Lidstrom:Sequencing with GeneWiz
From OpenWetWare
Jump to navigationJump to search
Back to Protocols
Price per rxn: $6 JM 10/2012
- This is the standard UW price, but volume discounts are possible if our lab starts using more.
SUBMIT TO UW By REQUEST Box A
- In summer of 2013 we got a GeneWiz box outside our lab. To use this box, select "UW Box A," the by-request box, so they know to pick up our samples. They don't come up to our box if nobody submitted a sample.
Prices
Current as of 12/8/2014 JM
- Premixed
- 1-47 samples: $6.00/reaction
- 48 or more samples in a 96-well plate: $5.00/reaction
- Predefined
- 1-47 samples: $7.00/reaction
- 48 or more samples in a 96-well plate: $6.00/reaction
- Custom (includes colonies)
- 1-47 samples: $10.00/reaction
- 48 or more samples in a 96-well plate: $9.00/reaction
- Note: "You can get this $1 discount for bacterial colonies by either suspending them in liquid in a 96-well plate OR just submitting regular plates. "That is the only exception that I am aware of for bulk sequencing orders that aren’t in a 96-well format." -- Tech support 12/8/2014 JM
Preparing Samples
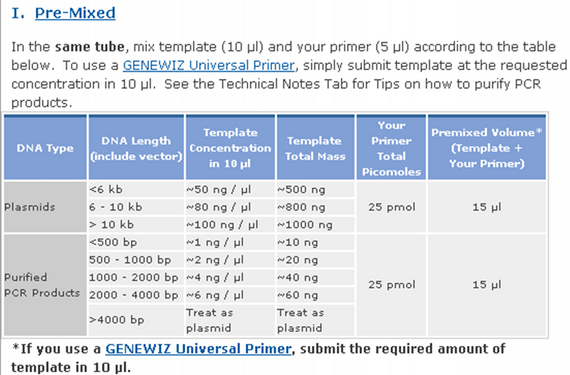
- 15 uL total including 5 uL of 5 uM primers.
- Keep a fresh stock of 5 uM primers that you take good care of (keep clean) and replace often with fresh stock.
- Red Flags:
- Low concentration minipreps (<70 ng/uL) frequently fail when submitted to sequencing. It may be due to impurities that co-elute; if you submit many uL of this to meet the 500 ng/sample requirement, then you will have more of these impurities present. -Janet 3/20/2012
Sequencing Un-Purified PCR products
- Submit 10 uL of PCR product and 5 uL of 5uM primer in a separate tube.
- Best to submit a printed picture of the gel, too.
- Submitting un-purified product costs ~$3/sample extra.
- pmol/uL = uM
Sequencing Colonies
- This option takes an extra day, and is more likely to give poor results relative to PCR products and minipreps. --J. Matsen 12 December 2014
- Instead, I recommend doing PCR products. Ideally the PCR primers are outside the priming region for the sequencing primer you submit.
- You can submit colonies that are randomly oriented on a plate (e.g. from a transform), or colonies streaked onto a grid.
- Either way, circle and number the samples.
- Make the name in the GeneWiz form and the circle description on the plate clear to match up.
- Submit 5uM primer in a separate tube and a plate with colonies.
- For each reaction you will want 5 µl of your primer at 5 pmol/µl (5 µM) in a labeled tube.
- GeneWiz keeps plates around for 5 days, so you can submit a different primer to be used on the same original plate a subsequent day.
- Submitting costs $4/sample extra relative to minipreps.
Order Submission Deadline
- The cutoff time for sequencing is 4PM.
- You can submit samples to the box a little after 4PM as long as you electronically submitted your order by 4PM. We are one of the last pickup sites so it takes a while for the person to get to our box.
Dropping of your samples to the Lab
If you miss the deadline, you can drop off samples at 1551 Eastlake Avenue E. They are on the first floor.
When Sequencing Fails
- There are several ways it can fail & you can see the GeneWiz help for causes/solutions below. You can also watch this GeneWiz webinar. Don't hesitate to call GeneWiz customer support -- their customer service is incredible.
- GeneWiz has very high standards for passing sequencing reactions. About 1/2 of your attempts may fail. Sometimes you can look at the trace file and decide it is good enough on your own despite a failed label.
- multiple are present on your DNA or you accidentally have a mixture of DNA templates.
- You can prepare a new miniprep, with the hope that your first one was contaminated with another plasmid. Otherwise, you have to assume the plasmid you built accidentally contains an extra priming site. Of course, miniprepping from single colonies will reduce the possibility of mixed templates.
- Note: a plasmid can contain an extra priming site via pcr/assembly fluke OR the insert you add can contain a mispriming site.
- hairpin present
- The clone has multiple priming sites (can be on accident)
- multiple are present on your DNA or you accidentally have a mixture of DNA templates.
- high background: there is a clear sequence but there is also a background sequence.
- remnants of chromosomal DNA
- primer degradation: recall that your sequencing is dependent on length. So if you have one population of primers that is 20 bp and one that is 18 bp, they will give two separate signals.
- top heavy: beginning sequence is strong; peaks may even be cut off. As it gets longer, the signal dies off. This is due to improper primer:template ratio; your primer may be "used up".
- lots of "N"s in sequencing result
- too much, too little, or no DNA
- no primer or sub-optimal primer concentration
- primer site not complementary
- carry-over inhibitors in the reaction such as phenol, ethanol, EDTA, or salts
- hairpin present, repetitive sequence, or high GC content can terminate the rxn sequencing. Note: they have several "difficult sequencing" protocols that can be used; select them in the form.
- too much DNA
- bubble in capillary
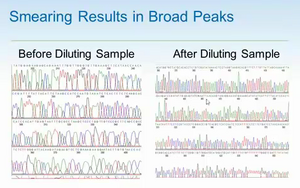
- can appear as a smear. You won't know when this happens, but customer support may be able to tell.
- dye blobs
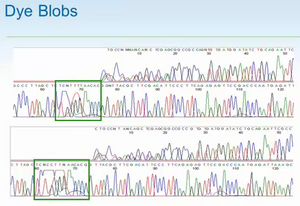
- too many terminating nucleotides remain after their purification -- see image on right. A big peak covers the sequence you want. They generally have it at the beginning, so you should start your priming ~ 100 bp away from the sequence of interest. They happen on PCR products and plasmid & are best visually edited (ignored.)
- what is the difference between High Background & non-specific?
- Customer support says: the degree to which the background sequence
- you can trust a sequence that fails because of high background more than you can non-specific
When it fails, try again!
- You can submit 2 samples per order for a free repeat, but they must be on the same templates you submitted.
- You can resequence as many as you like for 1/2 price and you have the choice to use new preps. This allows you to use a fresh miniprep or primer batch. If you want to submit fresh samples for 1/2 price, you should contact customer service and get approval. Write in the comments section at the bottom which samples are 1/2 price repeats, and which row & order the original failed sequence was in. Then call them with the order number for this new request/order. Janet 7/18/2012
- Whether or not your repeat will be successful depends on several variables such as:
- the concentration and quality of the sample -- if a "no priming" or "poor quality" samples is "no priming" or "poor quality" after you sequence again, the quality and quantity might be too bad to work the 2nd time.
- you also may have a GC-rich region, hairpin or general difficult template that is causing binding issues, a problem with the binding site, a problem with the primer, etc.
Vague primer design instructions
- "We recommend designing your sequencing primers in a region that is 100 bases upstream of your sequence of interest. If you do not have the luxury of having this buffer, the closest you want the primer to be to your area of interest is 50-60 bases. Anything closer and you risk missing a portion of your area of interest. The primers should be about 18-24 bases in length with a Tm of 56-60 degrees. The GC content should be about 45-55%. Many vendors that provide oligo synthesis services have software into which you can plug your primer sequence to check for Tm, GC content and homodimerization. For the oligo purity, desalting is all that is needed for sequencing" (link)
- Note that Finnzymes reports Tm higher than other calculators, so 62-65oC is probably good if using a Finnzyme calculation.
Sequencing un-purified PCR products
- send 10uL of PCR product and 5+uL of 5uM primer (separately).
- chose "custom" for the type of sequencing reaction
- you can leave the DNA concentration field blank
- you can't do nano drop on DNA that hasn't been cleaned up (says customer service 1/2014) so you would have to compare it to a known concentration of ladder anyway.
- pmol/uL [=] uM
- include a photo of your gel in with the sequencing form if available (not necessary).
- It costs ~$3 more per reaction.
- You can submit OneTaq 2X products, even though it has loading dye in it.
- [1][Genewiz instructions]: see III Custom (at the bottom).
Sequencing gel purified PCR products
Organic solvents inhibit/inactivate the Taq used in Sanger sequencing.
Tip sent by technical support to Janet 5/2013:
- Tips for Column Purification of Template DNA
- Here are a few guidelines to help aid in the effective column purification of your product.
- This will help to eliminate carry over of ethanol into the eluate.
- Prior to elution, there is a wash step that includes a high concentration of ethanol. If you are using a micro spin column, after the wash step, please spin samples at maximum speed in the microfuge 2 times for 2 minutes decanting the flow through completely after the first spin and placing the column in a fresh tube for elution after the second spin.
- If there is a rim around the membrane, carefully remove residual ethanol using a pipette tip after the second spin and then let the column sit for 2 minutes on the bench top prior to elution in order to promote residual ethanol evaporation.
- Carefully elute with EB or water heated to 50'C by adding drop wise to the membrane surface. (I usually let this sit a couple min before spinning -ALS)
- Spin at maximum speed for two minutes.
- Please check your 260/280 and 260/230 ratios prior to sending samples for sequencing.
- The 260/230 ratio is very important since small deviations reflect contamination with organic solvents that essentially deactivate the taq and preclude informative sequencing results.