User:Deanna Fleary/Notebook/Biology 210 at AU
3/19/15 Zebrafish Experiment
Purpose: The purpose of this lab was to observe the development of zebrafish embryos under the condition of rhodamine (specifically rhodamine 6G) and Deerpark water. Because of its use only as tracer and lack of evidence supporting health effects on larger level vertebrates, rhodamine will not effect the embryogenesis of the zebrafish. If there is a lack of developmental difference observed between the zebrafish from the control condition and those from the experimental condition then Rhodamine does not effect embryonic development.
Materials and Methods: 20 healthy zebrafish embryos were used for both of the conditions and placed in a petri dish with 20mLs of the condition liquid. Observational changes of the zebrafish were observed over a two week period including stage of development, motion, pigmentation, and mortality rate.
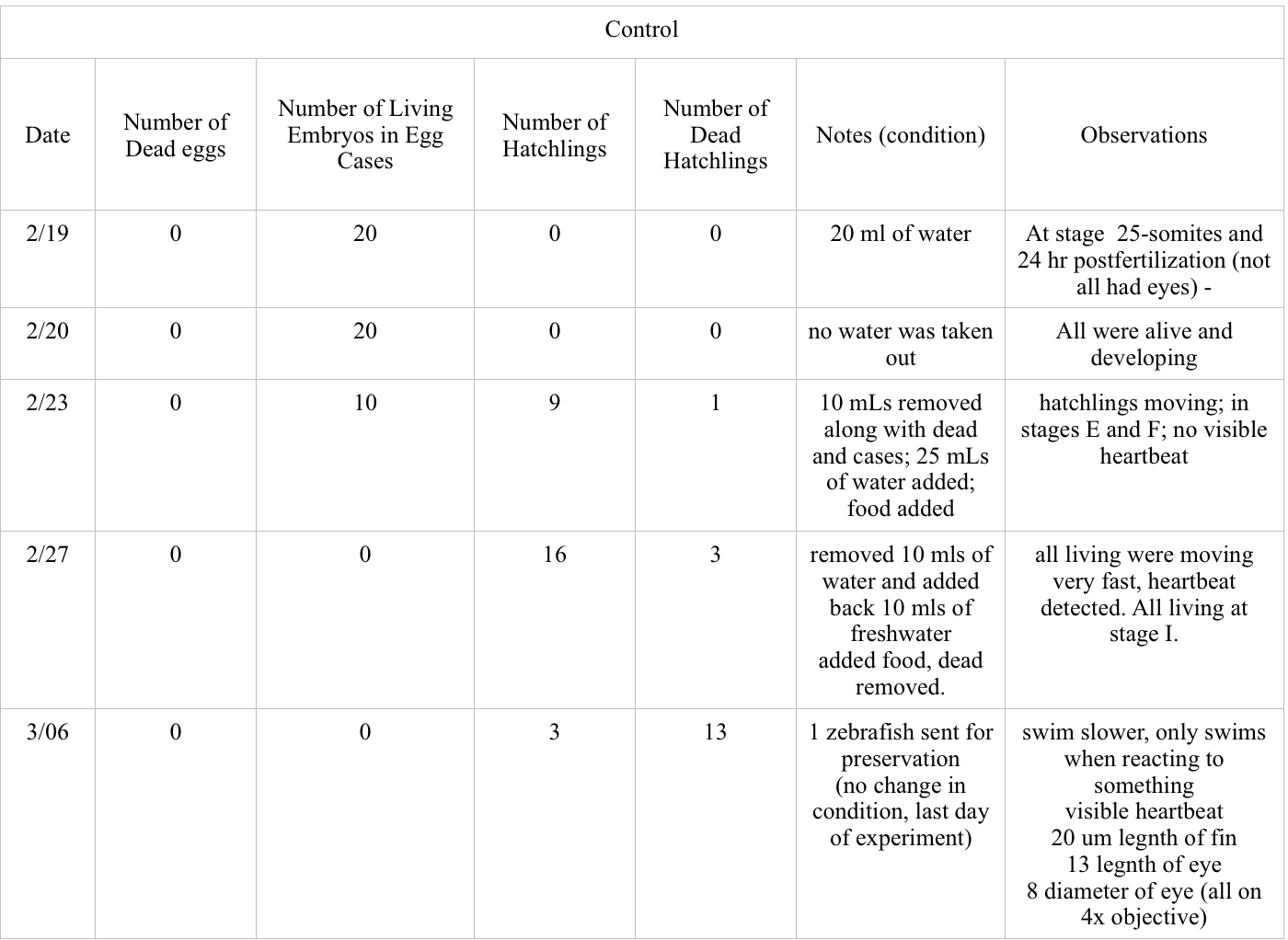
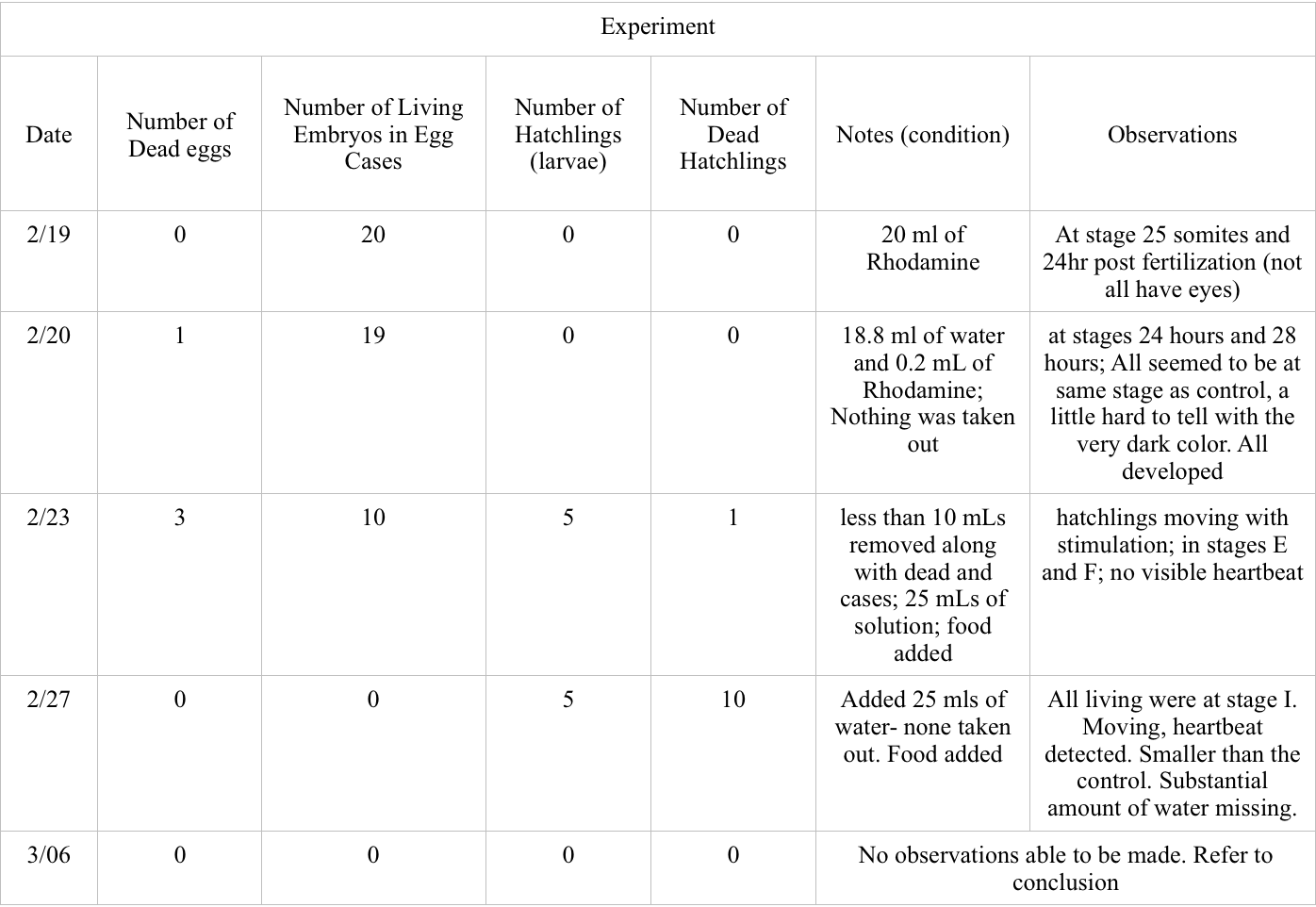
Data and Observations:
Conclusions and Future Directions: Based on the data collected between 2/19 and 2/27, there was a much higher occurrence of dead zebrafish in the experimental group. 4 of the experiment eggs died between this time as compared zero for the control, and 11 hatchlings died as compared to the 4 of the control. More of the control embryos reached hatchling with the highest amount of 16 on 2/27 compared to the 5 hatchlings in the experimental on the same day. Both groups had similar stages of development, the only observed difference being the change in observed size on 2/27.
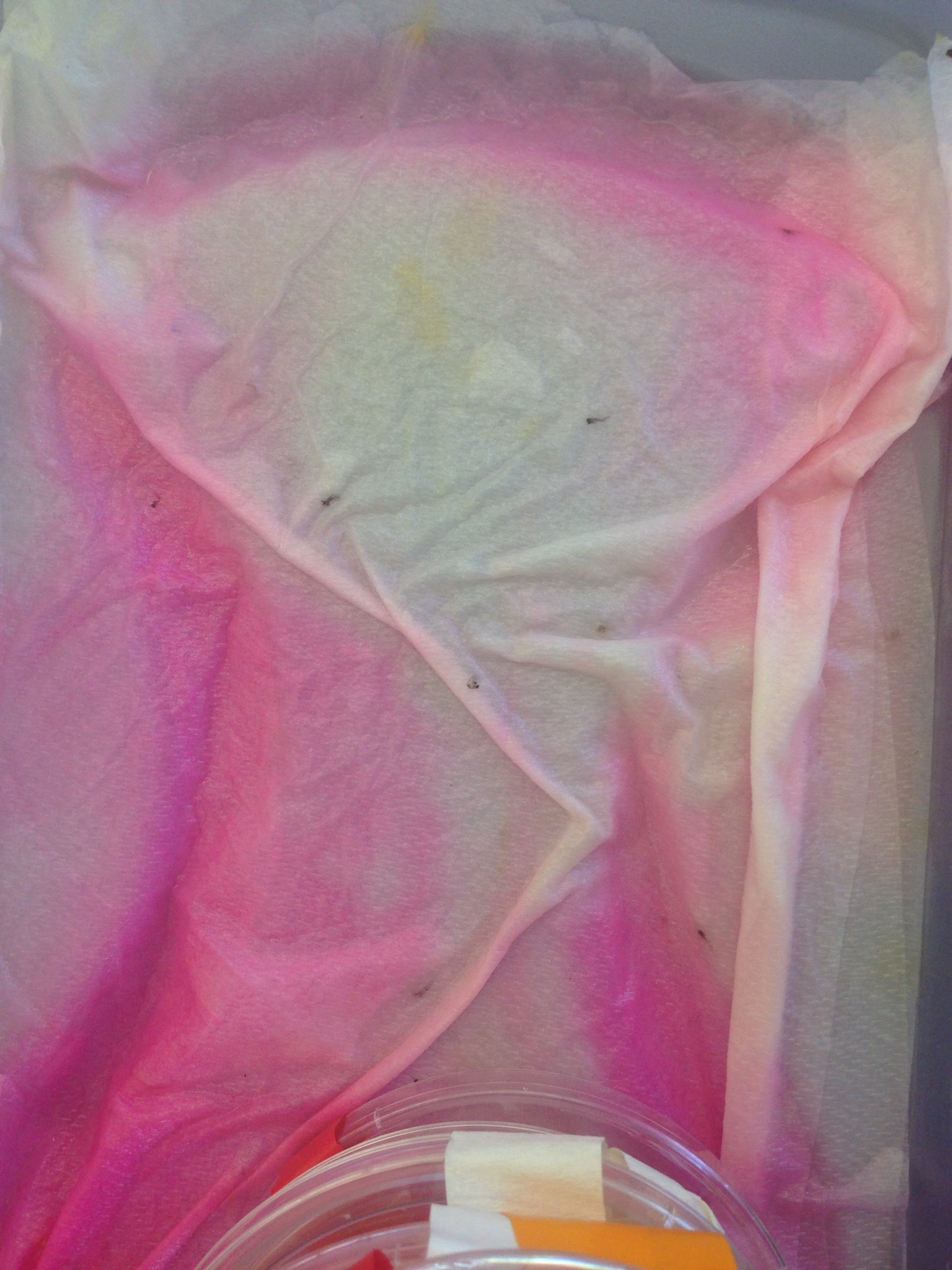
The environment in which the experiment was conducted greatly impacted the results making it difficult for non influenced data to be collected. The greatest interference that occurred when conducting the experiment happened between 2/27- 3/06 where it was found that the experimental petri dish had been emptied into the bin, shown in the picture.
Because our group was the only one to use any condition that had a color liquid, the samples at the bottom of the bin was ours. We can only assume that our experiment was accidentally tampered with when another group was collecting their samples since the petri dishes were piled in stacks within the bin. This experiment must repeated for any sound conclusions to be drawn on the effect of rhodamine 6G on zebrafish embryonic development.
Sources:
Bentley, M., Walters- Conte, K., Zeller, N.K. (2015) A Laboratory Manual to Accompany: General Biology II. Washington, DC: American University.
RHODAMINE 6G CASRN: 989-38-8, (2014). Retrieved March 3, 2015, from http://toxnet.nlm.nih.gov/cgi-bin/sis/search2/f?./temp/~s8SBlw:1.
DEF
3/5/15 16S Sequence Analysis
Purpose: The purpose of this lab was to identify the bacterial samples taken from the Hay Infusion Culture. Through PCR amplification of the 16S rRNA gene we were able to identify the bacterial species and observe whether or not our own observations of the DNA match.
Materials and Methods: Initial procedure for the DNA amplification is described below in the Microbiology and the Identification of Bacteria post. Once the 16S rRNA gene was amplified, two of the PCR products were sent for sequencing. Once the order results were received the sequence of each of the two tubes was reviewed. The entirety of this sequence was then put through nucleotide blast, which compares our sequence to known nucleotide sequences, which then gave the results of the bacterial species.
Data and Observations:
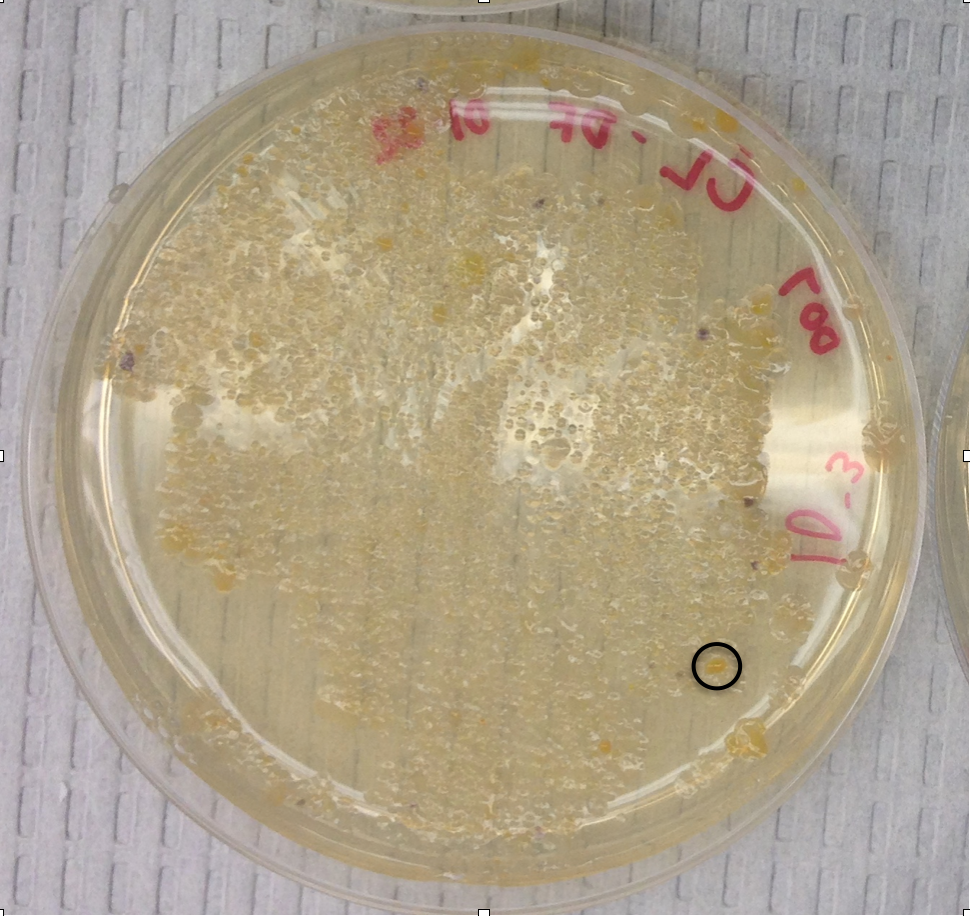
Sample 1 Nutrient 10-3:
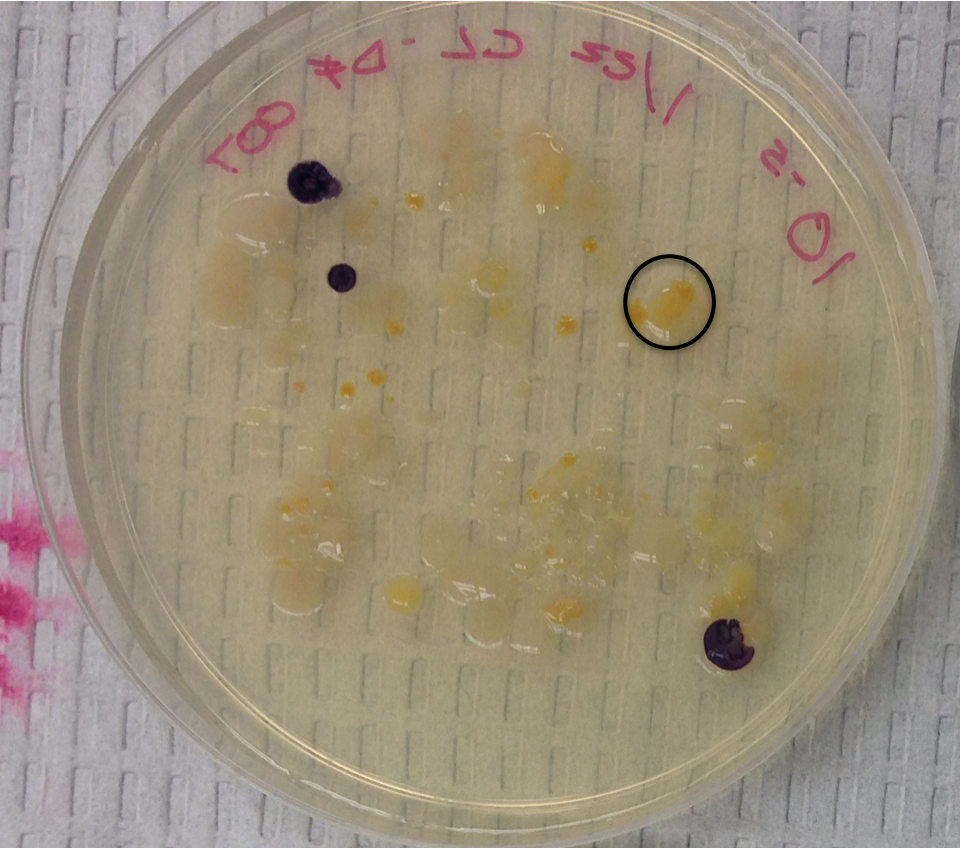
Sample 2 Nutrient 10-5
Raw sequence of sample 1 (Nutrient 10-3):
NNNNNNNNNNNNGNAGGCTANNNTGCAGTCNANCGGTNGTANNNCNNTANNGNATTCNTGANANCGCCGTAGAGGGGCGAATCATGNCTGCNACCCGGCCTGGTCGTGGGGATNNCCTTTCCAAAGGAANATCTNACCCCNTAACGTCCNANNGNANAAANTTGNGACTTCACTCCTTTGGATATCGANGGGCCNCGTCGGANNNNNTNNTGNNAGGGNNNTGGCCCACCNANGNATGNNCTTTAGGGGGNCGANAGGGTGATCCCCCCACTGNACNGAGCANNNNCNNACTCCNACNGNAGGCAGCANNGAGGANNATTGNACNATGGGTGAGAGCCTGATCCNCCATGCCGNNTGNNNGATGANNGCCCTATGGGTTGTAAACTTCTTTTGATNNGGATAAACCTTTCCACGTGTGGAAAGCTGAANGTACTATACGAATAAGCACCGCCAACTCCGTGCCNGCAGCCGCGGTAATACNGANGGTGCANNCGTTATCCGNATTTATTGTATTTAAAGNATCCNTANGCGGATCTGTAANTCAGTGTTGAAATCTCACNNNTTAACTGTNAAACTGNAATTNATACTGCAAGTCGNGACTAAAGTATANNTGACTNGAATAAAAANTGTAGCAATGAGTACAAANATATTACTTAGTACNNAAATTGCGAGNGCAGGTGGATGNCTCTCTGNNTGNNACCNACGAGA
Raw sequence of sample 2 (Nutrient 10-5):
NNNNNNNNNNNNNCNANNNTGCAGCCGAGCGGTAGAGATTCTTCGGAATCTTGAGAGCGGCGTACGGGTGCGGAACACGTGTGCAACCTGCCTTTATCAGGGGGATAGCCTTTCGAAAGGAAGATTAATACCCCATAATATATTAAGTGGCATCACTTGATATTGAAAACTCCGGTGGATAAAGATGGGCACGCGCAAGATTAGATAGTTGGTAGGGTAACGGCCTACCAAGTCAATGATCTTTAGGGGGCCTGAGAGGGTGATCCCCCACACTGGTACTGAGACACGGACCAGACTCCTACGGGAGGCAGCAGTGAGGAATATTGGACAATGGGTGAGAGCCTGATCCAGCCATCCCGCGTGAAGGATGACGGCCCTATGGGTTGTAAACTTCTTTTGTATAGGGATAAACCTTTCCACGTGTGGAAAGCTGAAGGTACTATACGAATAAGCACCGGCTAACTCCGTGCCAGCAGCCGCGGTAATACGGAGGGTGCAAGCGTTATCCGGATTTATTGGGTTTAAAGGGTCCGTAGGCGGATCTGTAAGTCAGTGGTGAAATCTCACAGCTTAACTGTGAAACTGCCATTGATACTGCAGGTCTTGAGTAAGGTAGAAGTAGCTGGAATAANTANTGTANCNNTGAAATGCATAGATATTACTTANAACACCAATTGCGAAGGCANGTTAGTATGTCTTAACTGACNCTGATGGACNANNNCGTGNGGAGCGAACANGATTGNATACCCTGGTNGTNNNCNCCATAAACGATGCTAACTCNTTTTTNGNCTTCNGCTTCAGANACTAANNNAAANTGANAAGNTTAGNNNCCTGGGAGTANGTTTCNCAGANTGATNAAANNAANTNACGGGNGCCCNCGCNCCCNCTNNAGTTATGTNNTTTATTNNTTNATACNGANANNNNNNNAGNNANNNNTAATGGCGANNGNGGGNGNNNGNNNNNGNGNGTGCNTTNNTCANNTTNGNNCTTNNTNCNTGGCNNNNGNGGGNNNGGTNNCCNNNNNNNNGNNNNNNGNNNNNNNNNCNGNNNGGNN
Sample 1 was identified as: Chryseobacterium sp. EECC-510. Sample 2 was identified as: Chryseobacterium sp. THG A18.
Conclusions and Future Directions: Both bacterial samples were identified as a part of the genus Chryseobacterium. The biological classification for the twos samples is listed below.
Kingdom: Bacteria Phylum: Bacteroidetes Class: Flavobacteria Order: Flavobacteriales Family: Flavobacteriaceae Genus: Chryseobacterium Species: EECC-510 or THG A18
Chryseobacterium are gram negative bacteria that are known to be filamentous, non-motile, and typically rod shaped. The colonies are observed to be round with a deep yellow coloring and tend to not exceed 2mm in diameter (Cunniff).
From data observations made on the bacterial sample in the lab, neither samples would have been identified as Chryseobacterium. Sample 1 (Nutrient 10-3) was observed to be gram negative like Chryseobacterium but the bacterial cells were identified as round not rod shaped. Contrasting this, sample 2 (Nutrient 10-5) as observed to be gram positive unlike the Chryseobacterium, but did have the same rod shaped bacterial cells. The colonies of the two samples were observed as a deep bright yellow similar to the description common to Chryseobacterium.
The reason for the lack of continual similarity between the bacterial samples and the Chryseobacterium characteristics could be because of our lack of experience performing a gram stain and observing bacteria. This was mine and my partners first time performing these techniques making it more likely that we made a mistake in the process.
Sources: Cunniff, J. (2012) Chryseobacterium indologenes. Microbe Wiki. Retrieved from https://microbewiki.kenyon.ed/index.php/Chryseobacterium_indologenes.
DEF
2/12/15 Vertebrates and their Individual Niches
Purpose: Both invertebrates and vertebrates make up an ecosystem. This lab aims to complete our understanding of life within our ecosystem by identifying the vertebrates present.
Materials and Methods: When the transect was visited during the morning on two separate occasions, only humans were presently visible within the transect. Other vertebrates were identified based off of the species which have been seen in other parts of campus closer to life. The bird species were identified by researching which species are common in the DC area.
Data/ Observations and Discussion
| Table 1: Vertebrates Present within a Certified Wildlife Habitat | ' | ||||||
| Vertebrate | Phylum | Class | Order | Family | Genus | Species | Biotic/ Abiotic Benefits |
| Human | Chordata | Mammalia | Primates | Hominidae | Homo | H. sapiens | bench, stone walkway, tree, info plaque |
| Mouse | Chordata | Mammalia | Rodentia | Muridae | Mus | Mus musculus | bush |
| Squirrel | Chordata | Mammalia | Rodentia | Sciuridae | Sciurus | Sciurus carolinensis | bush, tree |
| Song Sparrow | Chordata | Aves | Passeriformes | Emberizidae | Melospiza | Melospiza melodia | tree, soil, worms |
| Pine Siskin | Chordata | Aves | Passeriformes | Fringillidae | Spinus | Carduelis pinus | tree, soil, worms |
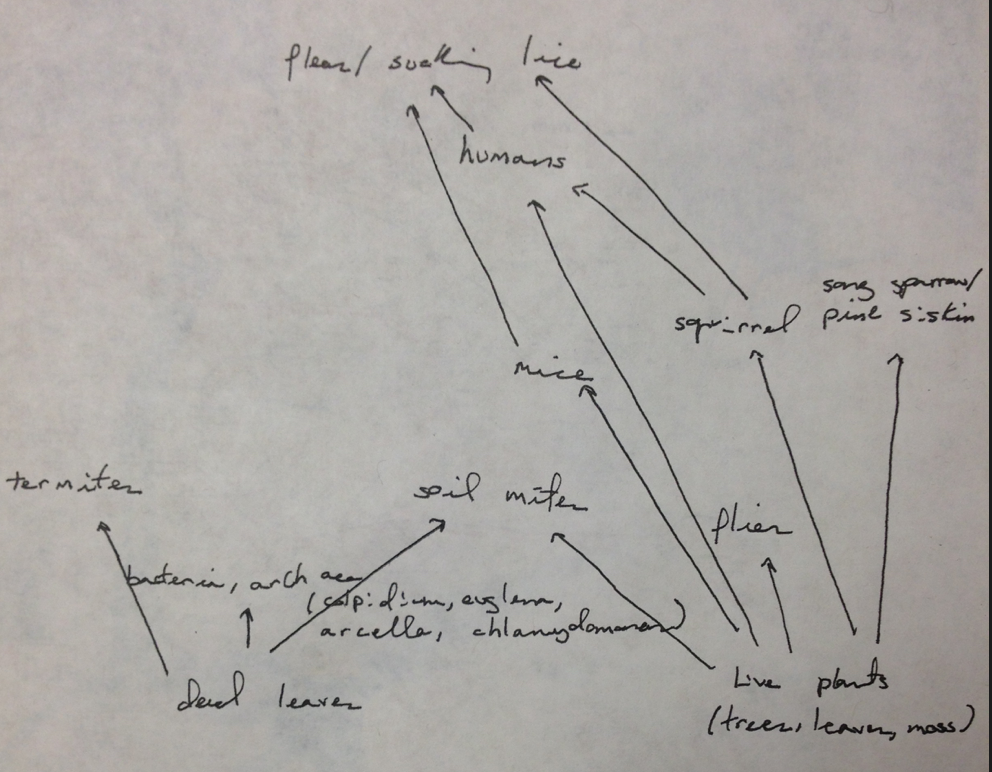
Image 1: Food Web of all observed organisms.
Each organism holds its own tropic level presenting their feeding level place within the ecosystem of the transect. All of the species observed in the transect represent a community in that they all interact with each other while directly or indirectly because they share the same living space. Each individual species observed live in the habitat because they are support by the resources present. When the resources can no longer support the species populations then they have reached the carry capacity for that habitat.
Conclusions and Future Directions: This lab as well as the others before it granted us with an understanding of the different lifeforms present within an ecosystem. The diversity of life which was observed represent a delicate interactions between the different organisms and how the sustaining of life for all is somehow connected to the life of another.
Sources:
http://anthro.palomar.edu/animal/table_humans.htm
http://www.itis.gov/servlet/SingleRpt/SingleRpt?search_topic=TSN&search_value=180366
http://bioweb.uwlax.edu/bio203/s2012/toellner_kayl/classification.htm
http://eol.org/pages/1052763/overview
http://eol.org/pages/1051055/overview
DEF
2/12/15 Diversity among Invertebrates and their Physical Characteristics
Purpose: In this lab different invertebrates were observed so as to understand the evolution and complexity of their organ system. By observing different species like the planaria and annelida the effect of organ structure, specifically of the circulatory system, was observed to impact motion. Other invertebrates within the transect were also observed further continuing the education on the diversity of life present within an ecosystem. The diversity of life aids in the stabilization of an ecosystem.
Materials and Methods: For the first part of the lab three different species (planaria, nematodes, and annelida) were needed in order to observe the difference and evolution of change of their organ system. Different tools like a dissecting microscope, compound microscope, and dissecting instruments also aided in the observations. for the second part of the lab different invertebrates were identified and observed. The invertebrates were first collected by using a Berlese Funnel. To set up the Berlese Funnel, 25mL of 50:50 ethanol/ water solution was placed with a conical tube (Lab Manual pg. 43). A screening material was then taped to the bottom of the funnel to stop leaf litter from falling into the preservative. Leaf litter sample was then placed into the top of the funnel and was then set up to a ring stand after the funnel was attached to the conical tube. The funnel/ tube were placed under a 40 watt light bulb and then covered with foil. The funnel was placed in this condition for a week allowing the invertebrates to move down into the preservative.
Data and Observations: The flat worm (planaria) move in a gliding motion. The overall movement is seamless and similar throughout the entire body. This could possibly be caused by that fact that it is an Acoelomate and does not have a body cavity. It has a solid body and a simple digestive system. The round worm (nematodes) moves in a S- shape motion. It forms a whiplike shape that moves throughout the body. This is because the round worm has an incompletely formed body cavity called pseudocoelom. Earth worms (annelida) move in a contraction motion in which their a part of the body becomes smaller as it moves closer to the other part. The is because it has a fully lined and filled body cavity not allowing it to move in one continuous motion.
| Table 1: Invertebrates Present within a Certified Wildlife Habitat | ' | |||
| Organism | Phylum/ Class | Length (mm) | Number in Sample | Description of Organism |
| Flea (Siphonaptera) | Arthropoda/ Insecta | 0.25 | 1 | without wings, body laterally compressed- legs made for jumping |
| Flies (Diptera) | Arthropoda/ Insecta | 1 | 1 | with wings, single pair of wings |
| Soil Mite | Arthropoda/ Arachnida | 0.75 | 1 | without wings, body not laterally compressed, minute insect, parasitic on birds and mammals, sucking mouth part |
| Sucking Lice (Anoplura) | Arthropoda/ Insecta | 0.5 | 3 | without wings, body not laterally compressed, minute insect, parasitic on birds and mammals, sucking mouth part |
| Termites (Isoptera) | Arthropoda/ Insecta | 3 | 1 | without wings, body not laterally compressed, rarely minute, not parasitic, social insect, soft body, abdomen broadly joined to theorize, development by simple metamorphosis |
The range of size of the collected invertebrates is .25mm to 3 mm. The smallest organism was the observed flea, and the largest was the termite. Both are part of the same phylum and class: arthropoda/ insecta. Of the invertebrates observed, the ones most common to leaf litter are mites that eat decaying plant material. Though not observed from the Burlese Funnel, common organisms include slug and snails which are apart of the mollusca phylum, and roundworms which are apart of the nematoda phylum (Catley, Johnson, 2002).
Conclusions and Future Directions: This lab encouraged a greater understanding of the diversity between invertebrates and how systems become more complexed. The presence of the invertebrates within the transect further increased the understanding of the diversity present within an ecosystem. Further experimentation can be done to understand the other forms of life present within the transect.
Sources:
Bentley, M., Walters- Conte, K., Zeller, N.K. (2015) A Laboratory Manual to Accompany: General Biology II. Washington, DC: American University.
Catley, K., Johnson, E. (2002) Life in the leaf Litter. American Museum of Natural History. Retrieved from file:///Users/deannafleary/Desktop/LifeInTheLeafLitter.pdf.
DEF
2/5/15 Plantae and fungi Present within the Certified Wildlife Transect
Purpose: The diversity and characteristics of plants were observed in this lab. Such diversity include height, presence of a visualization system, and the mechanism of reproduction. Fungi was then observed in order to understand their importance within the transect. Such observations increase the understanding of the diversity of life present within the transect.
Materials and Methods: In order to observe characteristics between plant samples within the transect, representative samples from five plants was collected from inside the transect. The samples and their seeds were then observed using a dissecting microscope in order to characterize them. Fungi sample from the 10-9 agar plate with nutrient and tet was then observed also using a dissecting microscope and comparing it characteristics against identified fungi samples.
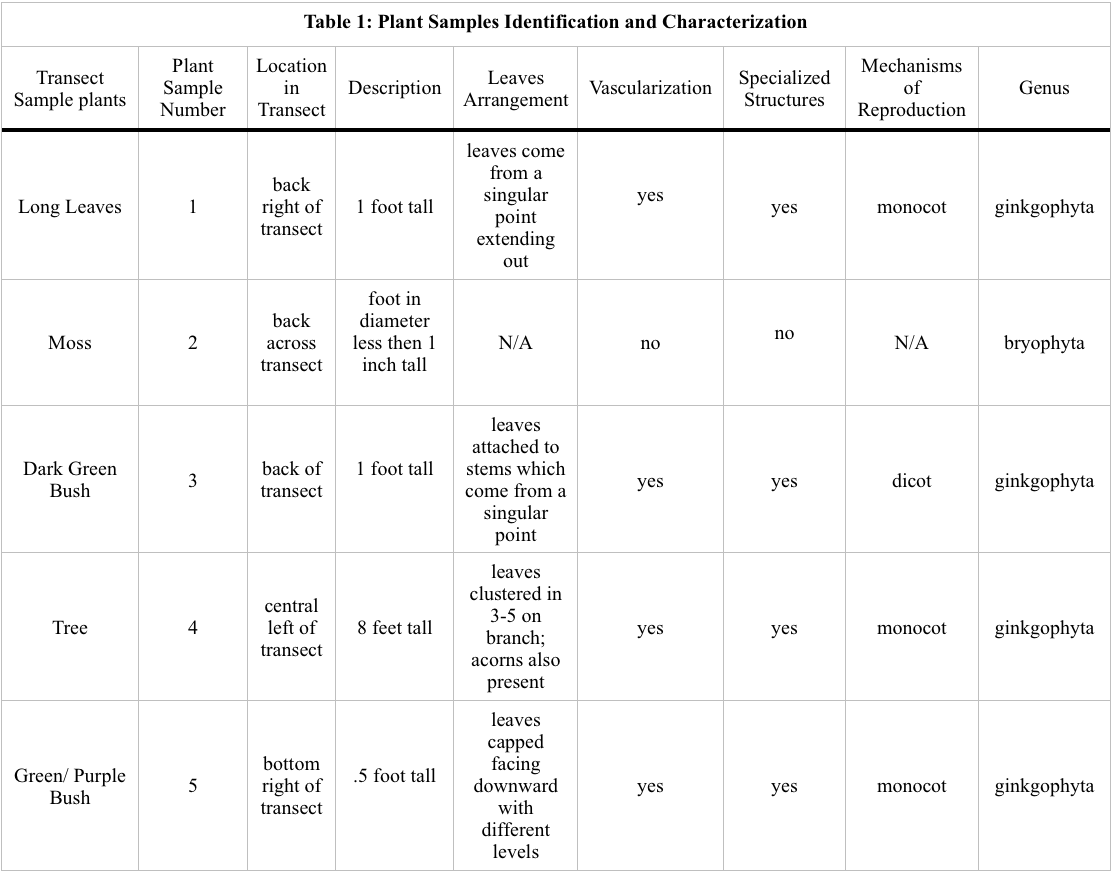
Data and Observations:
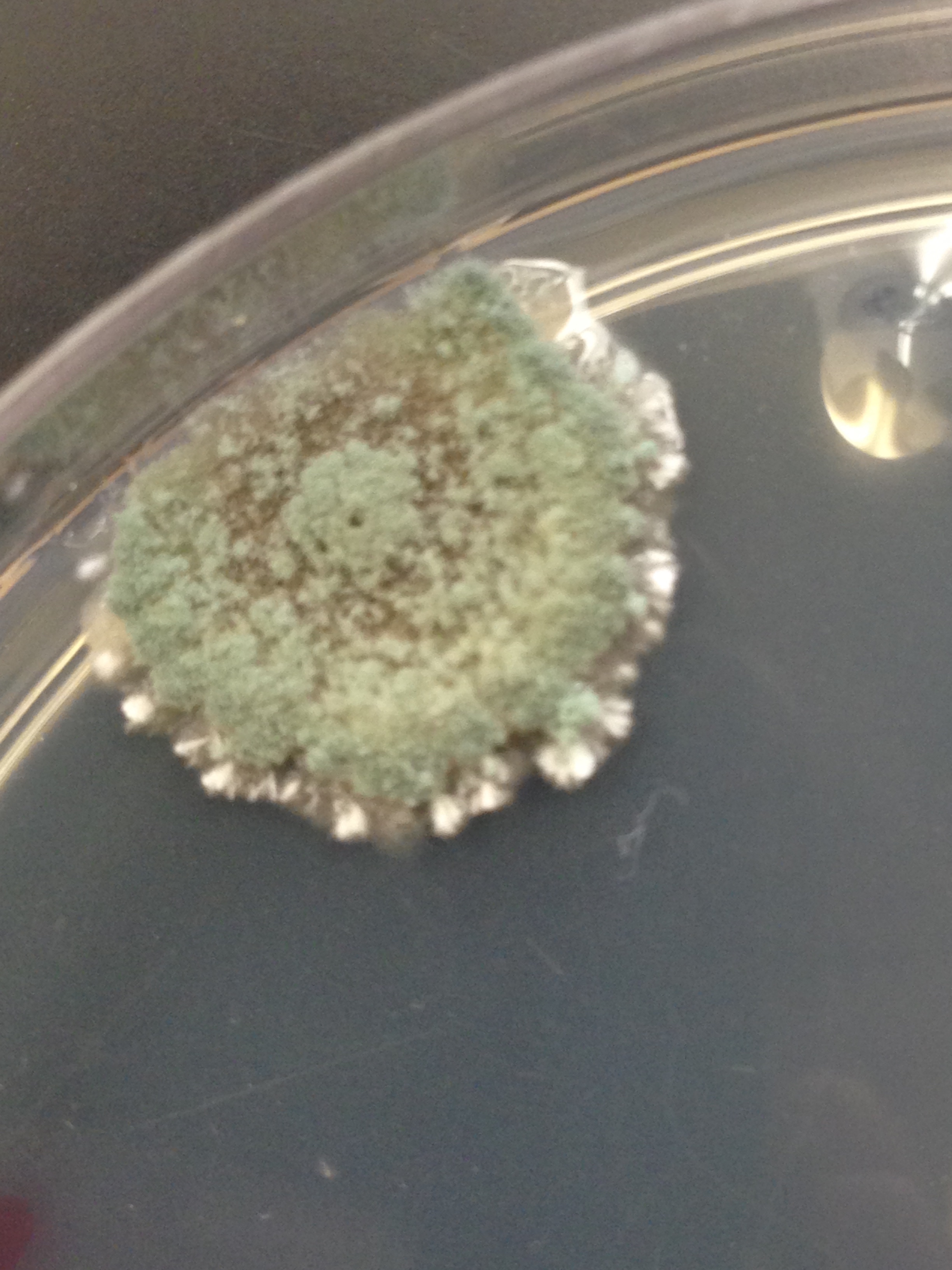
Fungi sporangia are round structures that contain spore cells. Such structures are important because the facilitate the reproduction of further fungi cells by opening and releasing the spores. The fungus observed from the 10-9 agar plate with nutrient and tet approximately four centimeters in diameter. The outside ring was white and the inside was green and black, but overwhelmingly green. The sample was classified as an ascomycota because of its similarity to known fungi samples.
Conclusions and Future Directions:
A high degree of commonality as found between the different plant samples. Such similarities was possibility caused by the plant samples taken and do not show a true representation of the diversity present within the transect. Further experimentation can be done increasing the amount of samples so that possibility a greater diversity can be observed.
DEF
1/29/15 Microbiology and the Identification of Bacteria
Purpose: In this lab, the bacteria within the Hay Infusion Culture was studied in order to understand bacterial characteristics, the effect of antibiotics on bacterial growth and how DNA sequencing can be used to differentiate between species. It is unlikely that archaea species will be identified on the agar plates because they tend to grow in extreme environments in which they are native. Numerous types of bacterial species will be observed during this lab. If different characteristics exist between the samples then the group’s Hay Infusion Culture contains different samples.
Materials and Methods: The experiment began by first observing the eight agar plates and the colonies that grew on them. This allowed for comparison of the plates between those with and without the antibiotic. Half of the bacteria was then observed on a wet mount slide so as to gain cellular description. A gram stain of the colony growth was then conducted in order to differentiate the different microorganisms. Slides were prepared for four of the bacterial colonies by using a sterilized loop to scrape up the sample from the agar plate and mixing it with water present on the slide. The slide was then heat fixed and using a staining tray was covered with crystal violet. Following this step the stain was then rinsed and a similar process was applied on the slide using Gram’s iodine mordant then 95% alcohol to decolorize, followed by with the safranin stain. Excess water was then removed from the slide and allowed to air dry so that the sample could be observed using a microscope. The final part of the lab was used to set up for the PCR amplification. For each of the characterized bacterial colonies, a single colony was placed into a sterile tube with water and incubated at 100°C for 10 minutes. The four tubes were then centrifuged fr 5 minutes at 13,400 rpm. After completion 5ul of each sample were added to the primer/ water mixture including the PCR bead and placed into the PCR machine.
Data and Observations: The Hay Infusion Culture contained appearance had change since it was observed last week. The waster level ahas decreased since before and is more dirty/ brown/ murky. Less green was visible in the culture and a translucent film was present on top of the liquid. The spoiled odor was greater than before, possibly due to the increased level of decomposition.
| Table 1: 100 Fold Serial Dilutions Results | ' | |||
| Dillution | Agar Type | Colonies Counted | Conversion Factor | Colonies/ mL |
| 10-3 | Nutrient | 300 | x103 | 300000 |
| 10-5 | Nutrient | 130 | x105 | 13000000 |
| 10-7 | Nutrient | 0 | x107 | 0 |
| 10-9 | Nutrient | 1 | x109 | 100000000 |
| 10-3 | Nutrient+ Tet | 160 | x103 | 160000 |
| 10-5 | Nutrient+ Tet | 2 | x105 | 200000 |
| 10-7 | Nutrient+ Tet | 0 | x107 | 0 |
| 10-9 | Nutrient+ Tet | 1 Fungus (black/white) | x109 | 0 |
Photo 1: Cultures w/oTetracycline
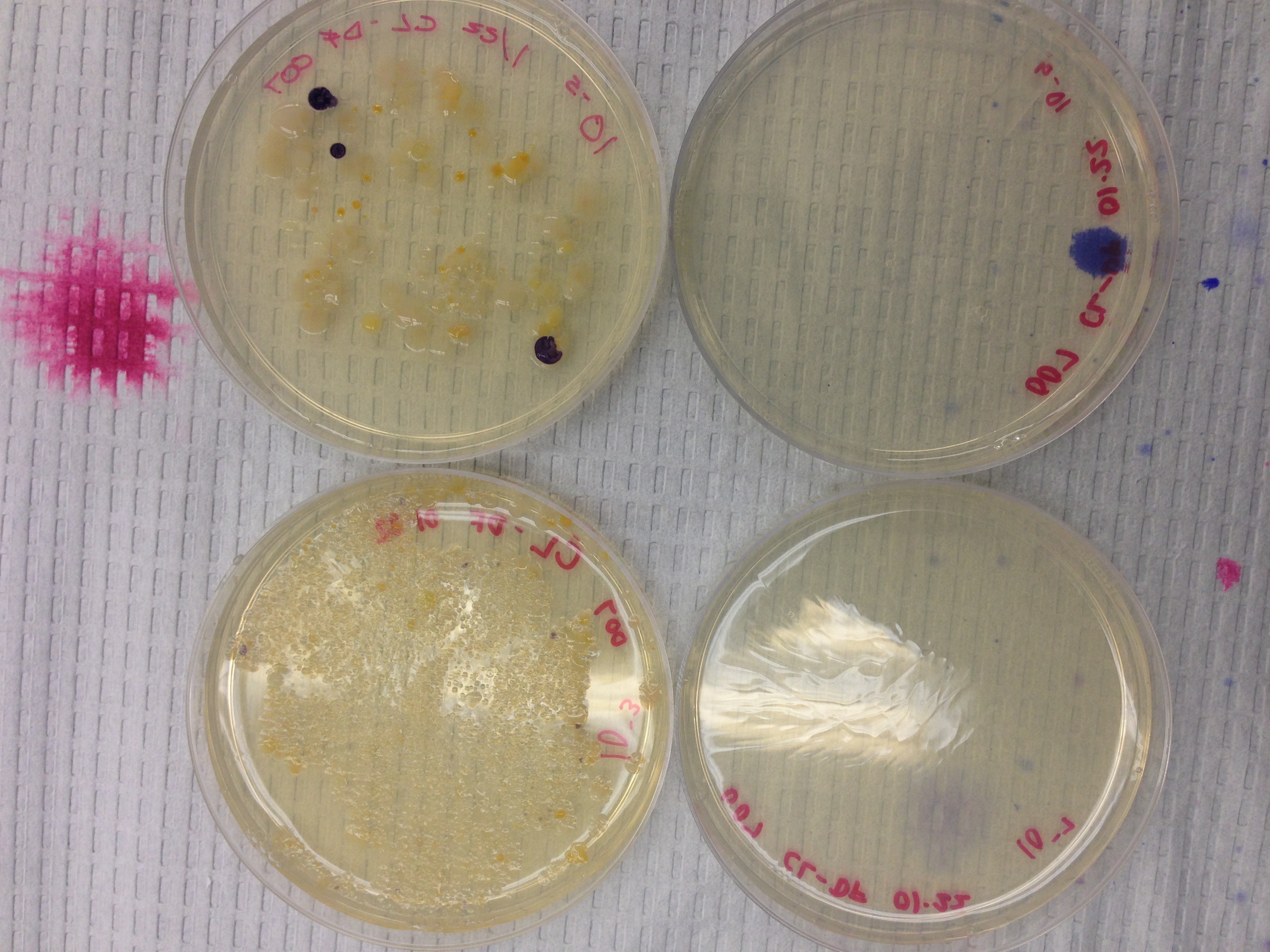
Photo 2: Cultures w/ Tetracycline
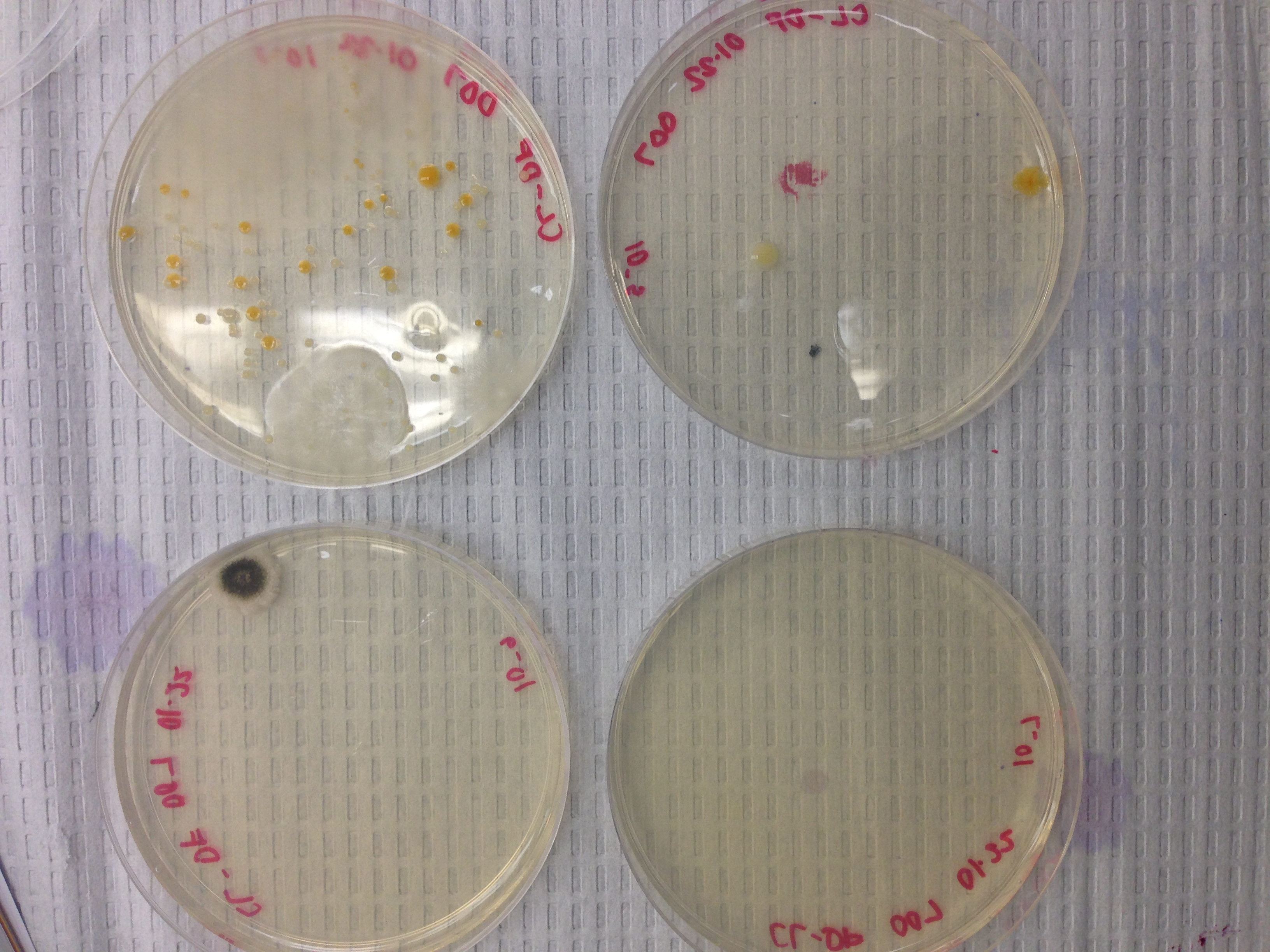
Colony amount was more abundant on the plates that did not contain the antibiotic tetracycline. This indicates that some of the bacteria present is greatly effected by the antibiotic because a clear decrease in colony amount was observed. Only one fungi appeared on the plate. Based off of this singular appearance a conclusion can be drawn that tetracycline does not effect fungi growth, but further experimentation would have to be done to draw such a conclusion. Tetracycline is effective against a wide range bacteria including both gram positive and gram negative. This means that bacteria that have extensive carbohydrate peptidoglycan in their cell wall and bacteria that have a thin peptidoglycan layer and outer phospholipid bilayer are both sensitive.
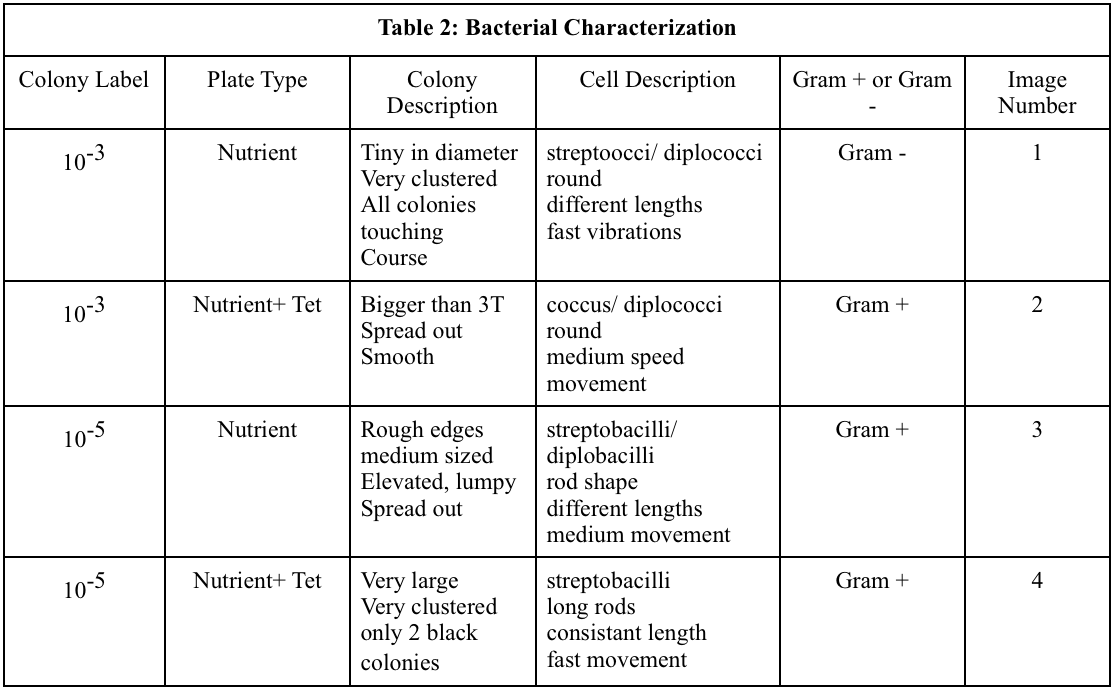
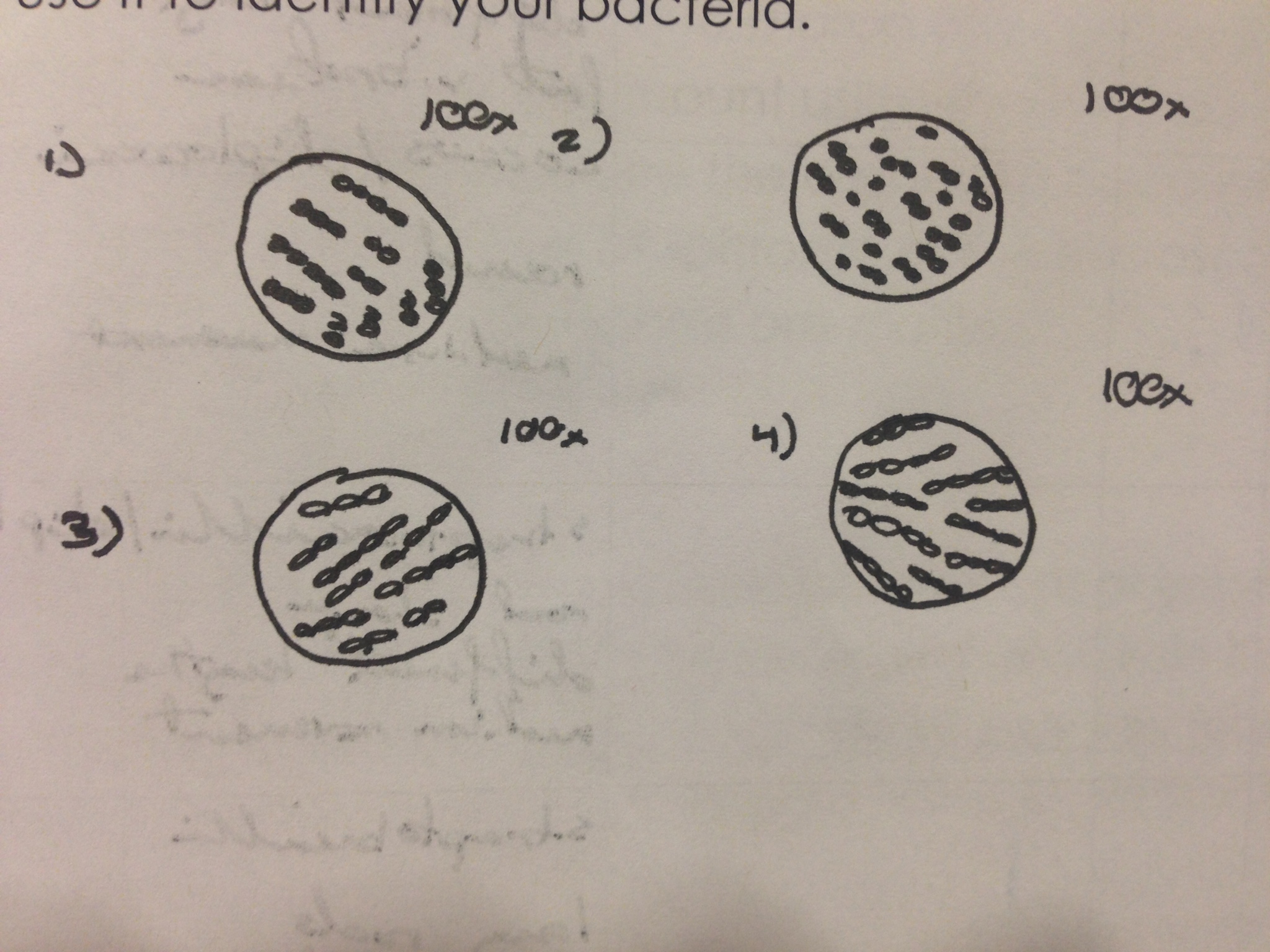
Table 2: Bacterial Characterization
Photo 3: Bacterial Description Drawing
Photo 4: Gram Satin 10-3
Photo 5: Gram Satin 10-3T
Photo 6: Gram Satin 10-5
Photo 7: Gram Satin 10-5T
Different characteristics were observed between the different bacterial samples. The largest variation between the different samples were the length of the bacteria and if they were cocci or bacilli. The gram staining revealed that both gram positive and gram negative cells were present and both are effected by the antibiotic.
Conclusions and Future Directions: From this experiment it was discovered that the bacterial samples were effect by the antibiotic Tetracycline. The antibiotic was shown to be effected against different types of bacteria. Further experimentation will be done to identify the different bacterial samples. Due to PCR amplification and the selective amplification of the 16S rRNA gene, we will be able to identify the different species.
Sources:
Bentley, M., Walters- Conte, K., Zeller, N.K. (2015) A Laboratory Manual to Accompany: General Biology II. Washington, DC: American University.
Chopra, I., Roberts, M. (2001) Tetracycline Antibiotics: Mode of Action, Applications, Molecular Biology, and Epidemiology of Bacterial Resistance. American Society for Microbiology. Retrieved form http://mmbr.asm.org/content/65/2/232.full.
DEF
1/22/15 Identification of Protist and Algae within an Ecosystem
Purpose:
Observe and identify free living protozoa present within different niches of a culture. Observations/ identifications are done so as to understand that different niches have unique carrying capacities for each species. If different species are identified in their respective niche, then the food and space provided by the niches are most optimal for each species.
Materials and Methods:
For the lab, the 500 mLs of liquid from the Hay Infusion Culture was used representing an ecosystem. Samples were drawn from two different niches, one located at the top near plant life and the other at the bottom. While under a microscope organisms were identified using the Dichotomous Key by comparing the seen organism against numerous physical characteristics.
Data and Observations:
From first observations the Hay Infusion Culture had a very pungent smell. A translucent film was located on the top and around the edges. Plant life was located throughout the infusion with most of the larger leaves floating near the top. Located at the top was also green spots. The bottom contained the majority of the plant life excluding the large leaves. The water itself was a cloudy yellow.
| Table 1: Organism Identification | ' | ||
| Niche 1: Top Film (near plant life) | |||
| Organism | Size | Characteristics | Photo Number |
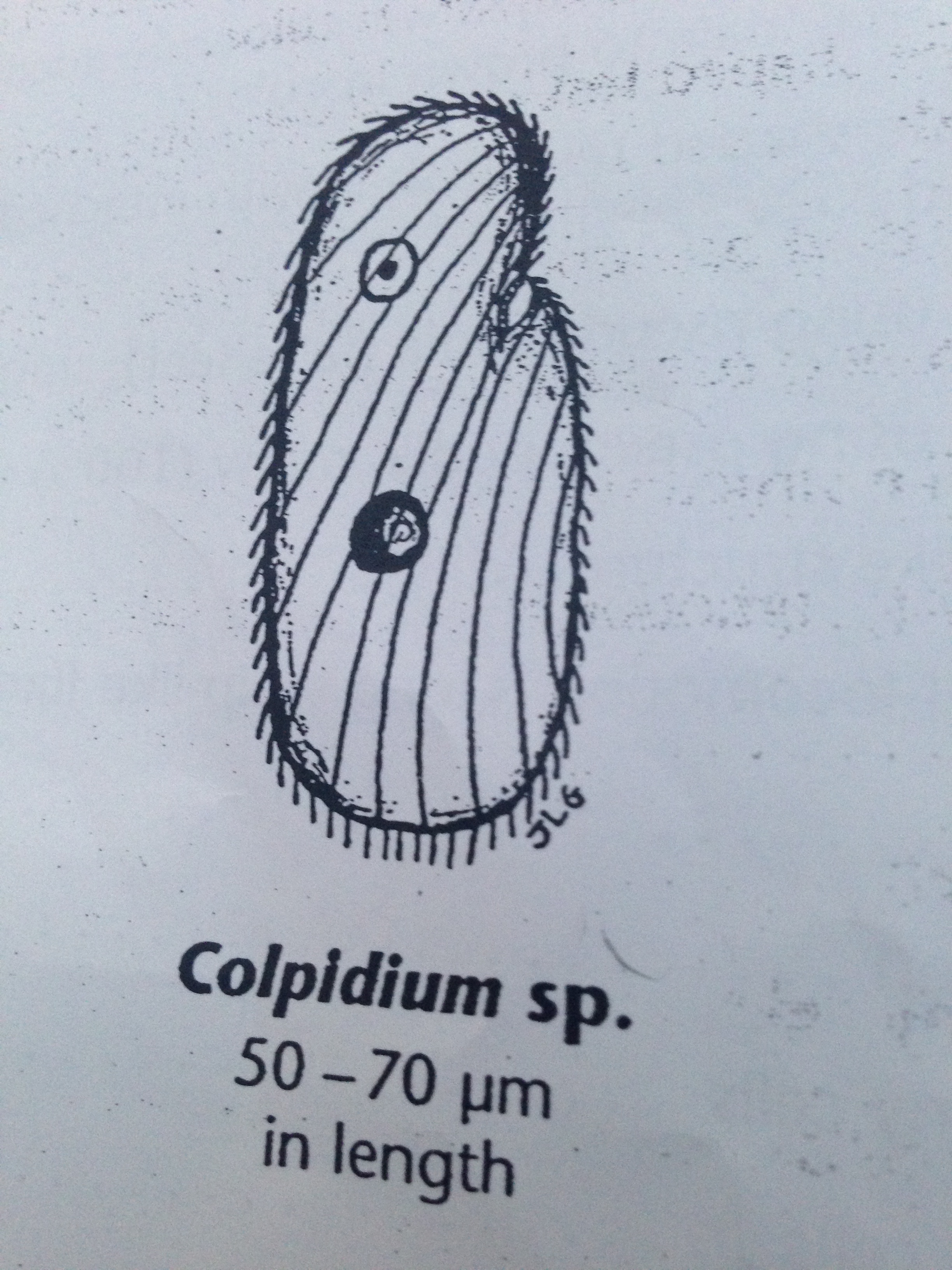
| Colpidium | 50-70 um | colorless, small body, oval shape, fast movement, cilia, | 1 |
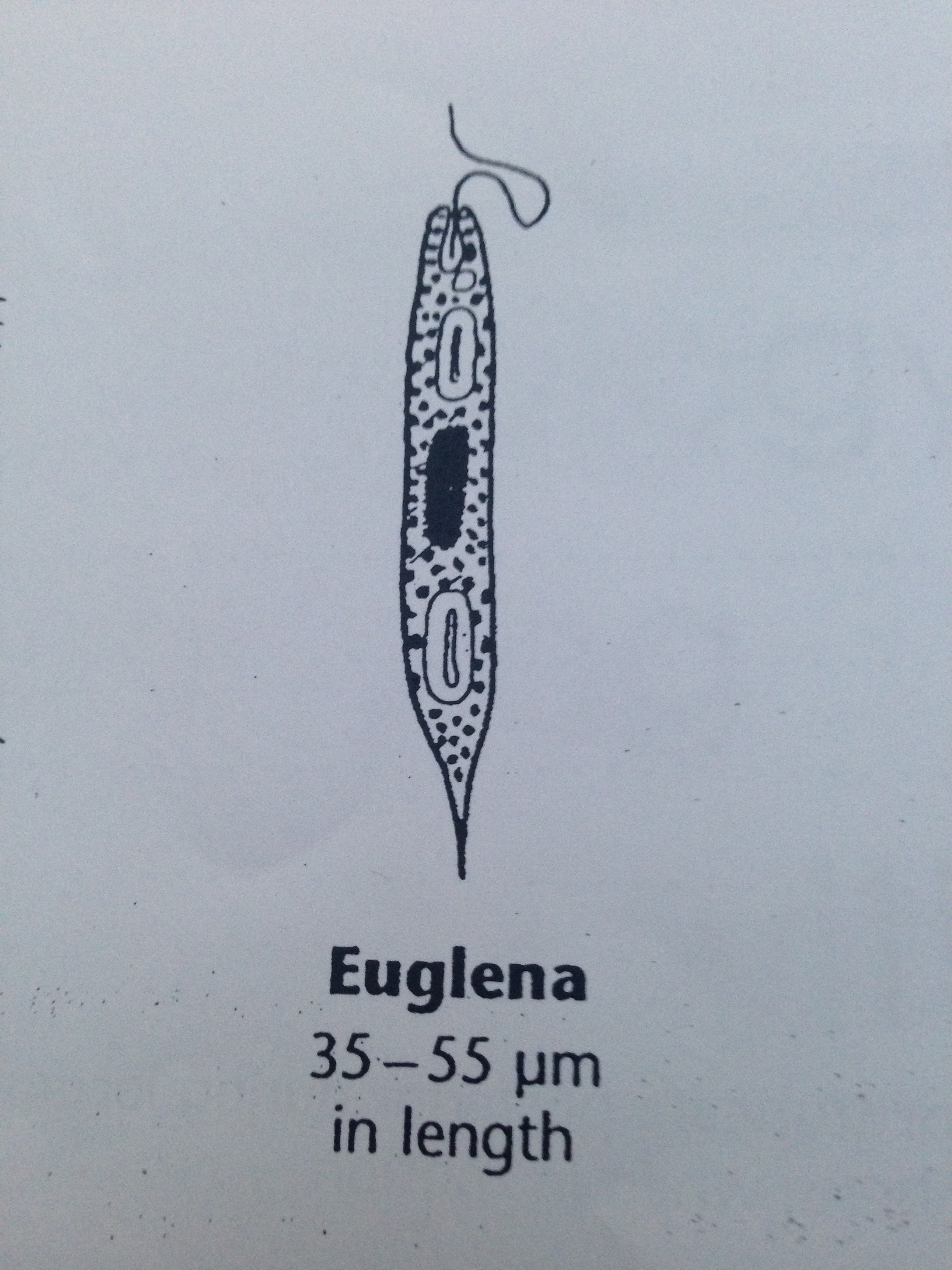
| Euglena | 35-55 um | green, single cell, cell elongated, one observed locomotor flagella, | 2 |
| Niche 2: Bottom | |||
| Organism | Size | Characteristics | |
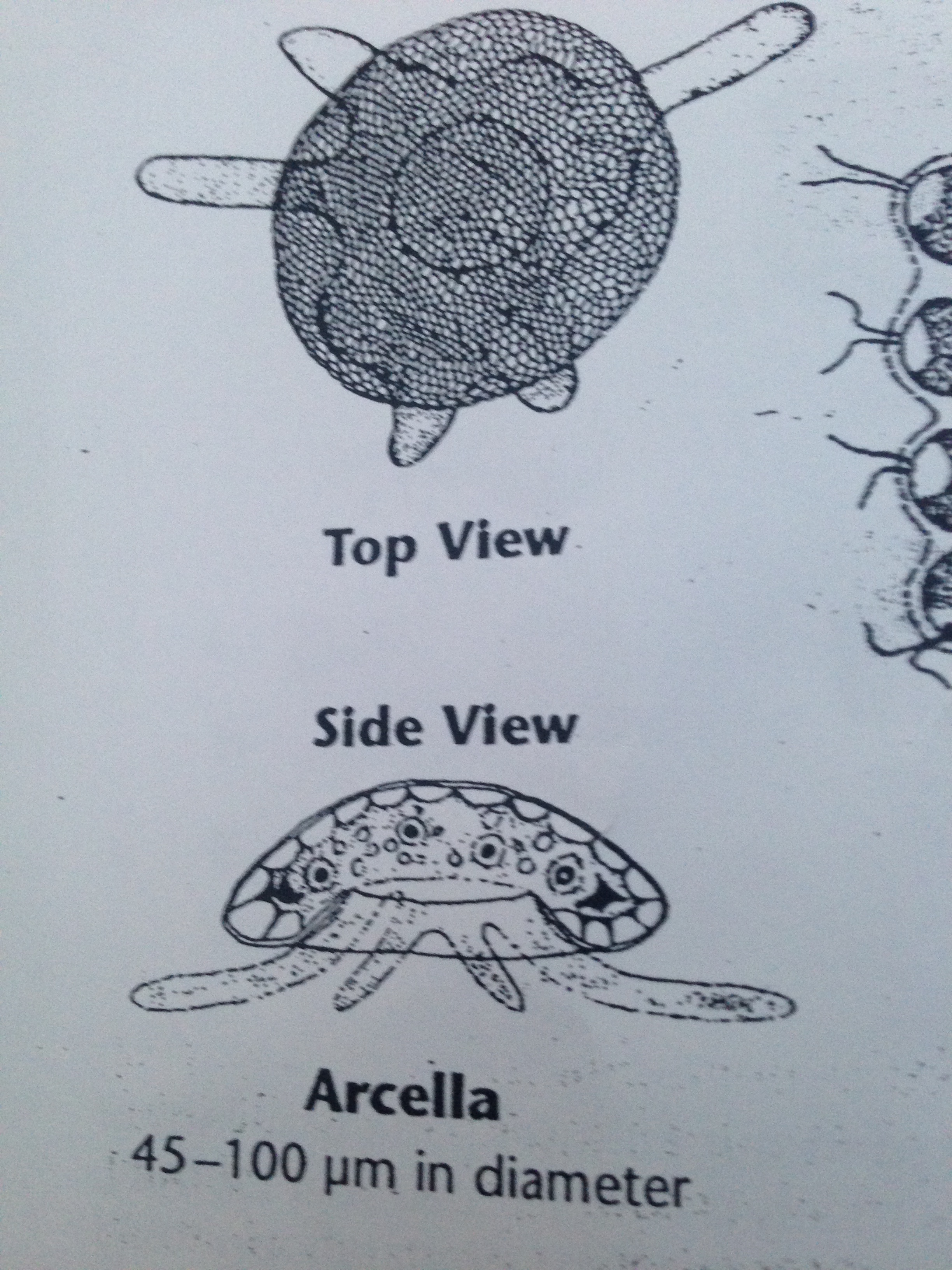
| Arcella | 45-100 um | colorless, consistent shape, not spherical, slow movement without apparent motion | 4 |
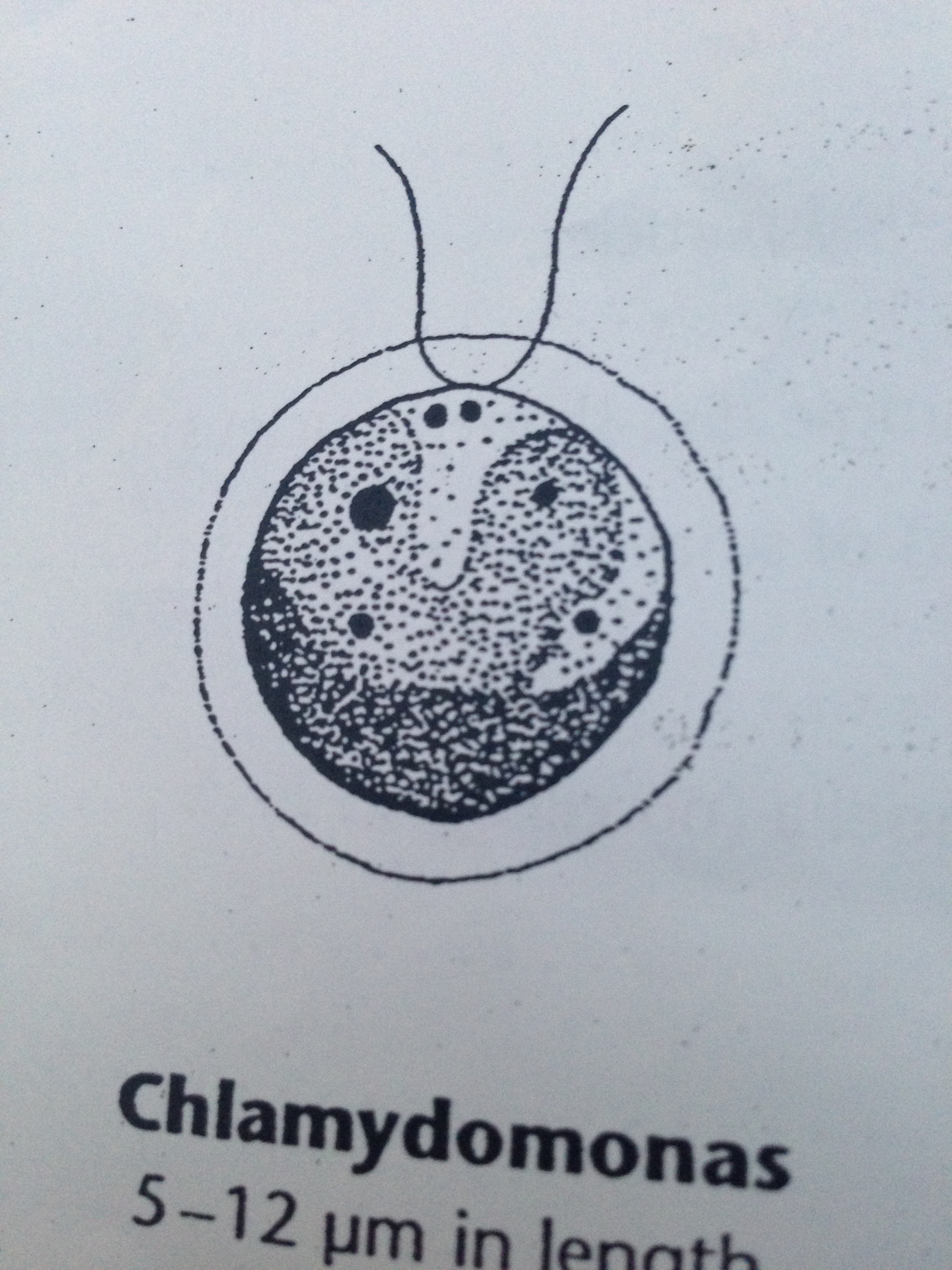
| Chlamydomonas | 5-12 um | green, single cell, motile, oval shape, two observed locomotor flagella, | 3 |
| Colpidium | 50-70 um | colorless, small body, oval shape, fast movement, cilia, | 1 |
Photo 1
Photo 2
Photo 3
Photo 4
Conclusions and Future Directions:
An organism might be located close to plant matter because of their necessary life function. If they require similar nutrients that plant do or are able to profit from the products of plants then their presence in the plant matter proximity would be understandable.
Euglena are single cell organisms often found in water without a current and an unpleasant smell like the ecosystem within the Hay Infusion Culture. Euglena meets the requirement of being classified as a life form because it 1) is a cell that regulates the material that passes through the membrane 2) performs photosynthesis to acquire and use energy 3) processes genetic information and respond to the environment 4) reproduces through binary fission 5) and is a product of evolution.
If the Hay Infusion Culture were to “grow” for another two months I would expect the life forms present and the abundance of these life forms to change. This would be in response to the change taking place inside Hay Infusion Culture. Because of the further decomposing of the plant life present and the further evaporation of water the available resources present in the culture now would change in two months. Further experiment could be done to observe the change in organism presence in a two month period.
DEF
12/15/15 Transect 2 at American University
Purpose: Observe the ecosystem within our transect. Observations are done so as to understand interactions and complexities present in different niches of an ecosystem. The characteristics of our transect will be composed of both biotic and abiotic features, which all when interacting together form an ecosystem.
Materials and Methods: For the lab, a 20 by 20 meter transect is needed which will be the focus of the observations. The general characteristics of the transect, including location and topography were described. An ariel view diagram was collected as well as a list of the biotic and abiotic features within the transect.
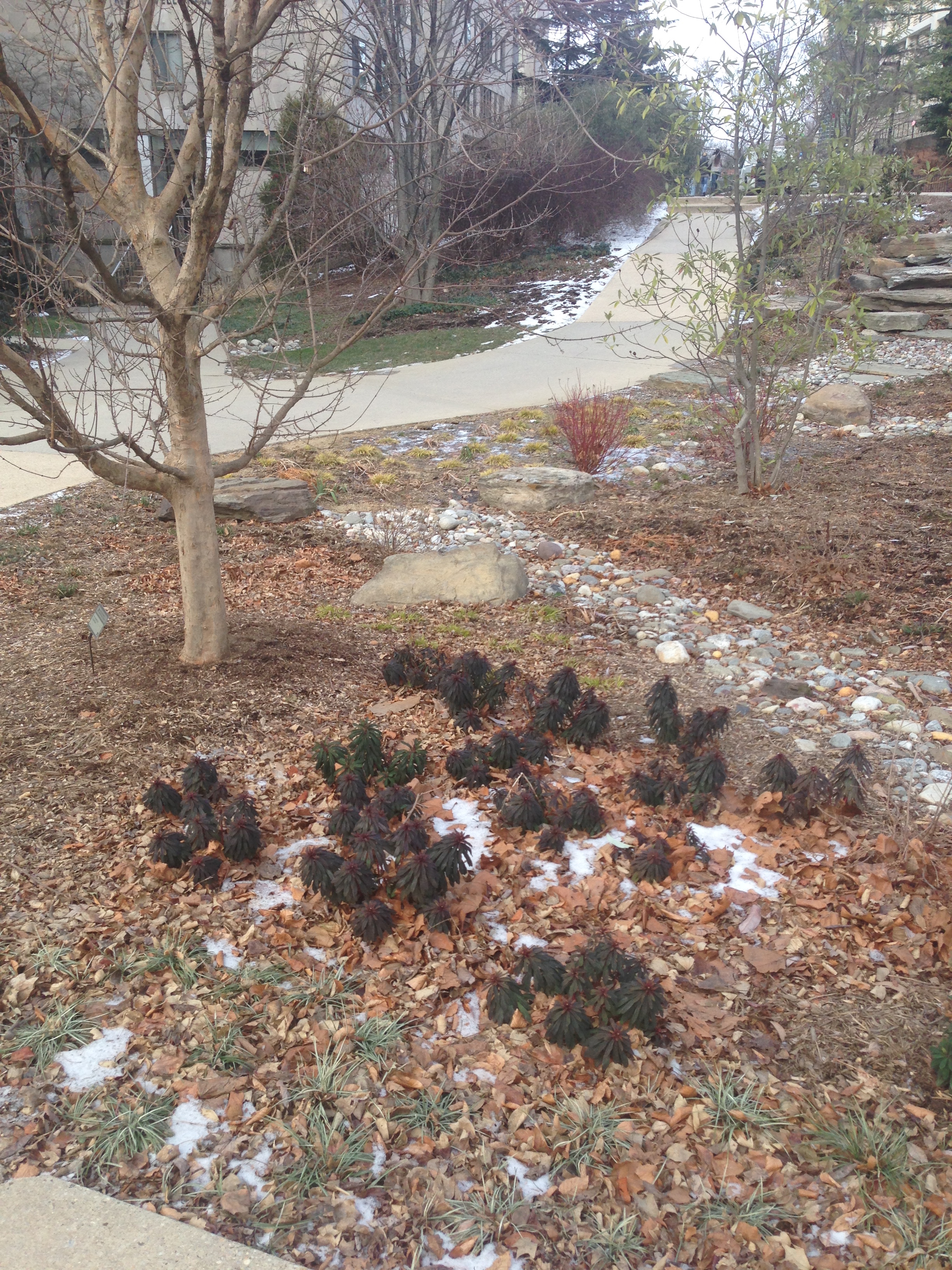
Data and Observations: Transect 2 is located on the Northside of the American University campus near Hughes Hall and McDowell Hall. The area is open with few trees and view obstructions, and the placement of many of the components were man chosen and did not appear there naturally.
| Table 1: Abiotic and Biotic Features with Transect | ' | |
| Feature | Characterization | Location Number |
| Tree | Biotic | 1 |
| Moss | Biotic | 2 |
| Leaves | Biotic | 3 |
| Bush | Biotic | 4 |
| Soil | Biotic | 5 |
| Informational Plaque | Abiotic | 6 |
| Bench | Abiotic | 7 |
| Stones | Abiotic | 8 |
| Snow | Abiotic | 9 |
| Cigarette | Abiotic | 10 |
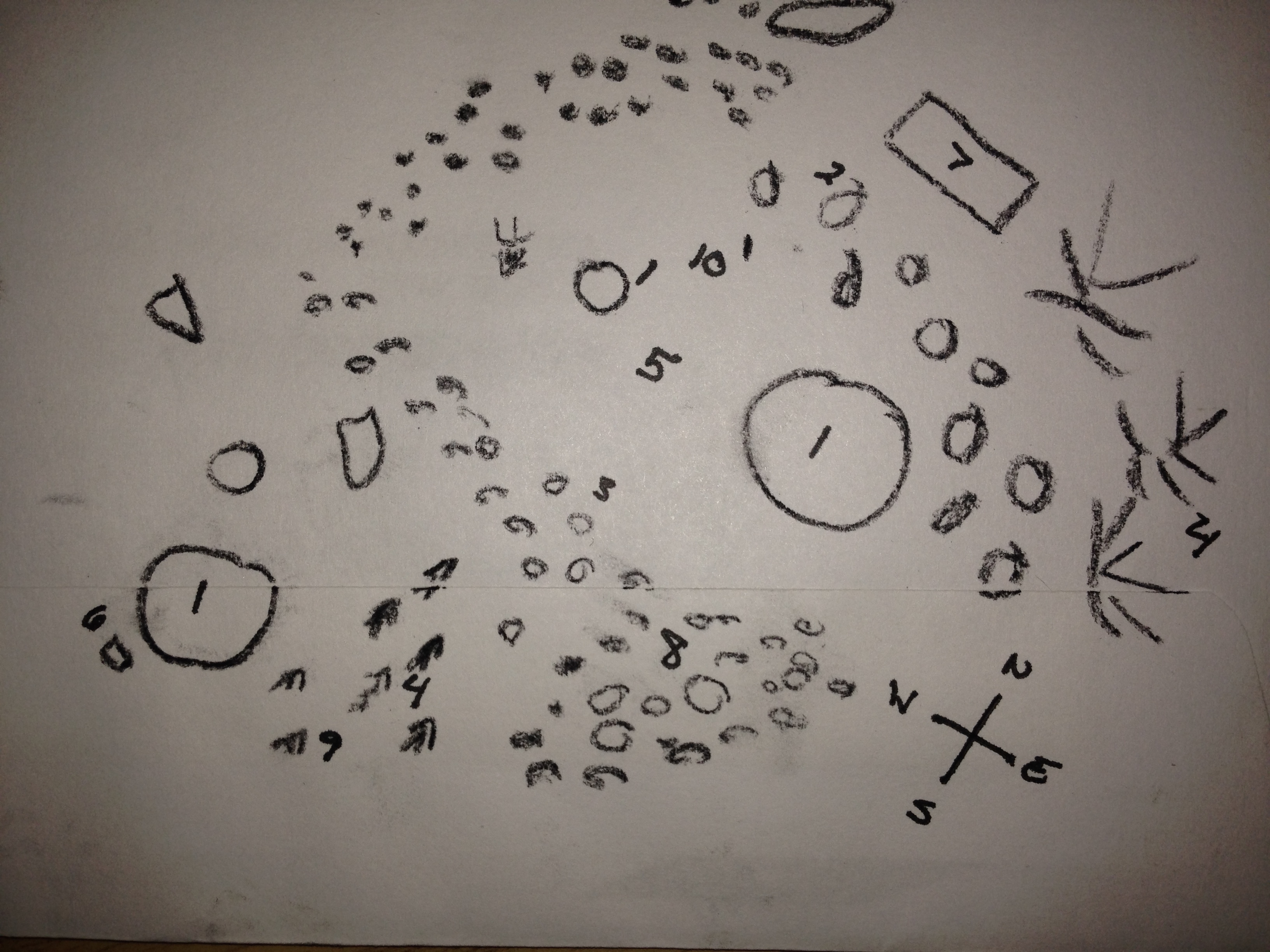
Aerial Transect View:
Photo 1:
Photo 2:
Photo 3:
Conclusions and Future Directions: The transect contained both abiotic and biotic features which together make up the ecosystem. Further experimentation will be done to identify the organisms (plants, bacteria, protist…) in the transect so that the interaction and complexities present within an ecosystem can be understood.
DEF
1/21/15
Public Health major, history minor.
DEF