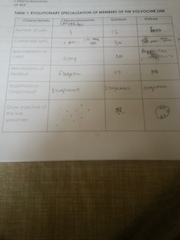
Material and Methods: When we observed our transect, my lab members and I were describing our location and topography. Second, we had to draw an aerial-view diagram of the transect and be specific in the observations we see. Along with this step, we had to indicate in the diagram north, south, east and west. Last, we had to list the abiotic and biotic components of our transect and indicate their position in the transect. For the second experiment we were doing, we had to use a microscope to view the Chlamydomonas, Gonium, and Volvex. We prepared a slide by using a pipet and dropped each organisms to our slide. We repeated this step for all three organisms. Second, we observed the numbers of cells present, colony size, specialization of cells, mechanisms of motility, isogamous or oogamous, and we drew a picture of the specimen.
In the first diagram, it is a picture of our transect. It is located in front of Kogod School of Business. My lab members and I notice that there was a drain close to the niche, so we assume that there was water present in the niche. The area is a pentagonal shape with all sorts of living and non-living things present. For example, we saw that cattails were present in the niche, and there was also red cardinal flowers. likewise, there were rocks present in the niche. When my lab members and I listed the biotic components we saw in our niche, we listed that there was cattails, grass, red cardinal flowers, bushes, and the living organisms in the soil. For the abiotic components, we listed that there was snow, soil, rock, a drain, and a Dunkin Donut cup present in the transect. The second image is a picture of the results when we saw the 3 living organisms under the microscope. When we observed the Chlamydomonas under the microscope, we notice that it is a simple cell with a flagella in order for it to move. As we continue to see the Gonium and Volvex under the microscope, my lab partner and I notice that the level of complexity increased. For example, the Gonuim now has a range of 4-32 cells present which is different from the Chlamydomonas. Also, the Volvex is different from the other two cells because it has specialized cells that help it to reproduce and in order to do the process photosynthesis.
Conclusion and Future Directions: Since in this lab experiment there was no hypothesis, the data does not refute or support the hypothesis. We took a sample of soil from the transect in order to make a hay infusion jar for next week experiment so that we can conclude what type of protist are there in our transect. For future directions, I may want to take a sample of soil from the middle of the transect instead from the side close to the cement because I think that more protists are living near the cattails. There may be protists that have a relationship with the cattails or the red cardinal flowers that is present in the middle of the transect.
- Erik Maradiaga 13:37, 25 January 2015 (EST):EM