Ideas for wiki archive
Back to the latest topics
Wiki Meeting 6/22/05 Notes
We decided to open the wiki up to other groups local to MIT and that openwetware was not a crappy name.
Before we bring other labs on board we decided that it would be good to have an example of a protocol page (or several pages) as we would like to see them develop. The (somewhat of a) consensus on this was:
- A general page such as DNA Ligation which gives background information as well as a version of the protocol where each step is explained with the relevant biology. Also, the "biological lore" which is usually not written down in standard protocol books could accumulate on this page. The version of the protocol on this page could be imagined as stepping towards a "standard" protocol or be imagined as a beautiful information resource about the biology behind the steps, depending on your philosophy.
- Links within that page to individual (or user) protocol pages. This will enable labs/users to be more likely to participate/view the general page and would be a source to glean information to be put into the general page (such as identify points where the protocols differ).
To Do list: (before inviting labs):
- Perfect the DNA Ligation protocol (and some others)
- Make a good openwetware front page
- Put my/sri's comments on Talk:Main Page
- Add an "ettiquette" type of page
- Done: Etiquette -- need to work on formatting of this page.
To Do list: (additional ideas): --Reshma 11:29, 27 Jun 2005 (EDT)
- Develop a logo to replace the media wiki logo in the top left corner of each page (perhaps just a simple OWW from the logo on the Main Page).
- Been working on it. I have one that might work perhaps. I'll put up a few options. --Sri Kosuri 11:49, 27 Jun 2005 (EDT)
- Add a redirect from the old Endipedia site to the new OpenWetWare site.
- There already is one: [1] --Sri Kosuri 11:49, 27 Jun 2005 (EDT)
- Perhaps there should be a redirect from http://model.mit.edu/endipedia/index.php?title=Main_Page as well? --Reshma 12:18, 27 Jun 2005 (EDT)
- There already is one: [1] --Sri Kosuri 11:49, 27 Jun 2005 (EDT)
- Add a google search of OpenWetWare pages only to the Main Page to improve search functionality.
- This might take some hacking. We also should change the sidebar on the left. --Sri Kosuri 11:49, 27 Jun 2005 (EDT)
Also, we gave some thought to a stable lab protocol resource, potentially independent of the wiki. I think this could potentially be useful but needs to be better specified. For instance, I think that Endy:Protocol could serve a similar purpose and live on the openwetware wiki. We could then create a locked version of these once a year (or whenever) to be sued locally by people who wanted a stable resource.
I will also be adding the BEiNG (biological energy interest group) front page in next couple days, and give them user access on Monday of next week. I don't think they will be creating many protocols at least to start; they'll probably just be adding content to their space. --Jasonk 20:11, 22 Jun 2005 (EDT)
Wiki Name Update
The need for a name other than endipedia is becoming more apparent. An idea that has been floating around is OpenWetWare (a la opencourseware). It is meant to be a repository of biological information in general. For now, the domains openwetware.mit.edu / openwetware.org / openwetware.com have been reserved. What are people's thoughts? --Sri Kosuri 12:47, 22 Jun 2005 (EDT).
At the 6/22/05 meeting, everyone agreed on OpenWetWare. --Reshma 12:18, 27 Jun 2005 (EDT)
Protocols
I think we should change how we store protocols on the wiki. I think we should have general entries into the protocol section. For example, the entry on DNA Ligation will contain only information on what DNA ligation is, and how it generally works. In addition, this general page will have a listing of specific protocols that can be different for any number of reasons (For example, see the DNA ligation entries for the Endy protocol or the Knight protocol). In general, people should post why (or how and what are the consequences) their protocol is different. Feel free to post personal protocols here as well. --Sri Kosuri 11:38, 21 Jun 2005 (EDT)
This should also alleviate some of the concerns about losing control of lab protocols if we open up the wiki to too many other people. We could try and have a type of etiquette where you don't edit pages that have "Endy:" if you're not in the endy lab, or don't edit "skosuri:" if you aren't sri. This wouldn't be a hard line edit permissions systems of course, but rather just something that tells people, "feel free to correct minor errors, but if you're going to make a major change you need to justify it well and probably make sure the page "owners" know about it." --Jasonk 12:08, 21 Jun 2005 (EDT)
- Additionally, I really don't forsee this being a major problem at all, I think if we just tell people "this is a shared space, be respectful of each other" then it will develop naturally. For instance no one has made major edits to my user page other than me since we started the wiki even though we haven't had any sort of sub-domain rules in place. The last thing I want to do is give the impression that you might edit something and screw it up/piss someone off because, again, the biggest problem we will have is people not participating. (not people breaking stuff) --Jasonk 12:17, 21 Jun 2005 (EDT)
- While I can understand the justification for Sri's scheme, I am inclined to disagree with it. The most appealing aspect of the wiki in my mind is the fact that you can revise other's protocols at will. By placing different versions of the ligation protocol on different pages and prefixing them with names, you are discouraging (albeit subtly) group editing of protocols. I personally prefer that people revise and annotate my protocols as much as possible so that I can learn either more streamlined methods of doing things and troubleshoot my own experiments better. If my protocols are labeled with Reshma or even Knight indicating that someone "owns" those pages, I think that you discourage this. This is why I was in favor of retaining centralized locations for technical information. And why I would be in favor of retaining a single ligation protocol page.
- The other issue that Sri's suggestion raises is do we really want everyone to be following a different protocol? One of the goals of Synthetic Biology at least is to turn what are currently experiments into simple procedures that even a monkey can do so that the engineering of biological systems can be done with ease. As Tom has often pointed out, assembling two pieces of DNA together should be a standard procedure not a new experiment each time as is often the case in biology now. My understanding is that this is the driving motivation for projects like the standard strain and BioBricks plasmids. Admittedly, I was one of the people who first started labelling different protocols as Knight lab versus Endy lab (mostly because of a lack of better distinction between versions of protocols). But I guess I thought that over time, these two protocols versions would merge into a single protocol that everyone uses (with possibly some tweaks to account for your personal experiment). Having N number of protocols that M people use seems contrary to much of what Synthetic Biology is trying to accomplish.
- Of course, the caveat to what I have said above is that not everyone is doing Synthetic Biology and thus not everyone necessarily has an interest in pursuing a policy of standardization. I don't intend these comments to be a critique of Sri's suggestion. I am just trying to point out that the organization of the wiki may help determine whether we progress towards a large and diverse web of molecular biology information or towards a standardized set of procedures for biological engineering. We should think about what our goal is (and there are advantages to both) and organize our wiki accordingly. As is likely obvious, I am in favor of the latter. --Reshma 13:03, 21 Jun 2005 (EDT)
- I also agree with Reshma that standardization of certain sets of protocols is a good thing that we should work towards, as long as the conditions in which the standards are to be used are defined. I think the point of this proposed change, is that we can now do both. Not only could this be a place for biological engineers to work on a shared, standardized protocol. It could be a great place to define/debug/improve those protocols based on the information being stored by individual researchers and what works for them. Practically, there is no reason that those SB standards could not be placed in a protocol such as SB:The Protocol. I think forcing people to contribute to a shared protocol will, in the short-term and long-term, discourage engagement of people who don't think they should change "the standard". In addition, it will discourage labs that think don't want to use the "standard protocol" for whatever reason (e.g., don't want to pay for a qiagen kit). As a nascent community, I would be more worried about this, than too many people putting their own individual protocols up. --Sri Kosuri 13:42, 22 Jun 2005 (EDT)
Wiki Changes incorporating SyntheticBiology.org and multiple entry pages
The process has started. See Main Page v2
Adding a figure to clarify what I was suggesting Endy 14:37, 12 Jun 2005 (EDT)
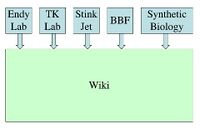
- that we rename the front page of the wiki (i.e., "endipedia")... by getting rid of the front page of the wiki
- that we have one wiki which serves endy lab, tk lab, and our synthetic biology working group.
- that the endy lab web page move in its entirety into a wiki-format, and that http://mit.edu/endy point to an internal wiki sub-page that is the new "endipedia."
- that the TK folks consider constructing a TiKipedia for the TK lab and that there be an internal wiki sub-page that is the TiKipedia port of entry.
- that the syntheticbiology.org website point to an internal wiki sub-page that is the new syntheticipedia, and that we move all information now on that website into wiki-format -- that site has gotten really stale, even though we are doing a lot more work.
- that a new URL (www.stinkjet.org) point to a still different internal wiki page that we can use to promote discussions of issues related to the development of second generation biological technology.
(emailed out by Drew 6/9/05)
- I think that one of the major advantages of the wiki as it stands now is the centralized nature of the technical information. By having having a single wiki page devoted to say "Ligations", everyone can contribute and place information about debugging problematic experiments in one place for the benefit of others. If we had separate ligation pages for the Endy, TK lab and Synthetic Biology then the barrier is raised to ongoing discussions between people about protocols. Therefore, if we do decide to change the wiki organization I would vote for maintaining a centralized place for technical information for all three groups. We can still eliminate the Main Page just by creating a new technical information front page that is linked off of all three groups.
- I prefer sbpedia to syntheticipedia.
- What is the difference between the synthetic biology wiki and a wiki about development of second generation biological technology? Shouldn't the latter be a part of the former?
- --Reshma 09:41, 10 Jun 2005 (EDT)
- We should definitely keep the central repository of information but I think the idea of having several "fronts" to that repository makes sense.
- This would probably require us to be better at crosslisting or categorizing our pages so that an interested visitor coming in through the sbpedia front page can get at the same pages that someone coming in from the Endy lab page can get at.
- Might be good to incorporate the iGEM team wiki also so that it is easier for them to use our info. and for us to give feedback on their plans.
- --BC 10:03, 10 Jun 2005 (EDT)
- agree with above, i think it might not be too difficult, we just need to make sure that we all point to the same centralized tech pages. I think one organizational decision is whether it is better to have another layer (i.e. a technical info front page) or multiple shared tech pages which can be linked to from any front page that would be relevant, for instance, tech pages:
- Protocols
- Equip
- Strains
- Vectors
- etc.
- Both Endy and Knight front pages would link to all of these, but SBpedia (or i might just call is 'SyntheticBiology.org') might only link to Strains and Vectors for instance.
- definitely agree with bringing the iGEM wiki page into the umbrella, will let them use (and improve) our protocols more easily.
- Stinkjet could be specific to forward looking BBF-type stuff, i.e. more focused on issues of open source biological engineering, safety when everyone has a 'stinkjet' on their desk, etc. SyntheticBiology.org might point more towards current issues, i.e. latest tech reports on BBs stuff, info about the registry, etc. They would of course point towards each other... maybe there is too much redudancy, but i do like the stinkjet.org domain name.
- --Jasonk 10:21, 10 Jun 2005 (EDT)
- I like the idea. I think we should begin building out the front pages though until we make the switch. Also, we should figure out a naming scheme for lab/group specific stuff, like group meetings and whatnot. Also, we should agree on early on about what parts are going to be jointly worked on, ie., protocols. --Sri Kosuri
- I also don't see the point of having multiple wikis. It just makes searching/management more difficult. Wikipedia has information about everything under the sun (except for synthetic biology) and yet they don't seem to have a problem with needing to split stuff into different encyclopedias. What about the concept of Wikipedia:Help:Namespace rather than completely separate pages? The only issue I can think of is for access control where certain pages should have different permissions than others. I don't know if that's possible but then the whole idea of a Wiki should be public access. So I'm all for a single wiki and if there's something that needs to be separated by labs, prefix pages with Endy: if needed.
- I've redirected syntheticbiology.org to this wiki. There was no Synthetic Biology page here?
- As for sbpedia, it sounds too much like expedia and I'd rather stay away from any Microsoft associations.
- --Austin 16:07, 11 Jun 2005 (EDT)
Communicating Changes to the Wiki
This has come up a couple of times. For those who check changes to the Wiki more frequently than checking for updates to \. I don't think its a problem. But it might be good to find a way to easily communicate changes to the wiki to all (who is all?). Possible ideas include -
- Just mail the endylab and tklab when you put up something. Easy but not very streamlined.
- Get the wiki to automatically send daily/weekly updates of non-minor changes to a mailing list of interested parties. Not sure if this is possible. Actually looks like enotif will do this, its a plugin for mediawiki. You choose to receive emails or not and which pages you want to be alerted to changes in. --BC 16:25, 26 May 2005 (EDT)
- Might be possible to automatically keep a recent updates section on the main page.
- Have everyone install RSS readers and subscribe to the RSS feed. Maybe a bit OTT!
Feedback?
--BC 16:15, 26 May 2005 (EDT)
Synthetic Biology Community on DSpace
I posted this below under Presentations/posters/papers but then thought it might not be very noticeable so I am reposting it with its own heading. It is a page about creating a Synthetic Biology community on DSpace which would enable us to post all digital materials produced by the group online and determine who gets access to them. Feedback much appreciated (you can also email me if you don't want to post on the wiki). --Reshma 15:54, 26 May 2005 (EDT)
User Wikis
Barry and I thought it might be cool to do an experiment evaluating "personal" wikis as a data organization tool. Similar to the local file structure that you would have in your computer home directory, but with better linking between sections and more transparent content. i.e. Not just word files in sub folders that I can't readily search or link between. (yes sri, i know spotlight can search them, but you still can't link easily)
I also think that if lab members format more of their personal remarks, etc, in wiki format they will be much more likely to make their way into the public wiki space. So will see how this works out. Of course this could all be done within the user pages on the current wiki, but i think there is some value to being able to take personal notes that aren't public domain. (but which eventually can move there easily).
Thoughts? Also, if you want to try it out for yourself we can set that up. Jasonk 14:41, 26 May 2005 (EDT)
- -Mac Users: if you want an easy walk through on how to set up your own wiki, go here. Takes about 30mins and sets up an Apache Web Server, a SQLl database, PHP and mediawiki. And its all OSS also! --BC 16:19, 26 May 2005 (EDT)
- -This seems like an interesting idea. One thing I am wondering is say I have a personal wiki and document a project on my personal wiki. How easy would it be for me, in the future, to move my wiki pages onto the lab wiki? Especially if I only want to move a section of my wiki to the lab wiki. If it is difficult to do this then it reduces the likelihood that people will eventually release their personal wiki pages to the public domain. Personal wikis might still be useful for the person but perhaps less useful as the documentation of a project that eventually gets released to the world. --Reshma 13:14, 27 May 2005 (EDT)
- -My thought on this is that right now if i had a project that i didn't want on the public wiki i would be documenting it on a word file on my personal computer anyway. So using a personal wiki instead just means that the formatting is already in place to just cut and paste to the public wiki when/if i'm ready to do that. So there may be a little work in moving it over (i.e. there's not just a "merge" button), but it should be less work then if i had just made a word file.Jasonk 15:31, 29 May 2005 (EDT)
- -This raises one of a few drawbacks to the personal wiki idea. The others are the difficulty of making a well ordered hard copy of your material, and embedding files is more cumbersome than when using Word or something equivalent. The merging of wikis problem might be partially solved by using a similar structure for a personal wiki as for the lab wiki and using the same list of categories, which would automatically integrate your pages into the lab wiki. For that to work, we'd need to rely on categories more than we do now. I'd be interested to hear from someone who knows about SQL databases and how easy it might be too merge two databases or portions of one database with another. I liked the personal wiki idea just as a way to organize my own material and make it easier to cut and paste individual pages to the lab wiki if I wanted. I think this would be really useful when developing a new protocol, it could live on a personal wiki until it works and then get moved over to the lab wiki. Time will tell though if it is ueful. --BC 15:37, 27 May 2005 (EDT)
Categories
I think we could use the Categories feature of the wiki more effectively. Having a list called Protocols and a category called Protocols seems redundant. What about using Categories to connect group pages across lists? For instance, a "Running systems in a μreactor" category could include links to ordering microfluidics (under Protocols), the Scope (under Equipment), media (under Materials) and other relevant pages. This approach would essentially use Lists and Categories as two separate "dimensions" along which pages can be grouped. --Reshma 21:13, 25 May 2005 (EDT)
- Yeah that's good -- I like the idea of grouping things from multiple lists in relevant ways, the question is when do categories become more useful then just creating a "Running systems in a μreactor" page and then linking to everything like that? For instance take the media (or the scope), how many categories could it end up being in? running a μreactor, running a macro chemostat, growing a batch culture, etc. In that case I think having seperate "group pages" or something that each individually pointed to the media might be better. Jasonk 00:55, 26 May 2005 (EDT)
- I think the categories might pay off better if we had a lot of protocols that were distributed accross different heirarchical list schemes. For instance if I started adding new protocols to my user page directly, in some sort of heirarchy that was relevant to me, i.e. "frequently used protocols" vs. "infrequently used protocols" or something, then I could just slap a category tag at the end and know that I had placed it into an appropriate spot in the aggregate general protocol area. Also, could imagine if you aggregated a few lab wikis together that had pre-defined heirarchies, the categories could serve to create a common protocol space without everyone having to conform to the same list scheme. So it might be that they are more useful down the line (or not), but it certainly doesn't hurt to give any system a try and see if it proves useful (can always change it easily). Jasonk 00:55, 26 May 2005 (EDT)
I guess i'd like to think about what other categories would be useful.
Lab Notebook
Not planning on throwing my lab notebook online just yet, but if you were to consider using the wiki for such a thing you might want to have some sort of encrypted date/user signature. This would potentially give the more paranoid among us more ammunition in their attempt at defaming the person who "scooped" them in this guerilla war we call academics. (sarcasm tags would also be nice) Jasonk 18:27, 23 May 2005 (EDT)
Some relevant links:(thanks ilya)
- Open Source Electronic Lab Notebook Software
- Also, the pharmaceutical industry has some pretty serious electronic signature rules, which are patent-safe. Expensive to implement and likely overkill for academia but:
Suggested new name: knendipedia
- I think knendipedia might be a more appropriate name for this wiki :). (I liked knendipedia better than enightipedia). What do you think? -Reshma 10:07, 11 May 2005 (EDT)
- Maybe we should just go with something like SynthBioWiki or something, would make it easier to add more friends down the line, not sure the knendipedia approach scales, shetkoscanknendipedia :) Jasonk 18:26, 23 May 2005 (EDT)
- How about SBpedia - pronounced speedia --BC 19:33, 23 May 2005 (EDT)
- My one problem with the above two suggestions is that not everyone does 'synthetic biology' ie me, francios, jeff, ty, heather... --Sri Kosuri 23:29, 25 May 2005 (EDT)
- good point, then how about some generic bio term, like geneipedia, or some such nonsense.Jasonk 00:53, 26 May 2005 (EDT)
- My one problem with the above two suggestions is that not everyone does 'synthetic biology' ie me, francios, jeff, ty, heather... --Sri Kosuri 23:29, 25 May 2005 (EDT)
User page / posting personal ideas
It would be nice to post pages which are non-editable for things like your personal opinion on something scientific. I.e. the type of stuff that might be found on your user page. This also becomes more important if we open the wiki to be world-writable.
We could accomplish this by making all users admistrators since admins have ability to "protect" pages and lock them from editing. This would work so long as members were all trusted, but if we decide to expand member base we might like to have a way of doing it without giving everyone admin access.
Anyone know of how to do it without giving admin access?
Presentations/posters/papers
One thing that struck me as potentially useful would be a place to archive presentations and posters. Might be useful to be able to look over others presentations to see how they presented things etc. Similarly, it might be nice to also be able to post works in progress like papers or various writeups etc. Of course, I can imagine several levels of release like private (probably wouldn't post), release to the Knight/Endy lab (wiki?) and release to the world (http://www.syntheticbiology.org?).
- Does the wiki have a good mechanism for posting documents? A quick look suggested to me that you could only upload pictures.
- i think you can only do pictures, best bet for docs seems to be store elsewhere and link - i like the idea of having a place on model where we can dump all wiki docs/ppts/etc in a centralized place.Jasonk 14:40, 28 Apr 2005 (EDT)
- Can now post whatever type of file you like Jasonk 18:30, 23 May 2005 (EDT)
- Can sections of the wiki be made off limits to those without user accounts. ie non-readable?
- Or alternatively people can post documents in their own public directories. Use MIT's certificate system to control who access the directories and then just put an external link on the wiki. This gives maximum control over access to the author but does not centralize documents as much. This also may be problematic as people graduate or leave the lab.
- This is a good idea for now. In the future, we could create a section on model to have a username and login for private files. --Skosuri 14:36, 28 Apr 2005 (EDT)
- Any better ideas?
Collaborators
Forgot to mention in lab meeting, but will be giving the Knight lab write access to the pages, talked with reshma and austin and they thought it made more sense to just have one wiki we could share rather than cross referencing seperate wikis. (might need a new name then).
Also, we could consider starting Collaborative Projects pages, for things such as the Standard BB strain, etc.
Ideas from Lab Meeting 4/27
Openness of the wiki
Options for public access:
- World-writable and readable
- World-readable only
- World-no access
- Hybrid
- World can write to discussion but not to main articles.
We decided that we should go with option 2 for the time being at least until the wiki stabilizes and then consider making the wiki world-writable.