LauraTerada Individual Journal Assignment Week 5
From OpenWetWare
Jump to navigationJump to search
Exploring Pathway Databases
SGD
- Which of these genes has a homolog (similar gene related by descent) in humans? What disease does a deficiency of this gene cause in humans?
- The GDH2 gene has a homolog in humans (GLUD1 and GLUD1), and deficiencies of these genes are linked to hyperinsulinism-hyperammonemia syndrome and several other neurological disorders.
- How is the expression of each of these genes regulated?
- GDH1 is levels are high by both ethanol and glucose, two carbon sources. GDH1 is regulated under fermentative growth conditions to undergo glutamate biosynthesis.
- GDH3 is regulated by ethanol and glucose concentrations. Ethanol induces GDH3 expression; however, glucose represses it. GDH3 is is required to balance levels of alpha-ketoglutarate, glutamate, and cell metabolism.
- GDH2 expression is regulated transcriptionally by elements that act as upstream activation sites and as upstream repression sites. GDH2 is also regulated under nitrogen concentrations; when nitrogen (glutamate) is high, transcription of GDH2 is repressed. Moreover, in situations of high ammonia intracellular concentration, GDH2 expression decreases. When extracellular ammonia concentrations increase, GDH2 transcription level decreases.
- GLN1 codes for glutamine synthetase, and it is regulated by ammonia concentration.
- GLT1 gene expression is regulated by glutamate repression and GLN3 activation. This regulation is ultimately dependent on nitrogen and glutamate concentration. With low amounts of amino acid, GLT1 expression is also regulated by GCN4.
- Using the compound search tool of SGD, search on "L-glutamate". How many pathways does it participate in?
- As either reactant or product: 56
- In reactions not assigned to pathways: 12
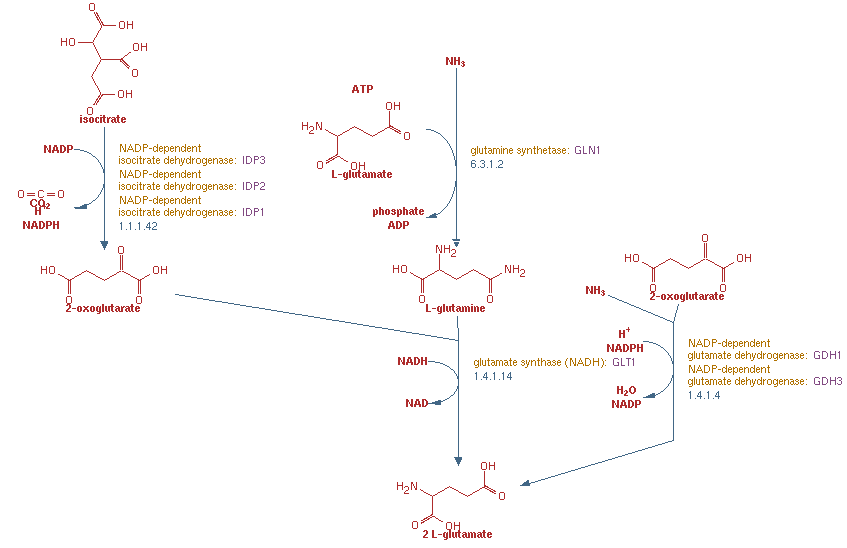
- Find the SGD representation of the pathway we are working on in class and attach a screenshot and hyperlink to your journal page. Choose the one that shows all of the reactions we talked about in class and make sure you can relate it to your notes, matching the genes, enzyme names, and reactants/products.
- What parameters for these reactions can you find using this database? HINT: the literature portion of the individual gene pages may be helpful.
- There is a large amount of literature on the database in regards to the pathways of Saccharomyces cerevisiae, and these studies can tell us about a variety of parameters. Such parameters include the conversion rate, nitrogen feed, and carbon feed.
KEGG
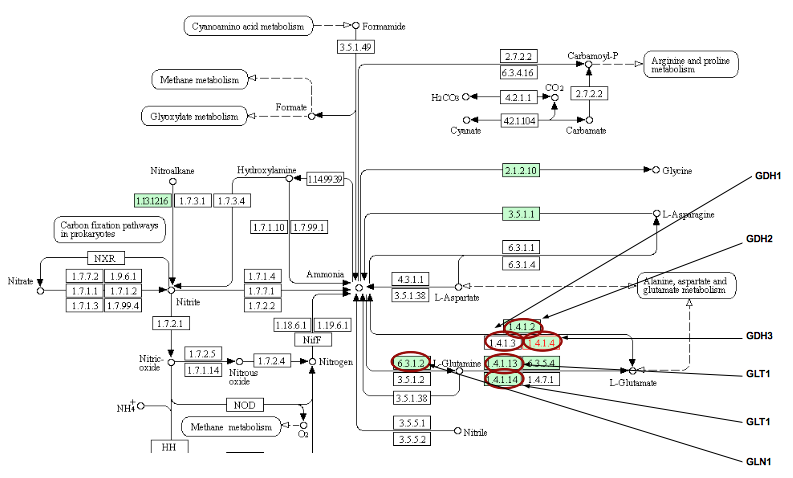
- KEGG also uses a system where a "master" summary pathway compiled from many different organisms is then highlighted with the organism-specific enzymes/genes. How many genes in this pathway exist in yeast?
- When looking at the entire pathway, called Nitrogen Metabolism, there are 9 green colored boxes. These 9 boxes indicate the enzymes/genes that are in both different organisms and in yeast.
- Click on each of the five enzymes of our pathway and read the individual enzyme pages. Is there any new information here that was not represented by SGD?
- Yes, KEGG shows both the amino acid and nucleotide sequence. It also had a section for the different pathways the gene was in and a section for class.
Reactome
- Reactome did not have the complete pathway on its database, but I did find one reaction of the pathway we have been discussing in class.