Karas Lab:Fragility Module
Fragility Processing Pipeline
Preface
First things first: if you don't have RAVE installed, follow these steps to do so: RAVE:Install Next, you'll have to pre-process your patient for RAVE. Tutorials for that are available here: RAVE:ravetutorials
Pre-Processing the Patient for Fragility
Step 1 on the left-hand side is to process the patient for the fragility module. I know you just did a bunch of pre-processing for the RAVE subject, but don't worry - this is very fast and is just to standardize and save some metadata in the patient files. There are two options to look at before you press the "Pre-process Patient" button: "Units of Recording", and "Halve Sampling Rate?"
- Units of Recording: The Fragility module defaults to working in units of microvolts (uV), so if your raw voltage traces are in units of millivolts (mV) or nanovolts (nV), this option is to convert them to microvolts before continuing to the next steps.
- Halve Sampling Rate: If your data is recorded at a high sampling rate (e.g. 2000 Hz), you have the option to halve it here to save on processing time. The fragility metric does not require extremely high temporal sensitivity so it is possible to halve the sampling rate for fewer time windows and lower processing time.
After you have set these options and clicked on the "Pre-process Patient" button, a [SubjectCode]_pt_info file will be generated inside your patient's module_data folder. This folder can be found within the patient's data folder (located in rave_data/data_dir/[ProjectName]/[SubjectCode]/rave). For future analysis of this patient, this pt_info file will be automatically loaded so that this step does not have to be run every time.
Generating the Adjacency Array and Fragility Matrix
Step 2 on the left-hand side is to generate the Adjacency Array. This is the time-varying linear model of the EEG data that will be used to determine the neural fragility of each electrode in the patient. It is generated by using a least-squares algorithm to linearly fit the raw EEG data. The time-varying aspect of this linear model comes from the fact that the entire time course is split into many smaller time windows, and a separate model is generated for each time window. The adjacency array generation process can be thought of as a "sliding time window" process, where the first time window model is generated before "sliding" over by a certain time step and moving on to the next time window. There are several input options:
- The size (in ms) of each time window. A smaller time window would mean a more accurate model (and surprisingly faster processing time too). Default is a time window of 100 ms.
- The size (in ms) of each time step. This refers to how far the "sliding time window" moves with each iteration; in other words, whether you want the sequential time windows to overlap or not. If the time step size is equal to the time window size, sequential time windows will not overlap. If the time step size is half of the time window size, each time window will be offset from the previous one by half a time window. Having overlapped time windows results in a more accurate model but slower processing times. See the figure below for an illustration of this concept.
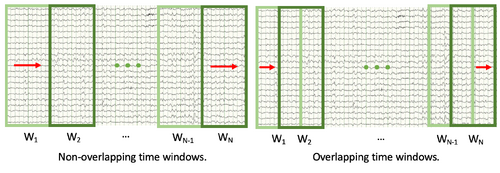
- Number of Lambdas. This refers to how many lambdas will be tested within the Ridge regression, which is the algorithm used to find a least-squares fit of the EEG data. More lambdas will result in a more accurate model but also longer processing times.
- Number of cores. How many cores to use on your computer for parallel processing. Each core will be used to test a single lambda value in parallel, so there is not much benefit to using more cores than number of lambdas.
- Seizure Trial for Matrix Generation. If your patient has multiple trials (i.e. multiple EEG recordings), this is where you choose which recording to generate the time-varying linear model for.
Clicking the "Generate Adjacency Array" button will load a pop-up with more information on the data you are about to process. Warning: this is the most computationally intensive step and may take a long time (many hours or even multiple days!) to run. This is because a new model is generated for each time window, and there are usually hundreds of time windows. Once the process is finished, a [SubjectCode]_adj_info_trial_# file will appear in your patient's module_data folder. This file will also be loaded for each analysis, so there is no need to generate the adjacency array again unless a different-sized time window or time step is required.
After the adjacency array for a trial has been generated, clicking the "Generate Fragility Matrix" button will calculate the neural fragility values of each electrode over time and generate a [SubjectCode]_f_info_trial_# in your patient's module_data folder. This file is used to create the figures and outputs on the right side of the UI.
In each case, once the files are generated and saved in the module_data folder, there is no need to generate them again unless different parameters are desired. Thus, even though the data processing may be time-consuming at first, it only needs to be done once for each EEG recording.
Viewing Fragility
Once at least one [SubjectCode]_f_info_trial_# file has been generated, these trials will become available for selection under Step 3, View Fragility.
- Seizure Trial(s) for Fragility Display. Choose which trial to view the fragility of. If multiple trials are selected, the fragility values will be averaged across those trials.
- Select Electrode. Choose which electrodes to view the fragility of. Only loaded electrodes (the electrodes you selected when loading in the patient on the Select Data screen) may be chosen.
- Sort Fragility Map By. Choose whether you want to sort the electrodes (y-axis) on the fragility map by electrode number or by average fragility over time.
- List how many for most/least fragile. Choose how many electrodes you want to list in the most/least fragile electrodes table to the right of the fragility map.
- Seizure Onset Marker. Adjust the location of the vertical black dashed line on the fragility map. Useful if you want to share the fragility map and need to mark where the seizure onset occurred in time.
- Exponentiate 3D Viewer Fragility. This option exponentiates the fragility data to the third or fifth power in order to increase contrast between fragile electrodes on the 3D viewer. This is to better fit the color scaling built into the 3D viewer.
- Exponentiate Fragility Map too. Choose whether you also want to exponentiate the fragility map for better contrast. Usually not as necessary as it is for the 3D viewer contrast.
- Auto-recalculate. Choose whether you want the fragility map, table, and 3D viewer to automatically reload when one of the input values is changed. Otherwise, you would need to click the "Calculate Fragility" button whenever you want the figures refreshed.
Other notes
- 3D Viewer: You will need to click the "Click here to reload viewer" button when viewing fragility on the 3D surface for the first time.
- The Refresh button under Re-check files is useful for if you are manually importing pt_info, adj_info, or f_info files into the module_data folder and want the module to recognize them.
