User:Anthony Salvagno/Notebook/Research/2009/09/22/PCR Cleanup and BstXI digestion
I hope this doesn't ruin the reaction, but I realize today based on Koch and previous notes that I used way too much pRL574 template DNA. In doing so I actually used up all the pRL574. Koch said he had glycerol stocks so I'll resupply us after talking with him, but I can't believe I didn't read thoroughly enough. I even saw the 1:100 dilution tube and thought "What could this be for?"
Cleanup
I wonder if I should run a gel of each tube before cleanup and while that is going clean the PCR reactions or just clean it and stuff. I'm just gonna do a nanodrop first. I have to do a gel later when I digest with BstXI so I'll wait for that.
NanoDrop
Tube# Reading
- 963.1ng/ul
- 966.7
- 958.1
- 971.3
- 961.1
I initially nanodropped all the samples one after another and each sample had high and higher concentration. Then I tried cleaning the pedestal between samples and got the results above. I used to clean the pedestal between samples, but then I read in the manual that that wasn't necessary. I guess it turns out that it is.
PS: PCR cleanup to come after class because I have homework to do. Finished early so cleanup commences.
Post Cleanup
I ran the nanodrop and got 1ug/ul in 30ul which is high for a column that can supposedly only hold 10ug of DNA. I will run a gel to try and distinguish the concentration.
Nanodrop Gel
Lane# ... Contents
- 1kb ladder
- 1:10 ladder
- 1:100 ladder
- 2x ladder
- .5x ladder
- anchor
- 1:10x anchor
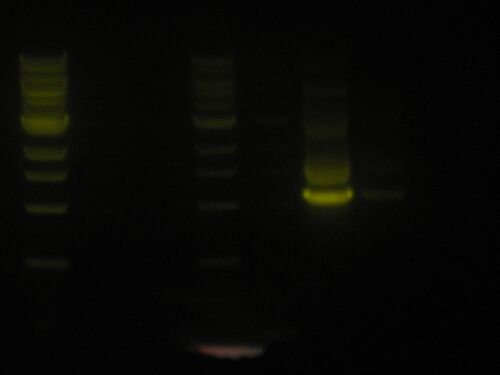
Commentary
It is a little blurred, but the picture will serve its purpose. I put way too much pRL574 in and this is what I reap. I will redo the PCR reaction tomorrow and after digestion with BstXI I will run a gel of this and that and gel extract the two. I can then choose to digest this PCR reaction and gel extract that. Although I don't know if the amount of DNA I will get from this will be worth it.
Andy helped me with some profile intensity (courtesy of ImageJ) and I found that my PCR product is roughly 4x as concentrated as the 2x ladder lane based on intensity. See here:
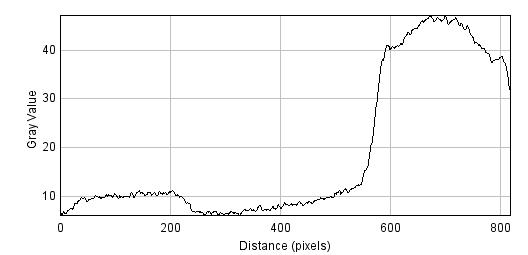
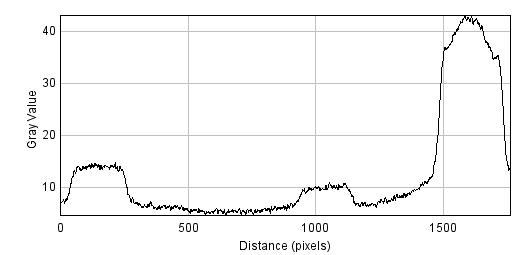
The bottom image is actually what I wanted. I realized looking at my gel image that the supposed 2x lane is not as bright as the 1x lane so I will have to go off that. According to intensity plot, the slice is 3x as bright so 3x as much DNA.
Well based on this new analysis I will just redo the PCR and not worry about today's product. 3x as much weight is only 132ng of DNA which is a tiny amount after gel extraction and then even less after digestion.