User:April Rose Schattner/Notebook/Biology 210 at AU
March 19, 2015 - Zebrafish Study & Observations
Purpose:
The purpose of this experimental observation is to study the effects of variations in embryonic and post-embryonic environments on the development of zebrafish. Zebrafish are often used in studying factors influencing embryonic development because the embryos are visible for measurement and observation through their transparent embryonic sac. In this study, we explore how caffeine influences growth and development of zebrafish between day one and day fourteen of their life. Our hypothesis states that zebrafish embryos exposed to caffeine will grow to be smaller in sized than zebrafish in a control setup of Deer Park bottled water.
Materials & Methods:
In this study, twenty zebrafish embryos were collected at random and placed in a 20 mg/L caffeinated aquatic environment. They were added to the aquatic environment between approximately hours 20 and 24 of their embryonic development. A control group of twenty zebrafish embryos were collected at random and placed in an aquatic environment of Deer Park bottled water. Observations of the zebrafish in the two groups were collected over a 14 day period. The water in each sample was refreshed on day four, seven, eleven, and fourteen to ensure the fish had fresh water. On day four of observations, the samples were checked for dead zebrafish embryos that might be molding in the samples. If any were present, the dead embryos were removed. On day seven of observations, the remaining living zebrafish in each group were fed with paramecium. Also on day 7 of observations, three zebrafish from each sample were extracted at random and euthanized for preservation and observation. Measurements of body length and eye color were collected, as well as notations on coloring patterns and appearance. On day fourteen, any remaining zebrafish in the samples were observed and, finally, euthanized.
Data & Observations:
| Day | Remaining Zebrafish: Caffinated | Remaining Zebrafish: Control | Notable Action |
| 1 | 20 | 20 | Experiment set up, experimental Zebrafish placed in 20 mg/L caffinated aquatic environment, control Zebrafish placed in Deer Park water environment |
| 4 | 19 | 19 | Water in both experimental setups was exchanged for fress 20 mg/L caffinated water and Deer Park water. |
| 7 | 0 | 12 | All remaining control zebrafish fed paramecium; Dead experimental zebrafish preserved in paraformeldehyde; 3 control zebrafish euthanized and perserved in paraformeldehyde; Dear Park water refreshed in control group |
| 11 | 0 | 7 | Deer Park water refreshed in control group |
| 14 | 0 | 0 | All zebrafish haved died - no remaining fish in experiment. |
Table 1: This table shows the number of zebrafish observed in each sample throughout the fourteen day observation period, as well as any notable action taken on the aquatic environments.
| Caffeine Treated Zebrafish | ' | ' | Control Treated Zebrafish | ' | ' |
| Fish ID | Body Length (µm) | Eye Diameter (µm) | Fish ID | Body Length (µm) | Eye Diameter (µm) |
| 1 | 437.5 | 45 | 1 | 400 | 37.5 |
| 2 | 362.5 | 47.5 | 2 | 375 | 30 |
| 3 | 425 | 37.5 | 3 | 387.5 | 35 |
Table 2: This table shows the measurements collected from the euthanized and preserved zebrafish on day seven of observations. Measurements were collected on body length as well as eye diameter of each fish.
| ' | Average Body Length (µm) | Average Eye Diameter (µm) |
| Caffeine Treated Zebrafish | 408.3 | 43.3 |
| Control Treated Zebrafish | 387.5 | 34.2 |
Table 3: This table compares the average body length and eye diameter of the zebrafish samples from the caffeinated environment and the control group. When comparing the two, the zebrafish collected from the caffeinated environment had body lengths and average of 1.1 times that of the zebrafish from the control group. The zebrafish from the caffeinated environment also had an average eye diameter of 1.3 times that of the zebrafish from the control group.
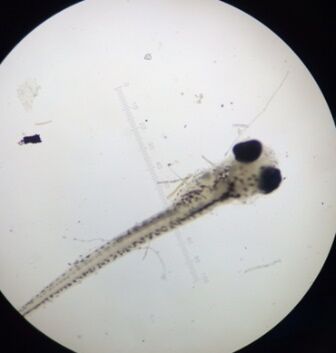
Figure 1: One zebrafish from control sample.
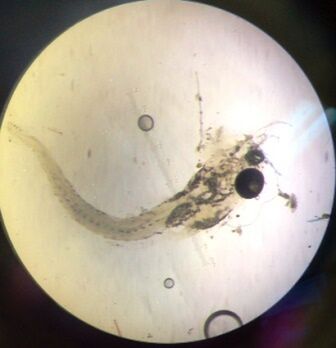
Figure 2: One zebrafish from caffeine treated sample.
Conclusion:
Overall it was found that features in the zebrafish treated in a caffeinated environment were slightly larger than those in the control group, and the average eye diameter was larger in the zebrafish treated with the experimental condition verses the control group. All zebrafish in the experimental group with treated with caffeine in their aquatic environment were dead by day seven of observations. All zebrafish in the control group were dead by day fourteen of observations. Our experiment disproved our hypothesis that zebrafish developing in a caffeinated environment would be smaller in size than zebrafish developing in the control environment.
March 2, 2015 - 16S Sequencing
Purpose
The purpose of sequencing the DNA of the bacterial colonies within Transect 5 is to determine the species of bacteria thriving in the manicured landscaping ecosystem on campus. This information provides us with a more complete analysis of the life in Transect 5.
Materials & Methods
A PCR reaction was run on four bacterial samples from Transect 5. Two samples were used from the nutrient agar plate at a dilution of 10E-7 from the Hay Infusion culture, and two samples were used from the nutrient agar + tetracycline plate at a dilution of 10E-7 from the Hay Infusion culture. The two samples from the nutrient agar plate were sent off to a lab for sequencing, and the resulting sequences were used in a database to identify the bacterial species that most closely matched the DNA from the selected samples.
Results
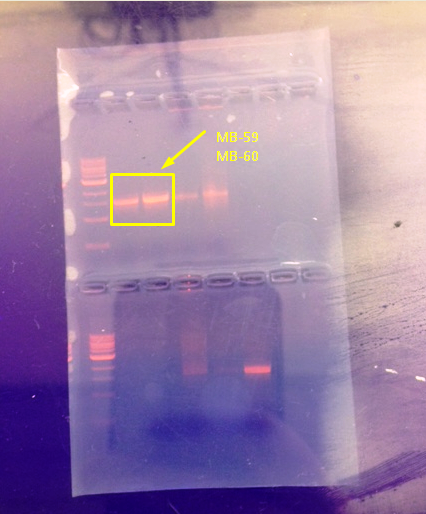
Figure 1: Aragose gel with products from PCR reaction showing the presence of 16S rRNA gene in Sample 1 and Sample 2 (MB_59 and MB_60) from nutrient agar plate at 1-E-7 dilution from the Transect 5 Hay Infusion culture.
MB-59: Nutrient Agar Plate 1 Raw Sequence (Forward)
NNNNNNNNNNNNNNNNNNAGCNNNGCAGTCGAGCGGANGANGGGAGCTTGCTCCTTGATTCAGCGGCGGACGGGTGAGTA
ATGCCTAGGAATCTGCCTGGTAGTGGGGGACAACGTTTCGAAAGGAACGCTAATACCGCATACGTCCTACGGGAGAAAGC
AGGGGACCTTCGGGCCTTGCGCTATCAGATGAGCCTAGGTCGGATTAGCTAGTTGGTGGGGTAATGGCTCACCAAGGCGA
CGATCCGTAACTGGTCTGAGAGGATGATCAGTCACACTGGAACTGAGACACGGTCCAGACTCCTACGGGAGGCAGCAGTG
GGGAATATTGGACAATGGGCGAAAGCCTGATCCAGCCATGCCGCGTGTGTGAAGAAGGTCTTCGGATTGTAAAGCACTTT
AAGTTGGGAGGAAGGGCAGTAAGCTAATACCTTGCTGTTTTGACGTTACCGACAGAATAAGCACCGGCTAACTCTGTGCC
ANCANCCGCGGTAATACAGAGGGTGCAAGCGTTAATCGGAATTACTGGGCGTAAAGCGCGCGTANNTGGTTTGTTNNGTT
NNATGTGAAANCCCCNGGGCTCAACCTGNGGAACTGCATCCCNNACTGGNNNNCTANANTACGGTGNANGGTGNTGNAAT
TTCNTGTGNNGNNNTGAAATGCNTAGATANANCATTAACAGCGGNGACNANGGGACNNCNTNNACTNGTTNGNNCCNCNN
ATGTGCGNNNNCNGGTGNNGNAAACANNNNNGAGNNCCNCCTTCGTCNNNGNNNNNNNNANNATNNNNCTACCCCNTTNN
NNNGNTNNNNGNANTTNCCNNGCCTNNCCNCNNGANNNNANCNNNNCGTTNNNNNAGCNCNCCNTNGNGNNNACNGGNNT
TTNNNAGNCNNNGNNNANNCAACNNNNNNAGNNNNNGTNGNNCNANTNNNNNNAAAANNNATNNNNNNNNCCNTNNNNTA
GNACGNNNGACGNNNNNNNNANNNNGNNNNN
MB-60: Nutrient Agar Plate 2 Raw Sequence
NNNNNNNNNNNGNNNGCTTNNNNTGCAGTCGAGCGGGGAGATGTAGCTTGCTACATTTCCTAGCGGCGGACGGGTGAGTA
ATGCTTAGGAATCTGCCTATTAGTGGGGGACAACGTTTCGAAAGGAACGCTAATACCGCATACGCCCTACGGGGGAAAGC
AGGGGATCTTCGGACCTTGCGCTAATAGATGAGCCTAAGTCAGATTAGCTAGTTGGTGGGGTAAAGGCCTACCAAGGCGA
CGATCTGTAGCGGGTCTGAGAGGATGATCCGCCACACTGGGACTGAGACACGGCCCAGACTCCTACGGGAGGCAGCAGTG
GGGAATATTGGACAATGGGGGGAACCCTGATCCAGCCATGCCGCGTGTGTGAAGAAGGCCTTTTGGTTGTAAAGCACTTT
AAGCGAGGAGGAGGCTACTAGTACTAATACTACTGGATAGTGGACGTTACTCGCAGAATAAGCACCGGCTAACTCTGTGC
CAGCAGCCGCGGTAATACAGAGGGTGCGAGCGTTAATCGGATTTACTGGGCGTAAAGCGTACGTAGGCGGCTTTTTAAGT
CGGATGTGAAATCCCTGAGCTTAACTTAGGANTTGCATTCGATACTGGGAAGCTAGAGTATGGGAGAGGATGGTAGAATT
CCAGGTGTAGCGGTGAAATGCGTAGAGATCTGGAGGAATACCNATGGCNAANGCGGCCATCTGGCCTGNTACTGACGCTN
ANGTACGAAANCATGGGGAGCANACAGGATTAGATACCCTGNTAGTCCATGCCNTANNCNATGTCTACTNNCCNTTGGGG
CCNNTGANNNNNTANTNNCNCAGCTCACGCNATAANTNNACCGCCTGGGNAGTGNNGNNCNCAGNNTANACTCAAATGNN
TTGACANNGNCCNGCACAANCGNTNGANCATGNGNNTTTNNNTCNATGCANNNNNANNNCCNTNACCTGCTNNTTGNNGN
TANNNNNNANCNNNNNNGGNANATNNNNTNGGNGGCGNTNGNTNANCTTNNNGNANANNTNNCNNCNTNGNNNNGNANGN
GCTNNNNGTCNNCNNNTGNNNGNNNANNNNNGGNNNAANNNNNNNNNNNNNNNNNNTNNNNNTNNNNNN
Species ID
MB-59: Nutrient Agar Plate 1 Raw Sequence: Pseudomonas putida
Pseudomonas putida is a rod-shaped, flagellated, gram-negative bacterium that is found in most soil and water habitats where there is oxygen ("Pseudomonas putida", 2013).
MB-60: Nutrient Agar Plate 2 Raw Sequence: Acinetobacter bouvetii
Acinetobacter cells are short, Gram-negative rods, measuring 1.0-1.5 by 1.5-2.5 microns during growth; they often become more coccoid during the stationary phase. Cells are found in pairs or small clusters; the groups form smooth, pale colonies on solid media ("Acinetobacter," 2010).
| Colony LabeL | Plate Type | Colony Description | Cell Description | Gram + or Gram - |
| Nutrient 1 | Nutrient Agar at 10E-7 dilution | yellow/round/shiny. Circular/Entire Edge | Non-motile, circular/coccus, clustered together/staphylocci | Negative. Viewed at 40x |
| Nutrient 2 | Nutrient Agar at 10E-7 dilution | Circular/matte/white-ish in color | Non-motile, circular/coccus, clustered together/staphylocci | Negative. Viewed at 40x |
Table 1: Colony and cell description of bacterial samples from nutrient agar plate, including gram +/- information.
Conclusion
The two species sequenced from Transect 5 were Pseudomonas putida and Acinetobacter bouvetii. Pseudomonas putida is rod-shaped, flagellated and gram negative. In our previous observation under the microscope, this bacteria was noted to be circular/coccus shaped and gram negative. The variance in description could be related to experimenter error in lack of familiarity with recognizing bacterial shapes under a microscope. Acinetobacter bouvetii are gram-negative rods that become more circular during the stationary phase ("Acinetobacter," 2010). They also form smooth, pale colonies. This aligns with our observations from the bacterial colony.
References Cited
Pseudomonas putida. (2013, January 15). Retrieved March 3, 2015, from https://microbewiki.kenyon.edu/index.php/Pseudomonas_putida
Acinetobacter. (2010, January 15). Retrieved March 3, 2015, from https://microbewiki.kenyon.edu/index.php/Acinetobacter
February 16, 2015 - Vertebrates
Purpose
The purpose of this experimental method is to collect information and identify vertebrate species in Transect 5, to complete an overall summary of life in this ecosystem on American University campus.
Materials & Methods
Observations were collected in Transect 5 in the late afternoon on a cold day in February. The temperature was well below freezing, at roughly 15 degrees Fahrenheit. It was a very windy day, and there was not a lot of activity on campus at this time.
Data & Observations
Conclusion
February 14, 2015 - Invertebrates
Purpose
The purpose of this lab is to explore the invertebrate species that live in transect 5, and discover their importance of their role in this ecosystem. We also explored out simple species evolved into species with more complex systems.
Materials & Methods
Last week, leaf litter was collected and placed in a Burlese Funnel underneath UV light to drive any invertebrate species into an ethanol solution to preserve them for observation. This week, sample species from different phyla of invertebrates were observed. First, samples of acoelomates, pseudocoelomates, and coelomates were examined to understand the different features of each species as it evolved. Next, five classes of the Arthropoda phylum were observed and identified to learn more about the basic difference across the classes of arthropod invertebrates. Lastly, the Burlese Funnel was carefully broken down and the organisms collected in the preservative solution were transferred into petri dishes for observation under a dissecting microscope. Five invertebrate species were observed and described in Table 1, and an invertebrate key was used to assist in identifying each species.
Data & Observations
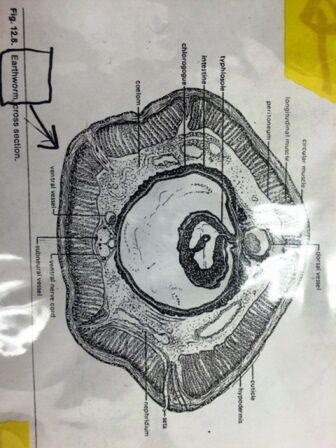
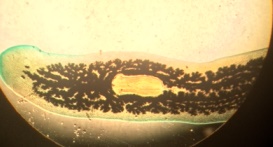
Figure 1 - 2: A diagram of the cross section of coelomates (Figure 1), and the observed cross section of a coelmate under a microscope (Figure 2).





Figure 3 - 8: Samples one through five of the observed species of 5 classes of arthropods: Insect, Diplopoda, Crustacean, Arachnida, and Chilopoda, respectively.
| Organism | Length (mm) | Number in Sample | Description of Organism |
| 1) Arthoropod, Insect | 3 | 1 | 3 legs on each side of thorax; has front antennas; brown in color with black spots on thorax; three sections of body |
| 2) Nematoda, Chromdaorea? | 0.5 | 1 | very tiny and thin; red in color; resembles a small string |
| 3) Arthoropod, Insect | 0.5 - 2 | 3 | 3 legs on each side of thorax; has front antennas; light purple in color with purple spots on thorax; three sections of body |
| 4) Arthoropod, Insect | 0.5 | 1 | winged; larg wings the size of entire body; brown with antennas; three sections of body |
| 5) Arthoropod, Insect | 1 | 1 | brown; wingless; structured similarly to Organism 3; dark brown; 3 sections to body |
Table 1: Descriptions of five invertebrate species collected from leaf litter in Transect 5. Four Arthropod species and one Nematode species were identified and examined.
Conclusion
We observed the cross sections of an acoelomate and coelomate underneath the microscope. The coelomate cross section (Figure 1 and 2) shows the fully lined, fluid-filled coelom. We also observed live Nemertea worms from the Nematode phylum. Their movement is very slithering; back and forth movement from the middle of their bodies to the ends.
We examined the different characteristics of arthropods through samples of five classes; insect, diplopoda, crustacean, arachnida, and chilopoda. In Figures 3 - 8 we used our knowledge of the characteristics of each of these classes to identify the five unknown samples of arthropods species.
We used the basics we learned in identifying the samples of arthropods to examine and identify some invertebrate species found in the leaf litter of Transect 5. We observed four insects, and one Nematoda worm species (possibly from the Chromadorea class). Table 1 shows their relative sizes. The largest species we observed was 3 millimeters, and the smallest invertebrate we observed was 0.5 millimeters; both were insect species. It appears that the majority of invertebrates living in Transect 5 are arthropods, and specifically insects. Given the constant activity that appears in Transect 5, through regular maintenance and foot traffic from humans, it seems that insects thrive in this ecosystem better than other invertebrates.
AS
February 11, 2015 - Plantae & Fungi
Purpose
The purpose of this laboratory procedure is to collect and examine the diversity of plant and fungi species in our transect and identify as much as possible based on our observations. We are specifically interested in vascularization, the presence of specialized structure, and the mechanisms of reproduction in our collected plant samples. We also collected leaf litter from transect 5 to prepare for next week's lab and the examination of the invertebrates living in this niche.
Materials & Methods
This week, five plant samples were collected from transect five and brought into the laboratory for observation. The layout of each species vascular bundles were examined and each vascularization was recorded in table 1. Next, the leaf of each plant was examined for special structures, and any notes around the cell specialization were recorded in table 1. Plant reproduction mechanisms in a lily flower were then observed and used to compare with the mechanisms of reproduction observed from the collected plant samples. Once the plant observations were recorded, samples of various fungi were observed and identified. Lastly, leaf litter was collected from transect 5, and a Berlese Funnel was set up using the litter to collect invertebrates for observation in next week's lab.
Data & Observations
| Transect Sample Plants | Location and # in Transect | Description | Vascularization | Specialized Structures | Mechanism of Reproduction | Identification of Plant |
| 1 | in soil - under the bushes | leaves are long, thin, and green; veins running parallel, 4 inch leaves | vascular bundles scattered | Root cells | monocot | ? |
| 2 | ground cover - weed growing next to stone path | some leaves bundled in groups of four; 1 inch leaves, broad vein network | network of veins; vascular bundles scattered; hairlike roots | hair like roots; leaves have hair like structures; broad vein network | ? | ? |
| 3 | Dead leaf from shrubs | Bush is large, flowering plant; 3 ft high; thorns | broad network of veins in leaf | flowers in summer as reproduction; roots; leaves | dicot | angiosperm |
| 4 | Bulb found in soil below bushes | circular; roots coming out of one end; white bulb; green plant growing out of one end | vascular bundles scattered | bulb structure; leaf bud out of bulb; roots | monocot | gymnosperm or angiosperm? |
| 5 | grass area in transect - grass | long thing green leaves; parallel veins; shallow roots; 2 inches tall per leaf | parallel veins | roots; flowering section of plant in summer | monocot | angiosperm |
Table 1: Description of plants collected from Transect 5

Figure 1: Transect sample plant #1 - bunch grass plant located underneath manicured bushes.

Figure 2: Transect sample plant #2 - small leafy ground covering plant growing next to stone pathway


Figure 3 & 4: Transect sample plant #4 - bulb found in soil below bushes

Figure 5: Transect sample plant #5 - grasses from manicured grass in transect
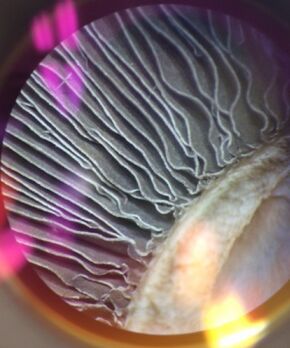
Figure 6: View of underside of mushroom - basidiomycota - to see spore-producing structures.
Conclusion
We collected 5 species of plants from transect 5 in this weeks lab. The plants were collected from a range of niches throughout the transect to increase the diversity of our samples. We collected samples from the soil under a manicured shrub, a sample from the shrub, some grass, and a bulb of a flowering plant in the landscaped bushes. We observed two angiosperms, one potential angiosperm or gymnosperm, and we were unable to identify two of the plants we observed. There were not may seeds to collect in transect 5, as the area is well groomed and maintained, and there are not seeds available during the winter months. The bulb, however, gave us an interesting look into the seed of the flowering plant.
We also observed fungi samples in the lab, noting their reproductive structures. In the samples of the sample of fungus from the Zygomycota phylum, we could see the sporangia structures. The sporangia is an enclosure that fungus creates to produce their spores for reproduction. In figure 6 above, the underside of a mushroom from the Basidiomycota phylum is pictured, showing it's reproduction function. The mushroom itself is the fruiting body of the fungus, and it's purpose is to produce and distribute fungus spores. There were no mushroom species in transect 5; however, there did appear to be some mold on the nutrient agar plate used to grow bacterial species from transect 5.
Lastly, we used the collected leaf litter to build a Burlese Funnel to collect invertebrate species that live in transect 5. We added the leaf litter to a funnel, and placed the funnel inside an ethanol preservative. We moved the funnel under UV lighting to encourage any invertebrate species in the leaf litter to move down the funnel, away from the light, and fall into the preservative for next week. I would expect to observe some invertebrates species in the next lab.
AS
February 4, 2015 - Microbiology & Identifying Bacteria with DNA Sequences
Purpose
In this lab, we observed some of the bacterial life from transect 5 and used the context of cell and colony morphology to make classifications about the bacteria species present. We also observed some antibiotic resistance from samples from transect 5, as we observed growth of bacteria colonies on agar nutrient + tetracycline plates, and set up four samples of bacteria colonies for a DNA PCR reaction to amplify the 16s rRNA gene. If the 16s rRNA sequence can be amplified in any of the four bacteria samples, the DNA from these bacteria will be sent for DNA sequencing, and we will learn the identification of these bacteria species.
Materials & Methods
In lab this week, my team observed the growth of bacteria on the agar nutrient and agar nutrient + tetracycline plates that were brushed with samples from the serial dilutions of the sector 5 Hay Infusion Culture. In the previous week, on January 29, 2015, four serial dilutions of the Hay Infusion Culture were created in steps. First the Hay Infusion Culture was mixed thoroughly, and 100 microliters of the solution were micropippeted into 10 mLs of sterile broth to create a [math]\displaystyle{ 10*10^-2 }[/math] dilution. 100 microliters of the [math]\displaystyle{ 10*10^-2 }[/math] solution were micropippeted into 10 mLs of sterile broth to create a [math]\displaystyle{ 10*10^-4 }[/math] solution. This process was repeated to create [math]\displaystyle{ 10*10^-6 }[/math] and [math]\displaystyle{ 10*10^-8 }[/math] serial dilutions of the Hay Infusion Culture. Once this was complete, 100 microliter samples from each of these serial dilutions were added to agar nutrient and agar nutrient + tetracycline plates, and the plates were covered and labeled with the corresponding dilutions of [math]\displaystyle{ 10*10^-3 }[/math], [math]\displaystyle{ 10*10^-5 }[/math], [math]\displaystyle{ 10*10^-7 }[/math], and [math]\displaystyle{ 10*10^-9 }[/math]. The agar plate sample were allowed to grow for one week, and the bacteria colonies were then observed, counted, and recorded.
To observed the bacteria samples under the microscope, wet mounts and gram stains were created. The wet mounts were created by using a sterilized loop to scrape a tiny amount of grown from four colonies onto slides (two from the agar nutrient plates and two from the agar nutrient + tetracycline plates). These slides were observed through a microscope. The gram stains were created by using a sterilized loop to scrape a tiny amount of growth from the same four colonies and mix the samples in a tiny drop of water on the slides (two from the agar nutrient plates and two from the agar nutrient + tetracycline plates). The water on the slides was then dried over a flame and crystal violet dye was added to the slide samples and allowed to sit for 1 minute. The slides were then rinsed with water, covered with iodine for 1 minute, and then rinsed with water again. 95% alcohol was poured onto the slides for 10 seconds and then rinsed with water. Lastly, the slides were covered with safranin stain for 30 seconds and then rinsed with water. Excess water was blotted and the slides were then ready to be observed under a microscope.
Once all observations were made from the bacteria colony samples, a PCR setup procedure was followed to prepare for a future lab. Samples from each colony were transferred into 100 microliters of water in a sterile tube, and icupabated at 100 degrees Celsius for 10 minutes. The four samples were then centrifuged for 5 minutes at 13,400 rpm. Once that was complete, 5 microliters of supernatant from each samples was added to a primer/water/PCR bead mixture in a PCR tube. The tubes were then added to the PCR machine and the products will be run in the next lab.
Data & Observations
Final Notes on Hay Infusion Culture:
jar smells very moldy; more than previous week
more growth of fungus looking hyphae on top from last week
some water has evaporated
some mold growing on jar wall
| Dilution | Agar Type | Colonies Counted | Conversion Factor | Colonies/mL |
| 10-3 | nutrient | Lawn | x10-3 | immeasurable |
| 10-5 | nutrient | 1500 | x10-5 | 1.50E+08 |
| 10-7 | nutrient | 24 | x10-7 | 2.40E+08 |
| 10-9 | nutrient | 0 | x10-9 | 0.00E+00 |
| 10-3 | nutrient + tet | 480 | x10-3 | 4.80E+05 |
| 10-5 | nutrient + tet | 0 | x10-5 | 0.00E+00 |
| 10-7 | nutrient + tet | 0 | x10-7 | 0.00E+00 |
| 10-9 | nutrient + tet | 0 | x10-9 | 0.00E+00 |
Table 1: 100-Fold Serial Dilutions Results - This table shows the range of growth on the eight agar plates inoculated from the Hay Infusion Culture from transect 5.
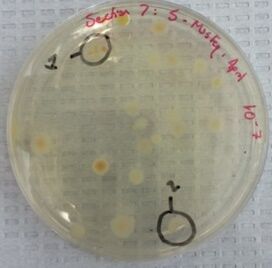
Image 1: A photo of the nutrient agar plate at 10E-7 dilution. The circled colonies 1 and 2 were used to observe cell morphology through the microscope.
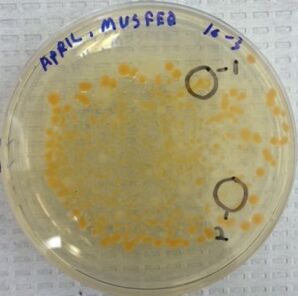
Image 2: A photo of the nutrient agar + tetracycline plate at 10E-3 dilution. The circled colonies 1 and 2 wereused to observe cell morphology through the microscope.
| Colony LabeL | Plate Type | Colony Description | Cell Description | Gram + or Gram - |
| Nutrient 1 | Nutrient Agar at 10E-7 dilution | yellow/round/shiny. Circular/Entire Edge | Non-motile, circular/coccus, clustered together/staphylocci | Negative. Viewed at 40x |
| Nutrient 2 | Nutrient Agar at 10E-7 dilution | Circular/matte/white-ish in color | Non-motile, circular/coccus, clustered together/staphylocci | Negative. Viewed at 40x |
| Tet 1 | Nutrient + Tet at 10E-3 dilution | Flat/Circular/Entire Edge, yellow/Orange in color | spiral shaped/all clustered together | Negative. Viewed at 40x |
| Tet 2 | Nutrient + Tet at 10E-3 dilution | Irregular edge/white-ish in color | circular/coccus | Positive. Viewed at 40x |
Table 2: Bacteria Characterization - This table describes the colony and cell morphology of four selected bacteria colonies from the samples; two from the nutrient plates and two from the nutrient + tetracyline plates.
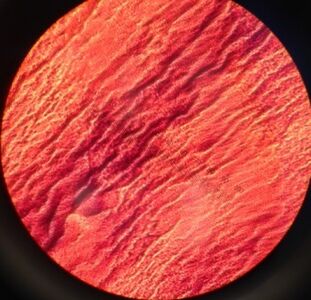
Image 3: A look at the "Tet 1" bacteria colony at 40x
Image 4: A look at the "Tet 2" bacteria colony at 40x
Conclusion
In this lab we learned about prokaryotes grouped into the domains Bacteria and Archaea. I did not anticipate seeing any archaea on the agar plates from the Hay Infusion Culture of transect 5, as archaea live in very extreme environments and would would not favor the environment at transect 5 at standard pressure. We were, however, able to study bacteria significantly in this lab, as there was a lot of bacteria colony growth on the agar plates from last week. Before observing the agar plates, we noted the changes in our Hay Infusion jar when compared to last week. The strong smell of mold/decay seemed to grow stronger from the previous week, and there was significantly more mold growth in the jar. This could be because different species are thriving as certain resources are used up in the contained jar environment, and the mold is now thriving on the decaying matter in the Hay Infusion.
As we moved onto observe the growth of bacteria on the agar plates, we noted that both the nutrient plates and nutrient + tet plate had fungi growth, although not a significant amount to interfere with the study. As noted in Table 1, there was significantly more bacteria growth in the nutrient agar plates than the nutrient agar + tet plates. The only growth on the nutrient agar + tet plates was on the 10E-3 culture dilution; the rest were empty. There were two distinctive colored colonies on the nutrient + tet plate; yellow-ish orange and creamy white. There were far more yellow-ish colonies on the nutrient +tet plate than cream/white colored colonies, as seen in Image 2. From this observation, it looks like there were two types of bacteria that are resistant to this antibiotic in the Hay Infusion culture from transect 5.
Under closer inspection using a microscope, we noted some observations and were able to capture some images of two samples from the nutrient + tet plate (Image 3 and 4). Three out of the four bacteria we inspected were gram negative; however, one of the bacteria resistant to the antibiotic was gram positive. This supports the method of action that tetracycline can traverse one or more membranes to enter the bacteria cell through an energy dependent process driven by the bacteria cell's method of maintain a proper pH gradient with it's surroundings (Chopra, 2001).
Overall, the study of bacteria in transect 5 was an interesting way to observe the variance in species across different areas of the transect. We observed antibiotic resistance in both gram positive and gram negative species. It appears that only two species thrived in the presences of the antibiotic when compared to the wide array of growth on the nutrient plates.
Works Cited
Chopra, I., Roberts, M. 2001, June. Tetracycline Antibiotics: Mode of Action, Applications, Molecular Biology, and Epidemiology of Bacterial Resistance. Microbiol Mol Biol Rev. 65(2): 232–260.
AS
January 29, 2015 - Identifying Algae and Protists in Transect 5
Purpose The purpose of this week's experimental method is to use a dichotomous key to identify the protists and algae in the hay infusion from the samples of soil and vegetation in Transect 5. One sample of material was collected from the bottom of the hay infusion culture, near the soil, and one was taking from the middle of the culture, next to roots of a plant. I make the prediction that we will observe more protists and life from the sample taken near the soil, than the sample taken in the middle of the culture near roots of a plant.
Materials & Methods
Last week, 11.7 grams of the sample of soil and vegetation from Transect 5 were added to a jar with 500mLs water and 0.1g dried milk. This jar was left uncovered for one week to be observed.The growth that appeared in the jar of the hay infusion from samples from Transect 5 were observed, and samples were collected from two niche areas of the sample; one near the bottom of the jar by the soil, and one near the middle of the jar, near roots. The samples were observed under a microscope to identify protists that were living in the hay infusion, and thus, in Transect 5. Once the observations were recorded, serial dilutions of the remaining sample in the jar from Transect 5 were plated on agar plates to observe the growth of prokaryotic organisms and fungi next week.
Data & Observations Description of Hay Infusion Culture There was a strong, bitter, pungent smell coming from the jar. It smelled moldy, and was very strong. There appears to be a mold or fungus on the top off the culture, across the surface of the water.
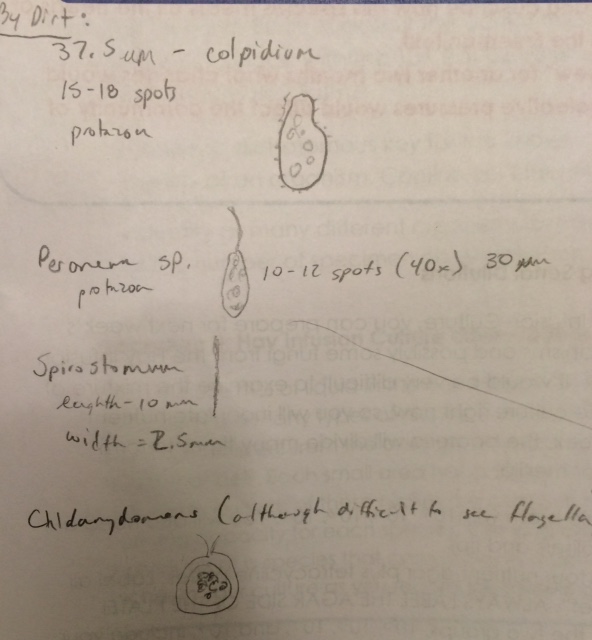
Observed Organism in Sample by Soil: Sample 1
Colpidium
37 micrometers
Motile - cilia
Peroneam
~30 micrometers
Motile - flagella...difficult to see
Spirastomum
Length:10 micrometers
Motile - difficult to see flagella or cilia
Chldamydomous:
Size was unattainable...
Motile - Difficult to see flagella
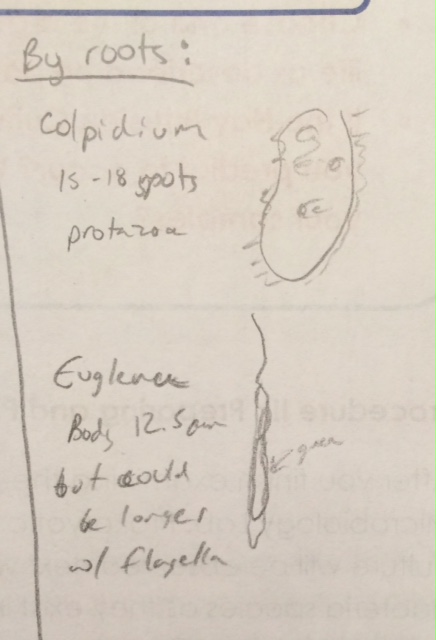
Observed near Roots: Sample 2
Colpidium
37-45 micrometers
Motile - cilia
Euglena
Body Length: 12.5 micrometers (but could be longer with flagella. Flagella was difficult to observe)
Spirastomum
Length: ~10 micrometers
Motile - difficult to see flagella or cilia
AS
January 29, 2015 - Biological Life at AU: Characteristics of Transect 5
Purpose This procedure was conducted as a means to discover Transect 5 at American University and make observations about the characteristics of the small section of land. Both biotic and abiotic materials were collected from the transect, for use in a hay infusion to later determine some of the living organisms in plot 5.
Materials & Methods
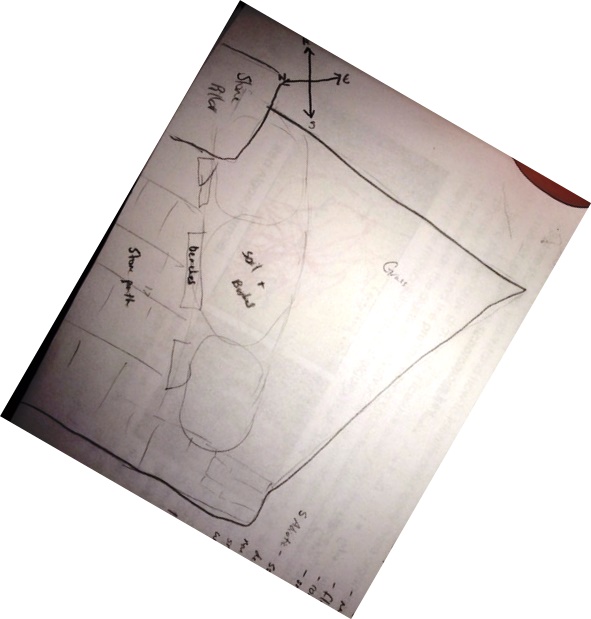
Once assigned to Transect 5 on American University's campus, the section of land was observed, diagrammed, and biotic and abiotic samples were collected from it and brought into the lab. 11.7 grams of the sample of soil and vegetation from Transect 5 were added to a jar with 500mLs water and 0.1g dried milk. This jar was left uncovered for one week to be observed in the next lab.
Data & Observations
General Observations: There is a lot of man-made materials in Transect 5; a stone hedge, stone path, and concrete benches. There is also manicured grass and maintained bush landscaping. While the grass is green, the landscaped bushes have fewer leaves at this time of the year.
Biotic Species:
Grass clumps,
manicured grass,
flower bush,
round berry plant,
small ground cover/weeds
Abiotic Species:
soil,
snow,
concrete bench/column,
wood chips,
stone path
Conclusion & Predictions
This lab was a useful introduction to Transect 5. I predict that as the Spring weather approaches the landscaping in this transect will blossom and change the nature of plot of land. I predict the hay infusion will produce a lot of bacteria and fungus, and we will be able to observe life in the next lab, given that the soil and vegetation was brought into the warm laboratory and fed through the dried milk.
AS
January 21, 2015 - Test Entry
Breakfast foods are the best foods.
AS