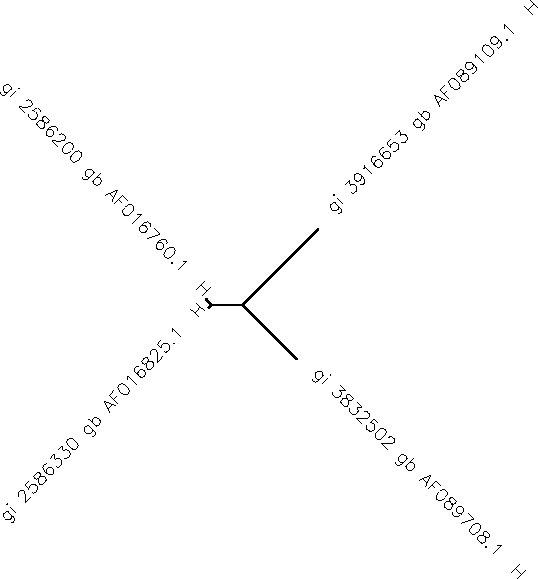Isaiah M. Castaneda Week 3
Part 1: PubMed The day’s activity started with an introduction to DNA databases and the FASTA format. This was done by examining sequences from an article entitled, “Patterns of HIV-1 evolution in individuals with differing rates of CD4 T cell decline.” This paper was found by performing a search on http://www.ncbI.nlm.nih.gov. I entered, “Patterns HIV-1 CD4 T cell decline Markham” to find this entry as the 12th result. The article could have also been found by simply using your average search engine. Another method, though not the best, would have been manually going through lists of Pub Med articles. There were several other articles on HIV-1 available as well.
Part 2: GenBank Unsure about how to find sequences that did not contain a link, I simply gathered information from the 4 clickable sequences available from the article.
I copied down the following sequences:
Accession number AF089708
>gi|3832502|gb|AF089708.1| HIV-1 isolate S15V4-10 from USA envelope glycoprotein (env) gene, partial cds GAGGTAGTAATTAGATCTGTAAAATTCACGAACAATGCTAAAATCATAATAGTACATCTGAATGAATCTG TAGTAATTAATTGTACAAGACCCAACAACAATACAAGAAGAGGGATACATATAGGACCAGGGAAAACATT TTATACAGGAGAAATAATAGGAAATATAAGGCAAGCACATTGTAACATTAGGGGTTCAAAATGTAATAAC ACTTTAAAACAGACAGTTAACAAATTAAGAGAACAATTTGTGAATAAAACAATAGTCTTTAATCATTCCT CA
Accession Number AF016760
>gi|2586200|gb|AF016760.1| HIV-1 subject 1 visit 1 clone 1 from USA, envelope glycoprotein V3 region (env) gene, partial cds GAGGTAGTAATTAGATCCGAAAATTTCACGAACAATGCTAAAATCATAATAGTACAGCTGAATGAATCTG TAGAAATTAATTGTACAAGACCCAACAACAATACAAGAAAAAGTATACATATAGGACCAGGTAGAGCATT TTATACAACAGGAGACATAATAGGAGATATAAGACAAGCATATTGTAACATTAGTAGAGCAGAATGGGAT AACACTTTAAAACAGATAGTTATAAAATTAAGAGAACACTTTGGGAATAAAACAATAGTCTTTAATCACT CTTCA
Accession Number AF08910
>gi|3916653|gb|AF089109.1| HIV-1 isolate S3V1-1 from USA envelope glycoprotein (env) gene, partial cds GATGTAGTAATTAGATCCGCCAATTTCTCGGACAATGCTAAAACCATACTAGTACAGCTGAATGAAACTG TAGTAATGAATTGTACAAGACCCGGCAACAATACAAGAAAAAGGGTAACTCTAGGACCAGGCAAAGTATA CTATACAACAGGACAAATAATAGGAGATATAAGAAAAGCACATTGTAACCTTAGTAGAGCAGATTGGAAT AACACTTTAAAAAGGATAGCTATAAAATTAAGAGAACAATTTCAGAATAAAACAATAGTCTTTAATCAAT CCTCA
Accession Number AF016825
>gi|2586330|gb|AF016825.1| HIV-1 subject 2, visit 4 clone 9 from USA, envelope glycoprotein V3 region (env) gene, partial cds GAGGTAGTAATTAGATCCGAAAATTTCACGAACAATGCTAAAATCATAATAGTACAGCTGAATGAATCTG TAGAAATTAATTGTACAAGACCCAACAACAATACAAGAAAAAGTATACATATAGGACCAGGTAGAGCATT TTATACAACAGGAGACATAATAGGAGATATAAGACAAGCATATTGTAACATTAGTAGAGCAGAATGGAAT AACACTTTAAAACAGATAGTTATAAAATTAAGAAAACACTTTGGGAATAAAACAATAGTCTTTAATCACT CCTCA
The label of the sequence in FASTA format contains information about the subject. The 1st two sequences listed were from subject 1, the 3rd from subject 3, and the 4th came from subject 2.
Part 3: Intro to the Biology Workbench I opened up the appropriate website to be confronted with the challenge of setting up an account. I performed this swiftly and quickly and then carried on. Worried that I might incorrectly save the 4 sequences I chose earlier, I copied and pasted the label into the label section and the sequence into its appropriate box for each sequence. I made sure that the program recognized that the FASTA template was being used as well. After running the multiple sequence alignment, I attained the following unrooted tree diagram.
Subject 2, visit 4’s sequence appears to be very similar to Subject 1, visit 1’s sequence due to their proximity and stemming off of the same point.
Activity 2
Part 1: Looking at clustering across subjects
The appropriate documents were downloaded off of the class wiki and uploaded to a new Biology Workbench session. Now, a more complex unrooted tree would be generated using 3 clones each from 4 different subjects. The subjects and clones used may be found in the excel link.
The following diagram was generated:
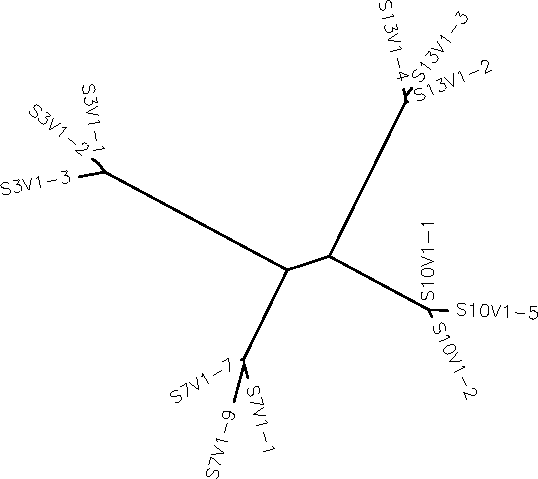
The clones from each subject do cluster together Subjects 10 and 13’s clones are the most clustered together. Such clustering perhaps is indicative of a close evolutionary relationship among clones. All of the clones of each subject appear to share a common ancestry, but the branching out of each could mean that the viruses are diversifying. None of the subjects cluster together, they are about equally apart. I take this to mean that each subjects virus is about equally different from the other.
Part 2: Quantifying diversity within and between subjects
I chose subjects 9, 7, and 5 to use for this piece. After counting the number of clones in each and recording this count into a table, I proceeded to run each subject through an alignment. For each subject, I counted the number positions for where at least one nucleotide was different across the collection of clones (a black nucleotide indicated difference). Using simple arithmetic, I calculated the denominator of theta for the subjects then applied some division to acquire the final theta calculation.
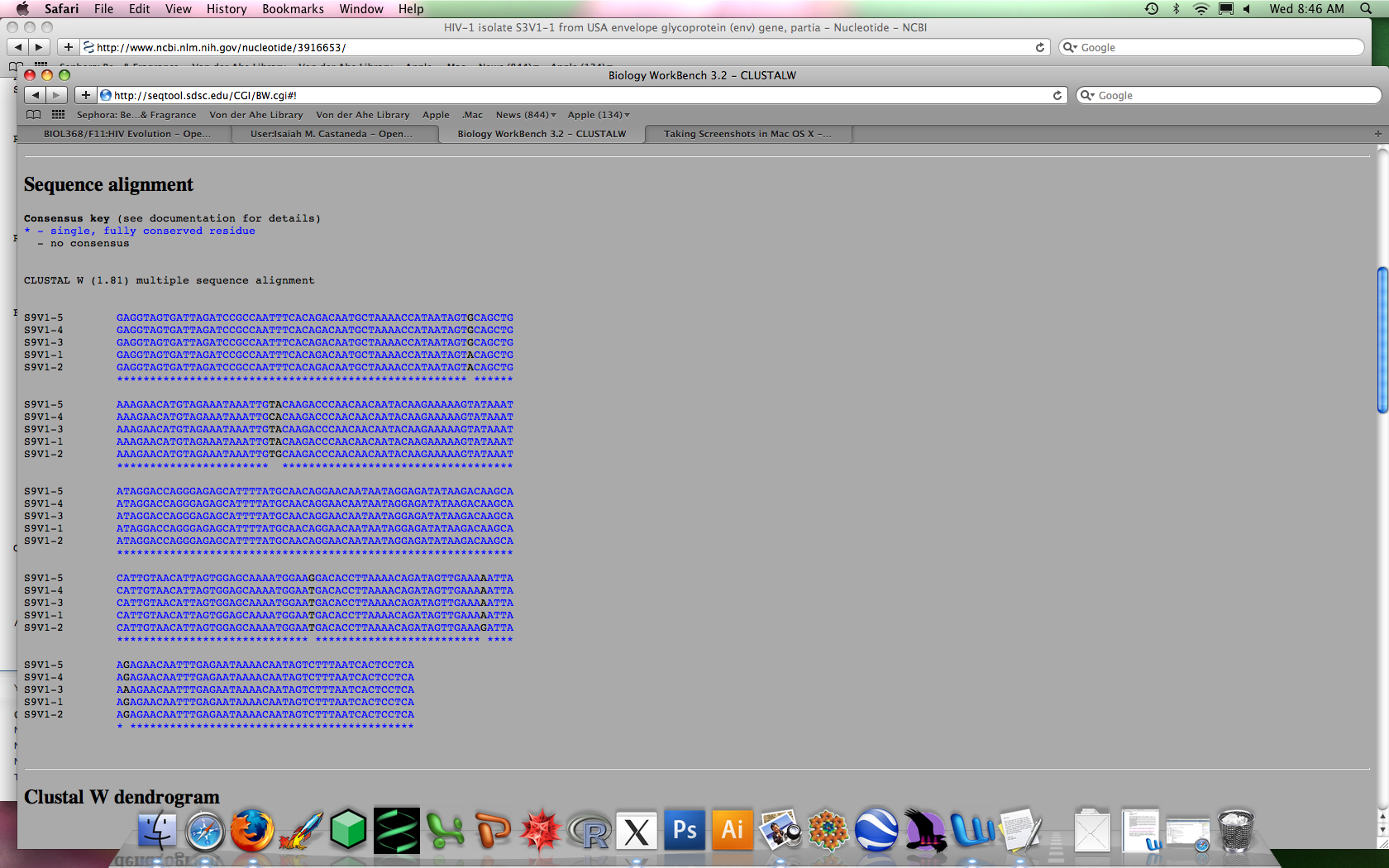
An example of what the alignment looked like is shown here
Min and max numbers were found further down on the page. The numbers were multiplied by 285 (the number of base pairs for each sequence) and then rounded to the nearest integer. By this point, I had become much more comfortable with the procedure and moved through the tools and controls very smoothly.
Min and max differences were then found on pairs of subjects for Table 4. Both tables 3 and 4 may be viewed from the same excel file link as before.