Talk:20.109(S07): Start-up expression engineering
From OpenWetWare
Jump to navigationJump to search
On Day 1 of Module 3, you did a few things:
- Learned about the SAGA chromatin-remodeling complex and chose a non-essential sub-unit of the SAGA complex to delete
- Designed primers to amplify the URA3 sequence with flanking sequence from upstream and downstream of the sub-unit chosen for deletion (On Day 2: This DNA sequence will be able to get into the nucleus of the yeast cells we are transforming after we have made them "competent" for DNA uptake. Once in the nucleus, the flanking sequences with homology to upstream and downstream sequence of your SAGA sub-unit will allow your DNA product from PCR to integrate into the yeast genome, via homologous recombination, kicking out your SAGA sub-unit sequence and replacing it with URA3 sequence)
- Set up PCRs with those primers to amplify the URA3 sequence off of the pRS406 plasmid DNA. I ran your PCR products out on a 1% agarose gel. The key for the lane loadings are below, so check out your results!
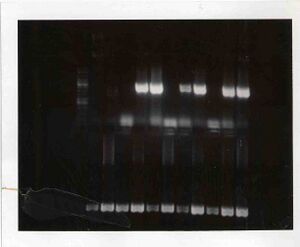
Gel lane loading order
Top row, Tue/Thur Section Samples
- empty
- ladder (100bp)
- purple team 1
- purple team 2
- blue team -template, good
- blue team +template, good
- red team +template, good
- red team -template, good
- pink team -template, contaminated (?)
- pink team +template, good
- green team -template, good
- green team +template, good
- yellow team +template, good
- empty
Bottom row, Wed/Fri Section Samples
- empty
- ladder (???)
- green team -template, contaminated
- green team +template, good
- red team -template, contaminated
- red team +template, good
- purple team -template, contaminated
- purple team +template, good
- pink team -template, contaminated
- pink team +template, good
- yellow team -template, contaminated
- yellow team +template, good
- blue team +template, good
It seems like there was probably some contamination of the PCR master mix we used with template DNA (pRS406) before the Wed/Fri section set up their reactions. From looking at the gel image, why do you think I say this? Also, can you tell from the gel if the contamination issue affected the Tue/Thurs section at all?