ISEEM Progress September 2008
| Home | Project | People | News | For Team | Calendar | Library |
I. Progress made toward completion of our goals
A. Personnel
Hiring is complete. The people funded by the iSEEM Moore Foundation grant are (with start date in parentheses if not started yet):
- Jessica Green
- Steven Kembel, PostDoc (11/1/08)
- James O' Dwyer, PostDoc (phasing in from another project)
- Katie Pollard
- Josh Ladau, PostDoc (10/22/08)
- Sam Riesenfeld, PostDoc (11/1/08)
- Jonathan Eisen
- Dongying Wu, Staff Scientist
- Martin Wu, Staff Scientist
- Sourav Chatterji, PostDoc
- Srijak Bhatnagar, Bioinformatics Engineer
The following students are involved in the project, but not funded by it:
- Liz Perry, PhD Student (Green lab)
- Marisano James, PhD student (Eisen lab)
B. Research
STAP pipeline expanded into a scientific workflow (Eisen lab)
We are working with CAMERA to convert our STAP rRNA analysis pipeline (which was described in the previous report) into a Kepler Workflow to be runnable from within CAMERA. We had a meeting with the CAMERA and Kepler groups to plan how to tackle this and work is underway to convert our initial Kepler Workflow into something that can be used by CAMERA.
Automated alignment masking (Eisen lab - Sourav Chatterji and Martin Wu)
One of the challenges for the iSEEM project and an project attempting to carry out phylogenetic analysis of many different genes is the need to assess the quality of multiple sequence alignments that are used for the phylogenetic reconstructions. In the past, including in the development of the STAP and AMPHORA software outlined above, we have carried out extensive manual examination of alignments to identify regions of the alignment that may not be useful for phylogenetic analysis. Known as "masking" this manual work takes both significant expertise and time.
We have developed ZORRO, a probabilistic masking program for taking into account alignment reliability during phylogenetic inference. Using posterior probabilities of a pair HMM based alignment model, ZORRO assigns confidence scores to each column in the alignment. Using either a cutoff or weighting based approach, these confidence scores can be seamlessly integrated into any phylogenetic inference pipeline.
Using the standard Balibase benchmark data-set, we were able to demonstrate that ZORRO is able to measure alignment accuracy with very high (>95%) sensitivity as well as specificity. We have also created a "benchmark" data-set of Alphaproteobacteria genes for comparing phylogenetic inference protocols in bacteria. Using this benchmark data-set, we were able to demonstrate that phylogenetic protocols using ZORRO showed statistically significant improvement over protocols not using any masking.
Automated Phylogenomics Inference Pipeline (AMPHORA) (Eisen lab - Martin Wu)
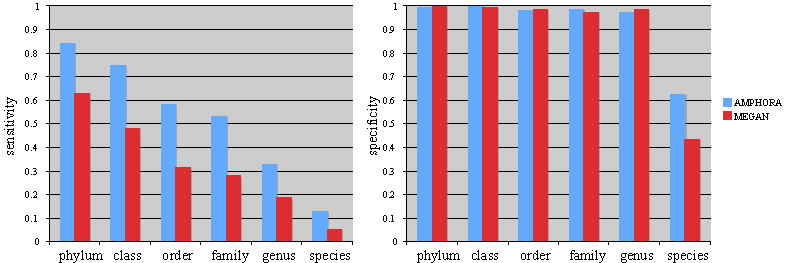
As part of the iSEEM project, Martin Wu is continuing developing AMPHORA, an automated phylogenetic inference pipeline for fast, accurate, and large-scale phylogenetic analyses of genomic and metagenomic sequence data. One of its main applications is to assign phylotypes to metagenomic sequences, which can be used to estimate the composition of a microbial community. Recently, Martin refined the tree-based phylotyping algorithm by taking into account of the evolutionary distance of a query sequence relative to the other reference sequences. In a large-scale, systematic phylotyping simulation study, we have shown that AMPHORA significantly outperformed a similarity-based method (MEGAN) in the sensitivity without losing the specificity. Our manuscript describing AMPHORA was recently accepted by Genome Biology and is in press. The software itself will be made freely available to the research community.
Gene family classification for phylogenetic marker identification (Eisen lab - Dongying Wu)
In order to identify potential phylogenetic markers for metagenomic studies, we are developing methods to analyze the gene distributions across genomes with compete genome sequences. We selected 85 bacterial and 15 archaeal genomes for the initial study. A maximum likelihood tree of 720 bacterial genomes based upon 31 concatenated genome markers were used for the bacterial genome selection, while the selection of archeal genome was based on a tree of the archeal radA genes. A program called maxPD was developed to automatically select taxa from a phylogenetic tree so the taxa are listed according to their contributions to the phylogenetic diversity based on the tree. The genomes that contributions most phylogenetic diversity were selected. In the selecting process, we only selected the genomes with publications and we also avoided genomes that undergone severe genome reductions.
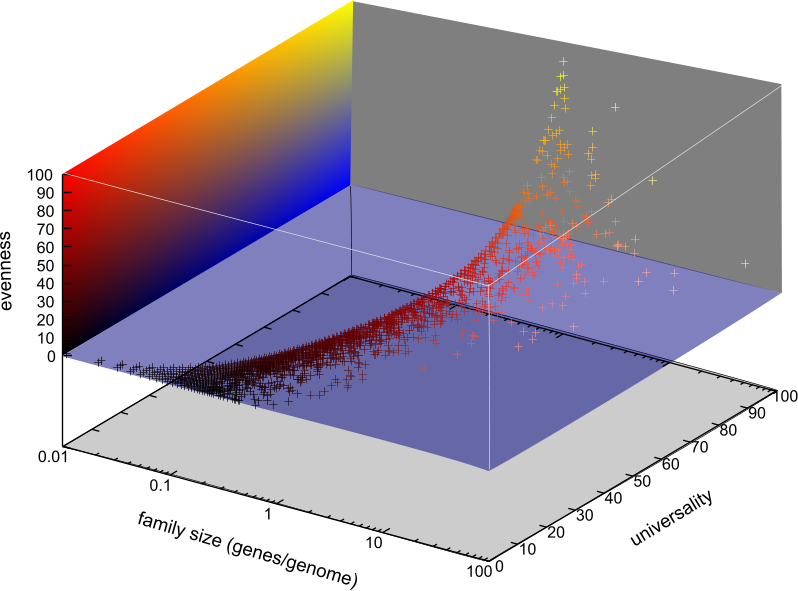
A protocol has been developed to classify gene families automatically. The protocol is base upon Blastall and the Markov clustering algorithm so the protocol are automated, robust. We use an e value cutoff of 1e-10 in our test run. For the 313139 genes from the 100 selected genomes, we identify 23336 gene families that include a total of 239453 genes. In order for automatically select gene families for potential markers, we've developed a program that not only evaluates the universality of the gene families, but also estimates the evenness of the distributions of the genes across a selected group of genomes (See the figure below).
Across all 100 the selected genomes, we identified 30 gene families that appear in almost all the genomes with even gene distributions, 12 were ribosomal protein subunits that were already included in our gene marker collections which indicate the approach works in principal. The rest of the candidates are mostly tRNA synthetases among others. Different cutoffs will be applied for more potential marker selection, and the evenness and distribution estimation program was developed in a way that potential markers for specific phylogenetic groups (for example, a phylum or a class) can be easily identified.
phyloP: phylogenetic p-values (Katie Pollard)
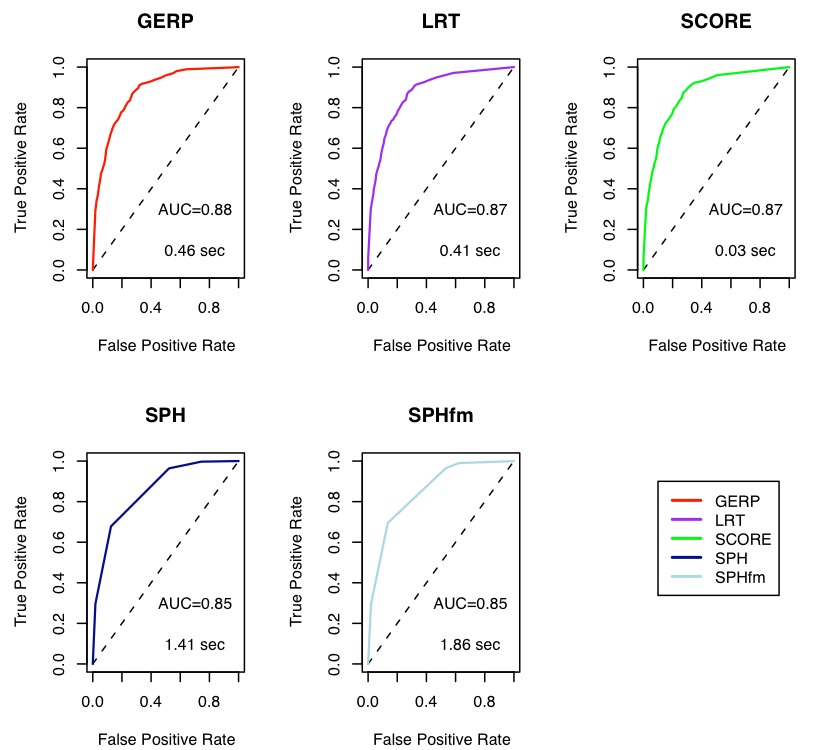
We are developing methods to statistically score multiple sequence alignments for changes in evolutionary rates of substitutions. This will be used in the iSEEM project for the studies of evolution of protein families. These scores are negative log p-values from several tests for acceleration and/or deceleration of substitution rate. Test statistics include likelihood ratio tests, score tests, and functions of the expected numbers of substitutions (two published methods: SPH and GERP). These tests can be performed on the whole tree or a subtree (i.e. clade) of interest, allowing detection of lineage-specific selection. Calculations are performed on single alignment columns, and when alignments are deep enough (i.e. contain enough species) simulations using parametric (time-reversible, Markov) models of sequence evolution indicate that we can detect subtle changes in evolutionary rates at single base pair resolution (See figure below).
Interestingly, the different methods have very similar performance. Scores from single sites can be combined to score genomic elements of various lengths, e.g. genes or operons. We are now conducting non-parametric simulations by resampling from real alignments with 4-fold degenerate sites representing neutral sequence and non-degenerate sites (second codon positions) representing sequence under negative selection. These data-based simulations will account for the effects of gaps and missing data in multiple sequence alignments, features that are not present in parametrically simulated data. This is joint work with Adam Siepel at Cornell University.
ComboDB development (Eisen lab - Srijak Bhatnagar)
For many of the precomputes we are doing as part of the iSEEM project, we are making use of a genome database developed by Martin Wu in the Eisen lab called ComboDB. ComboDB is a comprehensive database of complete genome sequences (including information on proteins, the genes encoding these proteins (CDSs), rRNAs, tRNAs, and genome assemblies) and their various attributes, such as taxonomy, GC content, gene locus, etc). In the past this database has been stored in a flat file format. This database has now been converted into a hybrid database, where the various information will be stored as a relational database and the sequences as a flat-files linked to this relational database. Such a database with add enhanced querying, easy accessibility and better interface for retrieval and mining of data. In essence, it will house the data used in various genomic analysis to develop the parameters for metagenomic analysis and will link to many tools for on-portal analysis, in an easy to access fashion. As of now, the database has been created and the front-end for it is under construction, following which various tools will be added to it.
C. Communications, Collaboration, Outreach and Education
Project group page at Open Wet Ware
We are building a iSEEM project page which will be housed at via the OpenWetWare site. A draft of the page is up at http://openwetware.org/wiki/ISEEM. After exploring many options for the "public" face of this project, we have chosen to use Open Wet Ware (http://openwetware.org/wiki/) because it is part of an "Open Science" initiative that is consistent with the open goals of the iSEEM project.
CiteYouLike
We are keeping a bibliography of relevant literature on CiteYouLike (http://www.citeulike.org/groupfunc/6072). This is a publicly available collection (although only group members can edit it).
Metagenomics educational materials
We are developing a wiki for iSEEM metagenomics educational materials. These will include figures and slides for use in courses, seminars, and conference talks by iSEEM team members. We were inspired in part by Yuzhen Yi, with whom Katie Pollard met in September at Indiana University. Yuzhen has taught a seminar course on metagenomics: http://mendel.informatics.indiana.edu/~yye/lab/teaching/spring2008-seminar.php
The second purpose of the wiki is to develop tutorials for sharing (meta)genomic data analysis methods and pipelines among team members. These will be particularly useful for incoming postdocs who have a variety of backgrounds and do not all have hands-on experience with genomics data. The interactive tutorial will take them step by step through the various tools & analysis involved in metagenomics. Our first tutorial will focus on sequence data analysis, from chromatogram processing through multiple sequence alignment. Srijak Bhatnagar is leading this effort.
Couses
- Jonathan Eisen is designing a metagenomics course for Spring 2008 at UC Davis. All course material will be made available online. In addition it will be audio and possibly video recorded and made available through iTunes.
- We are exploring the possibility of jointly offering a metagenomics class for the community at Bodega Bay in the Summer of 2008
Posters
- A Simple, Fast and Accurate Method of Phylogenomic Inference. Martin Wu & Jonathan A Eisen. The 16th International Microbial Genome Conference. Sept 14-18, 2008 Lake Arrowhead, CA
- Metagenomic Biodiversity and Biogeography: Theory and Practice. O' Dwyer JP. Perry EB, Green JL. James O’Dwyer and Liz Perry presented a poster at the 16th Annual Meeting on Microbial Genomics, at Lake Arrowhead. The aim of the poster was to outline a research program to develop predictions for metagenomic data using ecological theory. The data presented was based on the Sorcerer II Global Ocean Sampling expedition, and our analyses used the metagenomic similarity metric introduced by Rusch et al (PLoS Biology 2007). We demonstrated that genomic similarity shows significant distance-decay patterns along several important environmental gradients. The overall metagenomic similarity between samples is significantly correlated with geographic distance, temperature, chlorophyll a concentration, and ocean depth, while other environmental ‘metadata’ such as sample depth and salinity do not correlate with metagenomic similarity. The second part of the poster focussed on the development of new metrics to describe both within-sample metagenomic diversity and between-sample metagenomic similarity, and in particular how these metrics may be relatable to predictions of Hubbell’s Unified Neutral Theory of Biogeography and Biodiversity. We discussed number of subtleties and complications arising due to the nature of metagenomic data, and certain weaknesses with the metric used in our preliminary analyses. Despite these issues, we cited the clear spatial patterns in these data as indicative of important ecological processes at work, and therefore worthy of further investigation.
File:Arrowhead Conference vfinal.jpg
Publications
- Wu M, Eisen JA. A Simple, Fast and Accurate Method of Phylogenomic Inference. Genome Biology (in press)
II. Group meetings
Biweekly PI meetings
Eisen, Green and Pollard are meeting bi-weekly via conference call (for a while, Wednesdays 9:30 – 10:30 AM, moving this to Tuesdays 10-11 AM). During these meetings we are mostly dealing with logistical aspects of the iSEEM project. Notes are kept on the group wiki and are included as an attachment.
Biweekly iSEEM meetings
On the weeks that are alternate to the PI meetings we are holding iSEEM lab group meetings for all involved participants (including those at UCD, UCSF and UO). The purpose of these meetings is to discuss the scientific aspects of the iSEEM project and foster collaborations and interdisciplinary efforts between the PI lab groups. Notes are kept on the group wiki and are included as an attachment.
CAMERA-Kepler-iSEEM joint meeting
We had a joint group meeting with people from CAMERA, the Kepler Workflow Project, and our group. The meeting was held at UC Davis in the Genome Center and involved the following people
UCSD/CAMERA
- Amarnath Gupta
- Ilkay Altintas
- Jeffrey S. Grethe
- Paul Gilna
Kepler-Davis
- Bertram Ludascher
- Shawn Bowers
- Timothy McPhillips
- Sean Riddle
Eisen Lab/iSEEM
- Amber Hartman
- Jonathan Eisen
- Martin Wu
- Srijak Bhatnagar
- Dongying Wu
In the meeting we discussed how to work with CAMERA to take methods developed from the iSEEM project and integrate them as Kepler Workflows (http://kepler-project.org/) within the CAMERA system. We came up with a plan for the next few months of work which involves first working on a rRNA analysis workflow and using it as a test to see how to take workflows from the iSEEM project and develop them into CAMERA tools. Once this is done we will move on to protein analysis workflows.
Other meetings
- Katie Pollard, Joshua Ladau, Jonathan Eisen, and Jonathan Eisen's lab group met at UC Davis on September 22. Research directions for Josh were discussed, including development of optimal community phylogenetic statistical methods, and methods for inferring species richness and abundance from metagenomic data. Josh will be starting as a postdoc in Katie Pollard's laboratory on October 22.
- Jonathan Eisen, Martin Wu, James O'Dwyer, and Liz Perry held a meeting at the Lake Arrowhead small genomes conference to discuss methods for calculating metagenomic distances.
III. Any unexpected challenges that imperil successful completion of the Outcome
No significant unexpected challenges in the last quarter. The project is moving along quite well now that all personnel are in place. We are still exploring different mechanisms for group discussions and group interactions across the three labs, but this challenge was not unexpected. We have decided to try and coordinate more tightly the various projects at the different labs by trying to make use of the same data sets in our analyses (e.g, simulated data sets or real ones).