ISEEM Progress Final
1. GRANT OUTCOMES, OUTPUTS & ACTIVITIES
To the best of your ability, please address progress towards achieving each Outcome and Output listed below. In particular, please describe how progress towards accomplishing each Activity culminates in progress towards achieving a given Output. Mark the approximate percent complete for each Output. Also, in the appropriate space please describe how, in aggregate, the progress towards achieving these Outputs culminates in progress towards achieving the Grant’s Outcome.
Outcome1, Output 1: Characterize metagenomic microbial biodiversity and biogeography from CAMERA data.
1.1 Guidelines and weightings for using different gene families in metagenomic based diversity assays (Eisen).
1.1a. Generate alignments and trees for targeted set of 50-100 gene families; Deliver trees to projects 1b, 1c, 1d, 2a, and 2b; Generate scores for targeted set of 50-100 gene families; Test utility of scores using simulated and real metagenomic data sets.
- 100% complete
In order to build gene families for all archaeal and bacterial genomes, we started from 15 archaeal and 85 bacterial genomes. The 100 organisms were selected so that phylogentic diversities (PD) are maximized. For the bacterial genomes, we’ve identified 56 phylogenetic marker candidates, 25 of them are new bacterial markers.
To identify phylogenetic markers across Archaeal and Bacterial domains, we carried out gene family building and marker identification of a range of phylogenetic groups. 38 markers have emerged as good phylogenetic markers for both Archaea and Bacteria when we studied the universality and evenness in terms of genome distributions and phylogenies of the gene families we’ve built.
- Contributors: D Wu, J Eisen
1.1b. Generate alignments, trees, and scores for expanded set of ~500 families; Test utility of these scores using simulated and real metagenomic data sets.
- 100% complete
We’ve established a protocol to identify automatically phylogenetic marker candidates for any given phylogentic groups. The protocol uses BLAST and MCL clustering algorithms to generate gene families for a given group of genomes. Phylogenetic trees are built for the gene families and clades from the trees are automatically sampled and evaluated for universality and evenness in terms of their distributions. HMM profiles are built for the clades with genes distributed across the organisms with a single copy in each genome. HMM searching against the entire proteome of the group is applied to evaluate how distinct the gene families are. We have built distinct single-copy gene families that are evenly distributed within a phylogenetic group (we’ve studied the archaeal domain and 10 bacterial phyla). HMM profiles were built for 5133 families that can be potential markers for the lineages of interest. Clustering and tree building analysis of the consensus sequences from all the families reveals that we can identify 62 gene markers that each span archaea and at least 4 bacteria phyla, as well as 324 bacterial gene markers that each cover at least 5 phyla. The 5133 gene families prove the core of a marker database for phylogenomic and metagenomic studies.
Table DW1. Universal single-copied genes for different phylogenetic groups
| Phylogenetic group | Genome Number | Gene Number | Maker Candidates |
| Archaea | 62 | 145415 | 106 |
| Actinobacteria | 63 | 267783 | 136 |
| Alphaproteobacteria | 94 | 347287 | 121 |
| Betaproteobacteria | 56 | 266362 | 311 |
| Gammaproteobacteria | 126 | 483632 | 118 |
| Deltaproteobacteria | 25 | 102115 | 206 |
| Epislonproteobacteria | 18 | 33416 | 455 |
| Bacteriodes | 25 | 71531 | 286 |
| Chlamydae | 13 | 13823 | 560 |
| Chloroflexi | 10 | 33577 | 323 |
| Cyanobacteria | 36 | 124080 | 590 |
| Firmicutes | 106 | 312309 | 87 |
| Spirochaetes | 18 | 38832 | 176 |
| Thermi | 5 | 14160 | 974 |
| Thermotogae | 9 | 17037 | 684 |
- Contributors: D Wu, J Eisen
1.1c. Integrate scores with development of diversity assays.
- Status: 100% complete
Each protein family has been scored for various metrics including copy number, universality, phylogenetic consistency, and others. These metrics were used as a guide to select protein families for inclusion in AMPHORA 2 which is discussed in more detail below.
- Contributors: D. Wu; A. Darling; J. Eisen.
1.1d. Create and update database of score for different genes.
This deliverable was modified to focus on creating and updating a database of protein families from genome sequences.
1.1.d Modified: Building de novo protein families'
Family recall percentage distribution. Recall is defined as the percentage of sequences from a family that were correctly assigned to the family that they are part of when scanning the family HMMs against the starting sequences.Family precision percentages distribution. Precision corresponds to the number of sequences that were incorrectly assigned to the family profile when scanning the family profiles against the starting sequences. A precision value of 1.0 indicates that only sequences belonging to the family were assigned to the family by Hmmer.Pfam to novel familiy distributions. The majority of our novel families only have a common domain with only 1 Pfam.
- Status: 95% complete.
We clustered all proteins from 1,894 microbial genomes into protein families using Markov clustering (MCL algorithm) and a protein distance matrix based on a massive all-vs-all BLAST search. This de novo clustering procedure used 7.2 million full-length protein sequences. We identified 345,641 protein families. For each family, protein sequences were aligned and used to create a profile hidden Markov model (HMM) via the HMMER software. The resulting profiles for these homologous protein families will facilitate functional analysis of metagenomic sequences and annotation of new microbial genomes.
The precision and recall of each family was statistically assessed by screening each HMM against the protein sequences that were used to generate it. We find that approximately 66.78% of the families have a 100% recall rate and 62.9% have a 100% precision rate. These 47,602 families with "perfect" precision and recall can be reliably used in analyses to classify and annotate metagenomic sequences. We are currently tuning the parameters of the less-than-perfect families to improve their precision and recall.
- Procedure
- Download JGI's Integrated Microbial Genomes sequences (Dec 2009).
- Filter out the sequences that match phylogenetically diverse seed families from Dongying Wu's work.
- Build a percent similarity table using blast on the left over sequences with an 80% coverage threshold to both the query and subject.
- Cluster the sequences feeding the percent similarity table to MCL.
- Merge the newly created families with the seed families : 345,000 families.
- Build alignments and HMM profiles for each family.
- Scan the IMG squences using the HMM profiles for families' recall and precision values.
- Compare the de novo families with Pfam and TIGRfams.
In an effort to reduce computation time, a pre-filter (step 2) was done to remove the sequences that are part of very large phylogenetically diverse families. Automatic updates of the families can very easily be scripted by using the new HMM profiles as seed profiles when going through the pipeline. This approach enables the compute-intensive first four steps to only be run on sequences that do not match any already identified family in future iterations. These future updates will allow us to increase the diversity of the currently described families and to identify new families of sequences. We are currently describing the functional potential of our protein families using Pfam domains and annotations from sequenced genomes. Further analyses will try to identify relationships between our families with the Pfam families.
We are continuing to tune the families to improve the metrics presented above. We are interested in identifying sequences that were too short to be part of the families (i.e., they failed the 80% coverage threshold during the all-vs-all BLAST step) but should actually be included in one of the families. In addition, we are looking into determining relationships between families by comparing the family models and members (protein sequences) to each other. For example, a sub family could cover one domain of a multi-domain family and have been clustered separately.
A manuscript describing this work is in preparation.
- Contributors: G Jospin, T Sharpton, M Langille, D Wu, K Pollard, J Eisen
1.2 Searching for novel phylogenetic types in metagenomic data (Eisen).
1.2a. Develop an automated system for novel branches using rRNA sequences in metagenomic data.
- 100% complete.
We took several different approaches to this problem.
- One approach is described in previous reports and is in the STAP paper by Wu et al.
- A second version is in a paper on "Stalking the fourth domain of life" published in March 2011
- A third version of this is embedded in the PhylOTU method described in a paper in PLoS Computational Biology by Sharpton et al. in 2011.
- Contributors: D Wu, T Sharpton, J Eisen, K Pollard
1.2b. Integrate methods into CAMERA.
- Status: In progress
Methods have been provided to CAMERA for doing this type of analysis (e.g., the STAP software package) but are not yet fully integrated into CAMERA for reasons beyond the control of this project. We note, the STAP software is available for free and links are provided in the paper, and on the iSEEM and Eisen lab web sites.
- Contributors: D Wu, J Eisen
1.2c. Develop an automated system for searching for novel branches for protein coding genes.
- 100% complete.
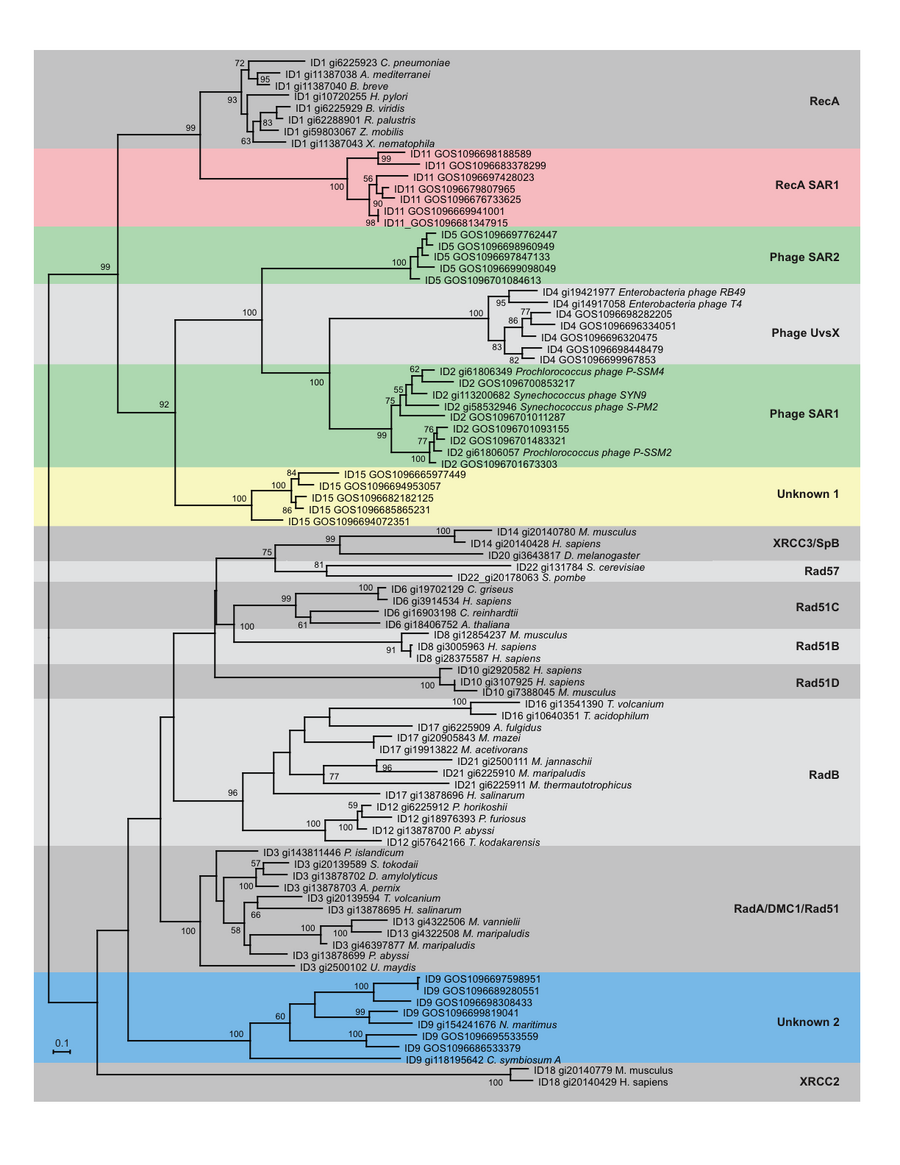
One version of this is described in the Wu et al. "Fourth Domain" paper published in 2011. In this paper, we describe the development of methods to scan through large metagenomic and genomic sequence data sets for sequences that fall into novel deep branches in a phylogenetic tree. We build data sets of all RecA and RpoB genes in genbank to serve as reference sequences. Gene families were built for GOS RecA homologs with RecA reference sequences using BLASTP followed by lek clustering algorithm. Representatives from each family were selected and a maximum likelihood tree was built. Based on the subfamily clusters and the tree structure, we’ve identified 15 major RecA groups. 5 novel deep branching groups included only GOS sequences at the time of initial analysis. With the additional help of neighboring gene studies of the metagenomic assemblies, we identified the novel RecAs as recA SAR1, phage SAR2, phage SAR1, a deep-branching archaea and an unknown group. Similar approaches have helped us identifying two novel rpoB groups as well. We demonstrated that using protein phylogentic markers to analyze metagenomic data is a robust and effective way to identify lineages that are still remains to be discovered. We are excited that two of the novel RecA groups (Phage SAR1 and the deep-branching archaeal group) were subsequently verified by other researchers.
A second version of this is embedded in the work being led by Tom Sharpton described in Section 2.
A third version is a component of the AMPHORA 2 software being developed in the Eisen lab - see below.
Amphora-2
- Status : Preliminary version 100% complete. Improvements needed in precision and functionalities.
Source code available from github.
AMPHORA is an Automated Phylogenomic Inference Pipeline for bacterial sequences (Martin Wu and Jonathan A Eisen. A simple, fast, and accurate method of phylogenomic inference Genome Biology 2008, 9:R151). From a given a set of protein sequences, it automatically identifies various phylogenetic marker genes. It then generates high-quality multiple sequence alignments for these genes and make tree-based phylotype assignments.
Amphora-2 is a complete rewrite of the pipeline in order to improve accuracy, speed and ease of use using 38 new marker genes. The pipeline takes as input nucleotides or amino acid sequences in fastA and fastQ format, and outputs an estimate of organism relative abundance with credibility estimates. One innovation in Amphora-2 is the use of RAPSearch (Yuzhen Ye, Jeong-Hyeon Choi and Haixu Tang. RAPSearch: a Fast Protein Similarity Search Tool for Short Reads. BMC Bioinformatics 2011, 12:159.) for short read sequence homology search, which proves to be 10-100x faster than BLAST or HMMer on short reads (< 1000 residues). Once reads homologous to marker genes have been identified, the reads are aligned to the marker's reference alignment using HMMer. These alignments form the basis for phylogenetic read placement using Pplacer (F.A. Matsen, R.B. Kodner and E.V. Armbrust. pplacer: linear time maximum-likelihood and Bayesian phylogenetic placement of sequences onto a fixed reference tree. BMC Bioinformatics, 2010.). The read placements on each branch of each gene tree are then counted and an estimation of the sample composition is output as a tab delimited text file listing the taxonomy, name, read counts and the percentage of total hits. Unlike the original version, Amphora-2 uses per-gene phylogenetic trees for read placement to accommodate historical lateral gene transfer in the marker genes. Depending on the options used, nucleotide alignments can be generated in addition to the amino acid alignments for each marker.
- sample text file output.
no rank CELLULAR ORGANISMS 43 1.0000
For each sequence placed onto the reference tree, the closest taxon is associated to the sample regardless of branch length. This can result in mistakes in the assignment of reads. We are currently working to develop a confidence value to add to the output for each hit or taxon.
superkingdom BACTERIA 43 1.0000
phylum PROTEOBACTERIA 40 0.9302
family MORAXELLACEAE 36 0.8372
class GAMMAPROTEOBACTERIA 36 0.8372
genus ACINETOBACTER 36 0.8372
order PSEUDOMONADALES 36 0.8372
species ACINETOBACTER SP. ADP1 36 0.8372
no rank UNCLASSIFIED PROTEOBACTERIA 3 0.0698
genus MAGNETOCOCCUS 3 0.0698
The pipeline is being tested using short reads from Illumina lanes on metagenomic data.
- Future work:
- Add a confidence value to all the taxon listed in the output file (Currently being implemented by Aaron Darling).
- Automatically update the marker genes when running new genomes through A2's isolate mode.
- Allow for longer reads to be used through Rapsearch.
- Allow multithreading and/or parallel computing of multiple data sets.
- Contributors: A Darling, G Jospin, J Eisen
1.2d. Identify novel branches for each gene family; Identify other genes linked to genes identified as novel; Develop a Unifrac-like system for identifying the amount of novelty in metagenomic data for a particular gene family; Compare the novelty of phylogenetic marker gene families versus functionally important families; Compare analysis of gene family novelty with metagenomic sample metadata to determine if any types of environments are enriched for novelty.
- Status: 30% complete - work was put on hold until because it was unclear if this could be done reliably.
One way of characterizing functional novelty is to search for novel protein subfamilies that are revealed by analysis of metagenomic data. The most statistically rigorous automated subfamily identification tool produced to date is SCI-PHY (Sjolander), which references a gene family phylogeny to cluster sequences into subfamilies. Unfortunately, the current implementation of SCI-PHY is inappropriate for metagenomic data because it only considers a neighbor-joining tree constructed at run time in a manner ill-equipped for handling short, non-overlapping sequences. We have begun modifying the open source SCI-PHY package to enable its accommodation of our high-quality metagenomic-read inclusive phylogenies. Once completed, we will use SCI-PHY to screen gene family trees (section 2.1a) for subfamilies comprised solely of metagenomic sequences.
- Contributors: T Sharpton, S Riesenfeld, K Pollard, J Eisen
1.2e. Integrate methods into CAMERA.
- Status: Removed as a deliverable
We tried throughout the course of our project to work with CAMERA and to integrate our methods into CAMERA. Unfortunately, CAMERA was unable and/or unwilling to commit to this and thus in consultation with GBMF we removed this as a deliverable from our grant.
1.3 Metagenomic analysis of community phylogenetic structure (Green)
1.3a. Metagenomic community phylogenetic analysis of rRNA genes.
- Status: 100% complete
Using the SSU rRNA gene identification and alignment pipeline described in section 2.1a, we have carried out community phylogenetic analysis based on 16S SSU rRNA reads identified from metagenomic data sets including HOT/ALOHA and GOS. We analyzed this data in the context of the larger community phylogenetic analysis of gene families identified in 1.1a (see section 1.3b below for details). Our analysis shows that SSU rRNA genes and the phylogenetic marker genes described in section 1.1a give concordant results when used to study community phylogenetic structure. This suggests that the use of phylogenetic marker genes with metagenomic data will give comparable results to traditional SSU rRNA gene based analyses of microbial community structure, and will dramatically increase the size of phylogenetic marker gene community data sets that can be derived from metagenomic data. A manuscript by Kembel et al. describing this approach and the HOT/ALOHA analysis is in press at PLoS One.
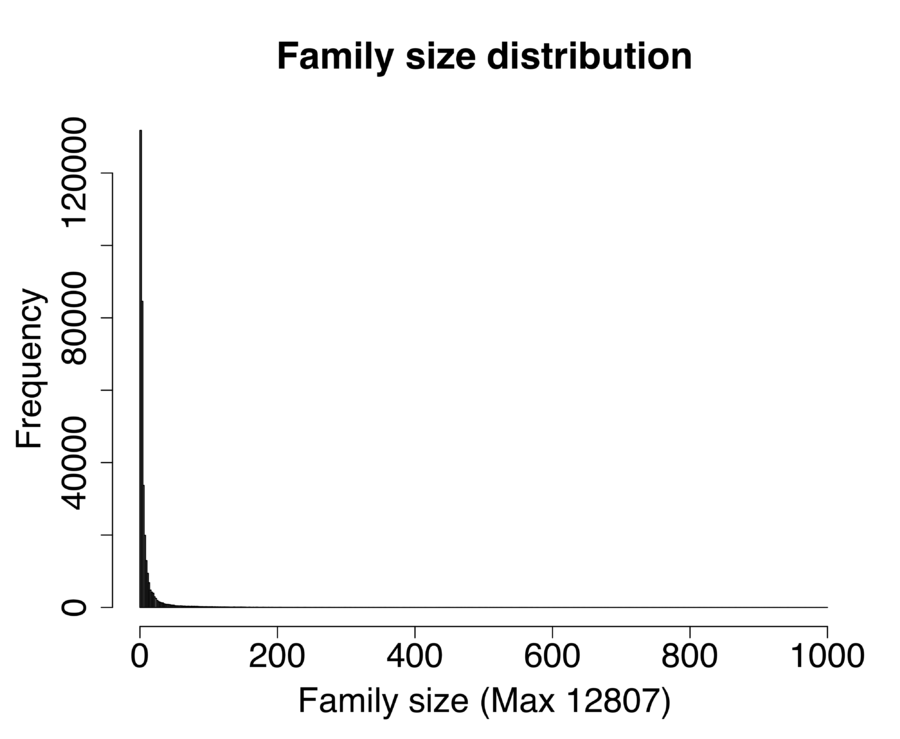
The SSU rRNA gene is widely used in studies of microbial ecology as a barcode to quantify community structure. The principal disadvantage of SSU rRNA genes as a barcode is that genomic rRNA gene copy number varies a great deal across the tree of life (Right Figure). Thus, variation in abundance of SSU rRNA gene copies in environmental samples can be attributed both to variation in the relative abundance of different organisms, and to variation in copy number among the individuals that are present. To improve estimates of community phylogenetic structured based on analysis of the SSU rRNA gene from PCR-based and metagenomic data sets, we developed an algorithm which uses information on phylogenetically related sequences to estimate copy number for environmental SSU rRNA sequences. This algorithm allows community abundance estimates to be corrected to reflect copy number variation (Left Figure), improving the accuracy of estimates of community structure from the SSU rRNA gene. A manuscript and software package implementing this algorithm are in preparation.
- Contributors: S Kembel, K Pollard, J Eisen, M Wu, J Green
1.3b. Metagenomic community phylogenetic analysis of all 50 gene families identified in 1a.
- Status: 100% complete
Analysis of community phylogenetic structure requires a phylogenetic hypothesis for the sequences or organisms living in a community. Construction of phylogenetic trees linking sequences in metagenomic data sets has been limited by the fragmentary, non-overlapping nature of metagenomic reads. We developed a novel approach for community phylogenetic analysis of metagenomic data based on combining the phylogenetic marker gene identification and alignment methods for metagenomic data along with reference data sets of full length gene sequences from fully sequenced microbial genomes described in section 1.1.
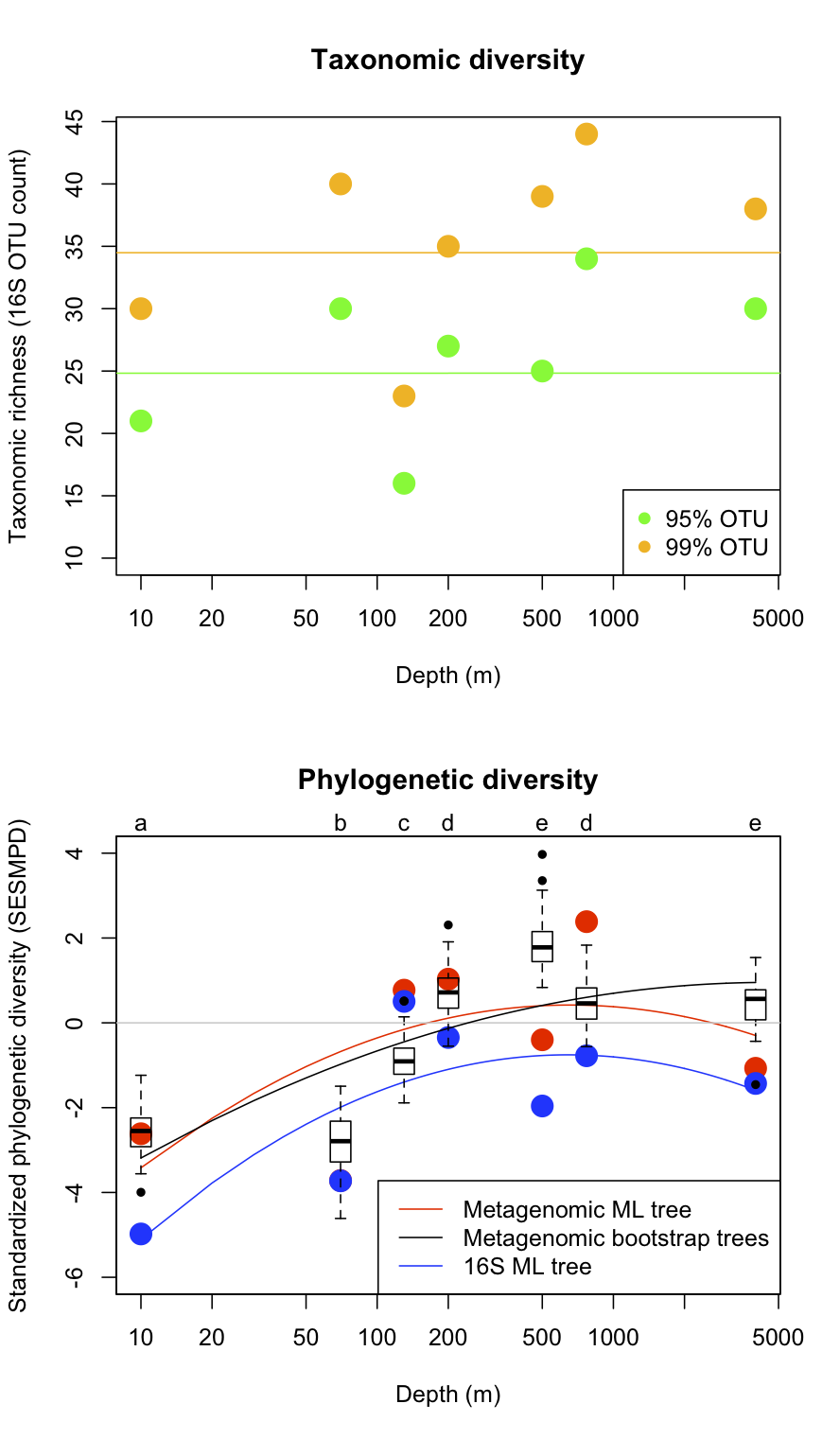
Our initial analyses using this approach focused on describing phylogenetic diversity in bacterial communities in the HOT/ALOHA data set. We quantified phylogenetic diversity and turnover of communities along the oceanic depth gradient sampled by this study based on a combined analysis of sequences from the 31 phylogenetic marker genes included in AMPHORA (section 1.1b). These analyses revealed that while bacterial taxonomic diversity (number of 16S rRNA OTUs) did not vary with depth, bacterial phylogenetic diversity (evolutionary branch length present in communities) peaks at intermediate ocean depths, and that bacterial communities in the photic and non-photic zones contain phylogenetically distinct communities. These results were recently published in in press PLoS One.
We applied this general approach to analyze microbial community structure in the GOS data set on CAMERA based on the 31 phylogenetic marker genes included in AMPHORA. We asked how phylogenetic diversity varies along environmental gradients in the ocean, and the relative importance of space and environment in determining phylogenetic turnover in these communities. Salinity and habitat type (e.g. open ocean vs. coastal waters) are the factors that explain most of the variation in phylogenetic diversity and turnover in oceanic bacterial communities.
- Contributors: S Kembel, S Riesenfeld, K Pollard, J Eisen, J Green
1.3c. Assess utility of gene families from community phylogenetic perspective.
- Status: 100% complete
A challenge of comparing phylogenetic hypotheses for the same metagenomic data set based on different gene families is that the organisms from which sequences were obtained are unknown; as a result it is not possible to directly compare phylogenetic trees for different gene families derived from metagenomic data. We have assessed the utility of gene families from a community phylogenetic structure perspective by comparing estimates of phylogenetic diversity and turnover obtained from the analysis of metagenomic sequences from different gene families, and comparing patterns of community phylogenetic structure based on separate analyses of individual gene families versus a combined analysis of phylogenetic relationships among sequences from multiple gene families.
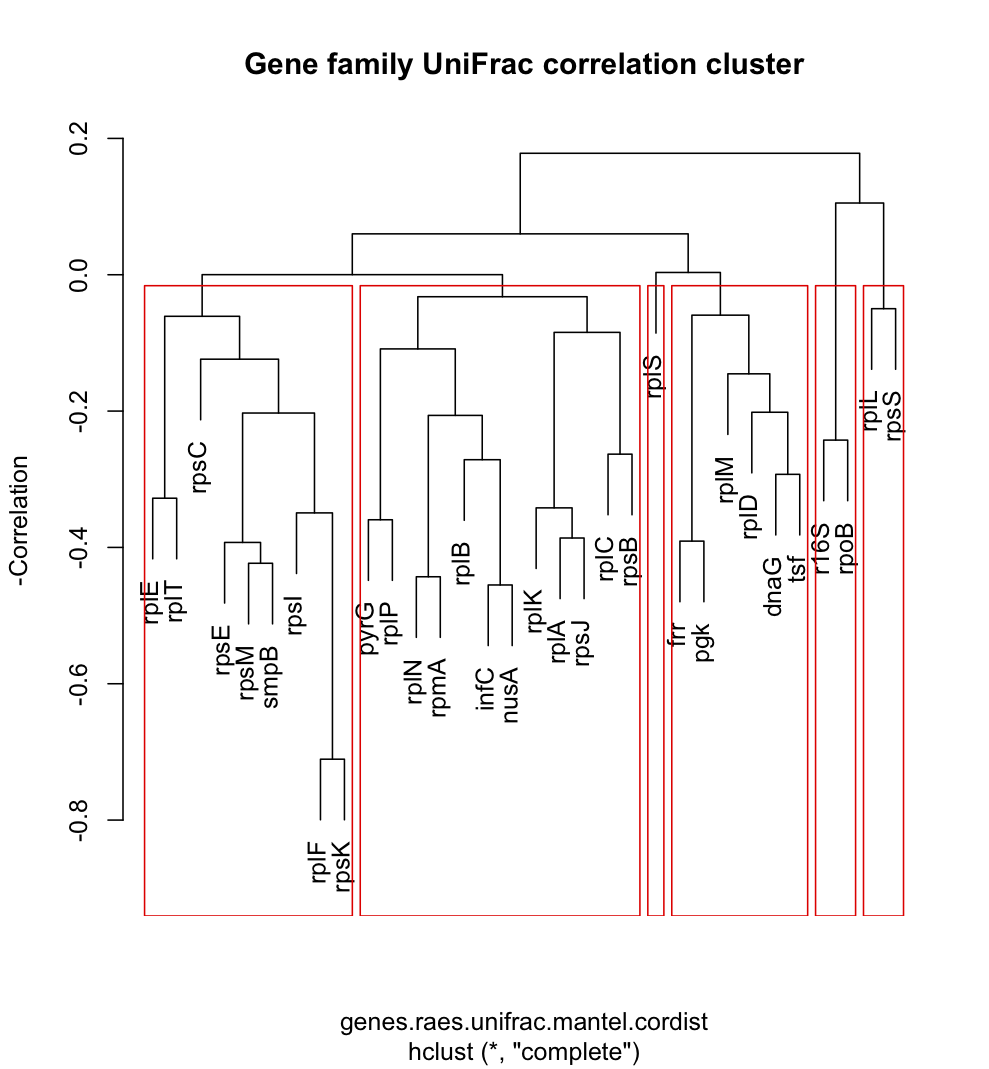
As part of our analysis of phylogenetic diversity in the GOS data set (section 1.3b), we contrasted estimates of phylogenetic diversity of environmental samples based on each of the 31 AMPHORA gene families, as well as for SSU rRNA sequences identified from the same data set (section 2.1a). Analysis of these different gene families gave broadly concordant results, with the phylogenetic diversity of different communities highly correlated regardless of the gene family used to estimate phylogenetic relationships. We identified groups of gene families (Figure Left) that provide the most similar picture of microbial community structure by measuring correlations among phylogenetic beta diversities measured using the 31 AMPHORA gene families. Analyses of the HOT/ALOHA data set indicated that estimates of phylogenetic diversity based on the 16S rRNA gene versus a combined analysis of 31 phylogenetic marker gene families also gave broadly concordant results (see section 1.3a above). These empirical analyses are being combined with simulation studies (section 3.1b) to evaluate the relative utility of different gene families for community phylogenetic analysis.
Our results have demonstrated that by selecting phylogenetic marker gene families that meet a set of criteria including universality and low copy number, we obtain consistent estimates of community phylogenetic structure across multiple gene families, and it is possible to greatly increase the number of sequences from metagenomic data sets that can be used to quantify phylogenetic diversity versus traditional single-gene marker approaches. A manuscript describing the comparison of 31 AMPHORA gene families is in preparation.
- Contributors: S Kembel, S Riesenfeld, K Pollard, J Green
1.4 Estimating biodiversity from metagenomic samples (Green).
1.4a. Assess the fidelity of currently used species (or phylotype) richness estimators.
- Status 100% complete
As noted in our previous annual report, to assess the fidelity of currently used OTU richness estimators, we analyzed the 16S sequence data available on CAMERA for the HOT/ALOHA and GOS data sets. We calculated commonly used OTU richness estimators: the Chao1, Jacknife, ACE, and boostrap. Our results corroborate those reported in Shaw et al. 2008, namely that all statistics yielded qualitatively similar diversity rankings of environmental samples for a given sequence similarity cut-off. Additionally, all the estimators gave qualitatively similar estimates of total richness. In isolation, these results suggest that the majority of OTU richness estimators are useful for ranking microbial diversity among metagenomic samples. But the fidelity of these estimators becomes questionable when considered in tandem with phylogenetic diversity measured across the same samples. Our analysis of taxonomic and phylogenetic diversity in the HOT/ALOHA data set indicates that OTU-based taxonomic diversity estimates did not vary predictably along an oceanic gradient, while phylogenetic diversity showed strong patterns of variation with depth (see Figure in section 1.3b). However taxonomic diversity estimators continue to be a widely used benchmark for comparative analysis of microbial diversity, and this is one reason we focused some of our biodiversity estimator research on taxonomic diversity.
- Contributors: S Kembel, J Green
1.4b. Develop and evaluate novel biodiversity estimators geared toward metagenomics.
- Status 100% complete
A basic question in microbial ecology is the extent to which habitat heterogeneity and dispersal limitation affect large-scale patterns of biodiversity. At one extreme, it is possible that most taxa occur everywhere, while on the other extreme, it is possible that the distributions of taxa are highly constrained by environmental conditions and inability to move between locations. Addressing this question has implications for understanding microbial evolution -– whether microbes are primarily adapted to specific niches or cosmopolitan –- and estimating regional to global microbial taxon richness.
Two characteristics of the distributions of taxa particularly bear on this question, taxa-area relationships and the shapes of ranges. Taxa-area relationships describe how the number of taxa occurring in a region increases with the area of that region. Thus, they can also allow regional or global taxa richness to be estimated. The shapes of ranges describe precisely how taxa are distributed: for instance, do they inhabit circular or highly elongate regions? With dynamical models of colonization, information about range shapes can allow strong inferences about the processes underlying microbial distributions.
Unfortunately, it is challenging to infer directly microbial taxa-area relationships and range shapes because censusing microbes in large regions is difficult. For instance, it is impractical to census all microbes inhabiting large volumes of sea water. However, distance-decay relationships can be constructed for microbes relatively easily. In addition to allowing direct ecological inferences, they can allow taxa-area relationships and range shapes to be inferred. Distance-decay relationships describe how the similarity between between pairs of communities (or samples) decays as a function of the distance between them; naturally, distant communities tend to share fewer taxa than proximate ones. To construct a distance-decay relationship, the taxa in numerous small samples must first be censused. For various subsets of the samples, a measure of similarity is then calculated -- for instance, the number of taxa that the samples share divided by the total number of taxa in the samples -- and this is regressed on the distance between the samples to give the distance-decay relationship. A rapidly decreasing distance-decay relationship suggests strong effects of habitat heterogeneity and dispersal limitation, while a slowly decreasing distance-decay relationship suggests cosmopolitanism.
To make ecological inferences from distance-decay relationships, and infer taxa-area scaling and range shapes from them, a quantitative understanding of how distance-decay relationships arise is necessary. We pursued two complementary approaches to this problem.
The Geometry of Taxa Distributions
One theoretical approach we have taken is focussed on understanding the size and shape of ranges of taxa across a spatial environment, and connecting this understanding to taxa-area scaling. Our theory assumes that the distributions of taxa can be approximated by polygons and disjoint points, and that samples are collected at random locations. Under these assumptions, we have proven that the average number of taxa (i) shared between a pair of samples, (ii) unique one sample in a pair of samples, and (iii) occurring in at least one sample vary as a the sum of a quadratic function of distance and a non-quadratic function, all divided by a function of the shape of the region that was sampled. We have also validated our theory using six large data sets. For these data sets, we have found that the observed distance-decay relationships conform remarkably well to the predictions of our theory.
Our approach for using this theory to infer microbial taxa-area relationships is described in the June 2009 Progress Report. To infer shapes of ranges, it can be shown that the quadratic function for the distance-decay relationship has a geometric interpretation. More specifically, a quadratic distance decay relationship means that the average similarity between pairs of sites is given by a0 + a1*r + a2*r^2, where a0, a1, and a2 are constants ("coefficients"), and r is the distance between the sites. If the distance-decay relationship is considered for just a single taxon, then a0, a1, and a2 can be shown to be proportional to the area, perimeter, and angularity of that taxon's range, respectively. Thus, by fitting the quadratic model to observed microbial distance-decay relationships, we can estimate these attributes of the shapes of their ranges.
A manuscript describing this work (Ladau, Green, Pollard) was submitted to Theoretical Ecology and is currently being revised for re-submission.
Individual-based, Stochastic Community Assembly
The second strand of theory focusses on the dynamics of individuals, taking into account birth, death, speciation and dispersal across a spatial landscape. This approach was described in the September 2009 progress report, and since then the theoretical framework has been published in a high-profile, specialist journal Ecology Letters, building on a framework developed earlier in 2009 and published in PNAS.
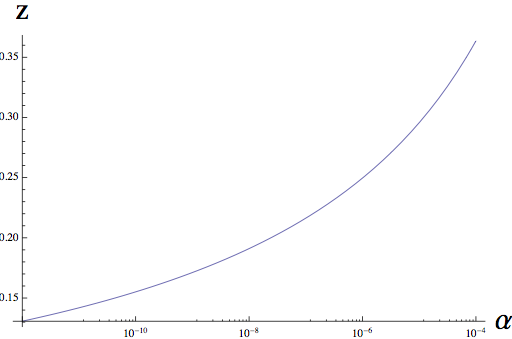
The central results of this paper are: (1) we find a classic power-law relationship over a wide-range of scales, so that taxa richness, T is proportional to a power function of sample area, A^z. This relationship has been observed in hundreds of empirical studies, including both terrestrial and aquatic microbes.
(2) We are able to relate the exponent of this power law, z, to the underlying rate of speciation in a community, alpha, shown below. This same parameter governs distance decay in this model, and so we can use turnover in taxonomic community composition (distance decay) to tightly constrain the form of the taxa-area relationship.
We are currently extending this framework in a number of directions. First, we are taking into account variation of the demographic parameters driving spatial patterns, as these parameters shift with variation in traits and environmental drivers, such as temperature. Adding this additional layer of biological realism may be crucial in applying theory to the full GOS dataset, which now encompasses significant variation in temperature. We are also beginning to integrate into our framework the effect of strong competition among individuals for available resources, to test whether this has a significant impact on large-scale patterns like the taxa-area relationship. Finally, we are seeking to integrate this theoretical framework with the geometrical approach taken by the Pollard lab. We are currently adapting tools from statistical mechanics to relate the dynamics of the shape and size of taxa ranges to the dynamics of the individuals underlying those ranges.
- Contributors: J Ladau, J O'Dwyer, K Pollard, J Green
1.4c. Develop and evaluate biodiversity theory to describe functional diversity.
- Status: 100% complete
The Green lab has contributed in three distinct ways to theoretical frameworks modeling the impact of organism function and environmental variation on patterns of biodiversity.
Impact of Organism Function on Patterns along an Environmental Gradient
James O'Dwyer at the Santa Fe Institute has continued to adapt a set of tools from theoretical physics known as field theory. In earlier phases of the iSEEM project, we explored two distinct applications of these methods: communities structured by body-size (O'Dwyer et al (2009) PNAS) and communities structured by space (O'Dwyer & Green (2010) Ecology Letters).
These applications have focused on how the properties of individual organisms (for example metabolic rate, or dispersal capability) feed into community-level patterns, and we have since been developing theory to include the effects of organism function on these patterns. Working with UO Undergraduate Eric Zaneveld, we modeled the impact of a temperature gradient on organisms with differing, temperature-dependent functional responses. We drew from metabolic theory, which shows a strong dependence of metabolic and demographic rates on temperature, and with Eric we demonstrated that metabolism can drive patterns in latitudinal diversity similar to those observed empirically, documented in his senior thesis.
Stochastic Niche Theory
More generally this approach takes an environmental parameter (temperature in this case) and uses information about organism function to predict patterns along a gradient. A parallel approach is to treat some or all environmental parameters implicitly, so that we look for average expectation values for patterns driven by environmental gradients. We think of this second framework as a stochastic, neutral theory of environmental niches; neutral because we are averaging over many different effects, and as a first step treating species and their responses to environmental gradients neutrally; and niche because we are assuming that patterns are driven by environmental constraints rather than e.g. dispersal limitation, as in our previous work.
We have derived novel results for spatially-implicit models of community assembly. The processes we have built into these models include allopatric speciation, invasion, and environmental stochasticity---in contrast to existing neutral theories, which focus on point speciation, birth and deaths of individuals, and demographic stochasticity, without considering interactions between species or with the environment. We plot an example of the contrasting predictions for relative species abundances in the figure below.
We are currently developing a spatially-explicit version of these models, based on the dynamics of species ranges in space, and using this we will make niche-based predictions for spatially-explicit patterns like the taxa-area relationship and distance decay. We plan to evaluate this model with metagenomic microbial data. The longer-term goal of this project is to combine dispersal and niche processes into a single framework, and to pick out explicit environmental parameters (like temperature, or salinity) when we need to make specific predictions about patterns along those gradients. We also plan to link our results with current empirical work by all three labs to map microbial taxon ranges from sequence data ("niche mapping"; see below), and aim to provide a mechanistic underpinning for iSEEM work in review on the impact of species range size and shape on spatial patterns.
A manuscript describing this project is in preparation.
Correlation analysis of metagenomic data
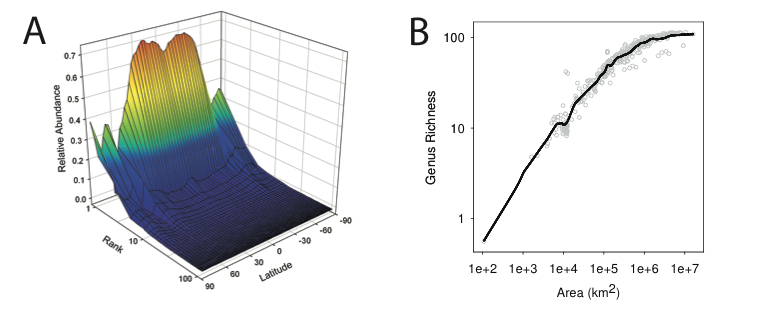
As described in more detail in Section 3.1c and Section 3.1d, the iSEEM project has applied the techniques of ecological niche modeling to infer the ranges of taxa of marine microorganisms. The conceptual basis of the analysis is that certain key environmental parameters select taxa based on their function. The Green lab has contributed to this project by analyzing the Taxa-Area Relationships and Rank-Abundance Distributions predicted by the ecological niche model, providing a contrast with the neutral predictions for these same patterns explored in our earlier papers.
The genus-level taxa-area relationships that we observed are consistent with the power-law form often found and predicted for taxa- area relationships. We note that the genus-area relationships are rather steep (slope =0.39), which may particularly indicate relatively high rates of endemism because our analysis was biased toward widespread taxa. Sequencing depth also influences the slope of the relationship.
A manuscript describing these findings is in review at PNAS. Data and range maps are publicly available on the Pollard lab website.
- Contributors: E Zaneveld, J Ladau, T Sharpton, S Kembel, G Jospin, A Koeppel, J O'Dwyer, J Green, K Pollard
Outcome 1, Output 2: Identify evolutionary dynamics of microbes in nature as illustrated in CAMERA data.
2.1 Molecular evolution of gene families (Pollard).
2.1a. Develop and evaluate molecular evolutionary methods for metagenomics.
- Status: 100% complete
We have undertaken three different projects aimed at rigorously quantifying rates and patterns of sequence evolution.
PhyloP: Statistically evaluating changes in rates and patterns of molecular evolution
New methods and software (PhyloP) for quantifying rates of molecular evolution were described in the 2008 Annual Report. We have subsequently extended these methods from working on DNA sequences to being directly applicable to amino acid sequences, using codon models. We have also implemented models and statistical tests for detecting changes in the pattern, rather than rate, of molecular evolution.
A manuscript describing this project was published in Genome Research (Pollard et al., 2009). All methods are implemented in the freely available PHAST software. This work is a collaboration with Adam Siepel (Cornell University).
PhylOTU: quantifying taxonomic diversity and discovering novel organisms
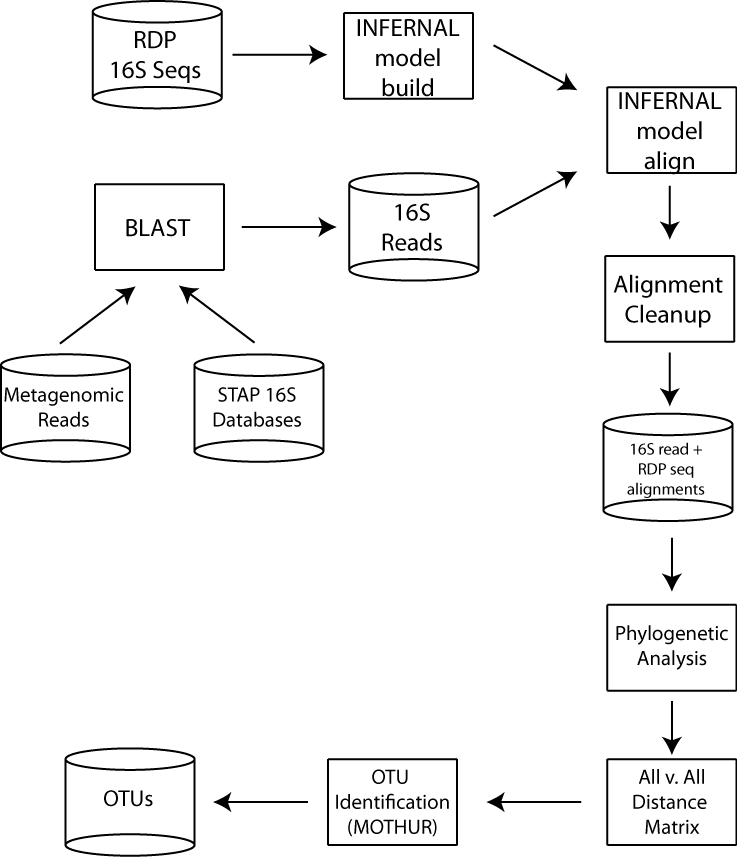 The identification of novel microbial species is great interest to microbial ecologists and evolutionary biologists. To overcome the challenge of taxonomic characterization based on short, non-overlapping metagenomic seequencing reads, we developed PhylOTU, a computational workflow that identifies Operational Taxonomic Units (OTUs, or a corollary for microbial species) directly from metagenomic data. PhylOTU can identify OTUs from metagenomic data with relatively high affinity and can reveal the presence of microbial species that are missed by PCR-based investigations given various methodological biases.
The identification of novel microbial species is great interest to microbial ecologists and evolutionary biologists. To overcome the challenge of taxonomic characterization based on short, non-overlapping metagenomic seequencing reads, we developed PhylOTU, a computational workflow that identifies Operational Taxonomic Units (OTUs, or a corollary for microbial species) directly from metagenomic data. PhylOTU can identify OTUs from metagenomic data with relatively high affinity and can reveal the presence of microbial species that are missed by PCR-based investigations given various methodological biases.
In order to accommodate very large metagenomic libraries, we had to streamline the PhylOTU software in several ways, including implementing more memory/storage efficient distance calculations and enabling the algorithm to be deployed across multiple parallel nodes in a computer cluster. These substantial modifications to the software radically improved its efficiency and yielded a dramatic improvement in PhylOTU's throughput; processing the GOS library (~10 million sequences) now takes several hours, compared to several days.
This work was published in PLoS Computational Biology (Sharpton et al. 2011). We made the PhylOTU source code publicly available here.
Profile based classification of metagenomic reads into protein families
The general goal of this project is to develop a phylogenomic methodology that will classify, or annotate, metagenomic sequences into gene families. Classified sequences will be used to explore biodiversity, discovery novelt, and describe trait-based community assembly.
Phylogenomic annotation is a well documented process, requiring the classification of sequences into homologous families, sensitive and specific family member sequence alignment, and characterization of the evolutionary relationships between members. Unfortunately, all aspects of this generalized process are challenged by the short and fragmentary nature of metagenomic sequence. To circumvent these limitations, we developed a bioinformatic strategy that first references well-annotated whole genome data to characterize known gene families as probabilistic models (section 1.3d) and then uses these models to guide the classification, alignment and phylogenetic analysis of metagenomic sequence. This generalized strategy must be tuned to accomodate the evolutionary properties of the various biological molecules under investigation (e.g., DNA, RNA, protein). To date, we have developed workflows that enable the classification of metagenomic sequence as SSU-rRNA and protein families. The endpoint of the workflow is a relational database that catalogs the presence, abundance, and phylogenetic diversity of each gene family in a metagenomic sample. We evaluated this method's sensitivity and specificity using simulations. Ongoing work aims to tune the parameters of the workflow to maximize the sensitivity of classification.
We are currently leveraging our protein family database and read classifier to analyze marine metagenomic data (e.g., GOS)(section 2.1b).
A manuscript describing this project is in preparation. The protein database and read classifier software will be freely available to the research community.
- Contributors: T Sharpton, J Ladau, S Riesenfeld, S Kembel, J O'Dwyer, J Green, J Eisen, K Pollard
2.1b. Focused assessment of global evolutionary patterns and trends.
- Status: 100% complete
We applied our new molecular evolutionary tools to the GOS and HOT/ALOHA marine microbial data sets to investigate questions pertinent to the characterization of diversity and the discovery of genetic and taxonomic novelty. Additional analyses are ongoing as larger data sets become available. These future projects will include studies of correlations between evolutionary and environmental data.
Taxonomic diversity
We applied PhylOTU (section 2.1a) to the GOS data set. From the 10,133,846 Sanger sequenced reads in the library, PhylOTU identifies 14,320 Bacterial SSU-rRNA homologs, of which 12,020 passed the method’s filters and could be used for OTU discovery. PhylOTU clusters these reads into 833 OTUs at a clustering threshold that is an approximately species level grouping. We also identified 192 Archaeal SSU-rRNA sequences, 79 of which pass the quality control filters and cluster into 7 OTUs.
Comparison of metagenomic and PCR based taxonomic diversity in the GOS data set using PhylOTU
The GOS project also generated 6,413 full-length SSU-rRNA sequences via targeted sequencing of PCR products from six of the 73 geographical sites surveyed. We evaluated the ability of PhylOTU to discover novel taxa in shotgun data by comparing the OTUs identified from metagenomic reads to those identified from the full-length PCR data. This analysis revealed that while PCR libraries identify more OTUs per sequenced residue, metagenomic libraries recover a greater taxonomic diversity of OTUs. By comparing sequences to databases of characterized SSU-rRNAs, we discovered novel species, genera and families in the metagenomic libraries, including OTUs from phyla missed by analysis of PCR sequences, perhaps due to mutations in the "universal" PCR primer sequences for these taxa. Taken together, these results suggest that PhylOTU enables characterization of part of the biosphere currently hidden from PCR-based surveys of diversity.
PhylOTU is also being leveraged by the Human Microbiome Project to identify novel microbial taxa living in and on the human body.
Functional diversity and novelty
Taxonomic diversity, as revealed by the analysis of SSU-rRNA sequences, is only one aspect of biodiversity. The current characterization of protein family diversity, or functional diversity, is relatively poor given that few families can be easily amplified by PCR and few organisms can be cultured for whole genome sequencing. Indeed, to directly survey functional diversity from the environment requires the use of metagenomic sequencing, which in turn requires a means of classifying short, fragmentary metagenomic sequences into their proper families. To meet this need, we developed a metagenomic read classifier (section 2.1a). We estimated a phylogenetic tree for each family and used these to compute diversity statistics across environmental samples.
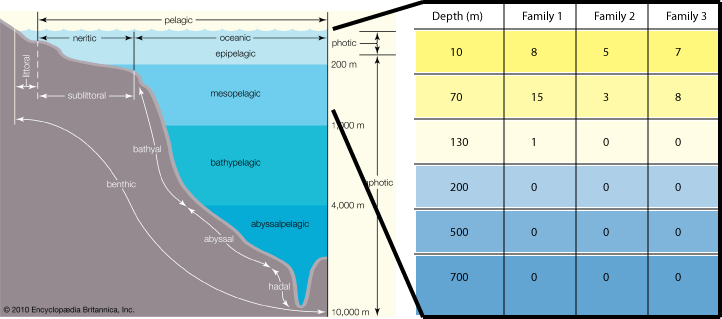
As a test case, we evaluated the performance of our method by searching for photosynthesis-related metagenomic protein sequences along a marine depth column (HOT/ALOHA). Specifically, we screened for sequences homologous to the Photosynthesis Reaction Complex (PhotoRC) families. We expect to find no PhotoRC-related proteins beyond the limit of the photic zone, which is approximately 200 m below the surface of the ocean. As expected, we find PhotoRC-related metagenomic sequences only in samples corresponding to photic zone depths. While this positive control is certainly not an exhaustive test of our method, it provides cause for additional investigation and development.
Ongoing work aims to apply this methodology to the GOS and HOT/ALOHA data sets in order to identify those gene families whose diversity correlates with changes along an environmental gradient (e.g., salinity, temperature). Such families may be important functional components of the communities associated with these particular samples. Another goal of these analyses is to discover novel protein subfamilies represented in metagenomic sequences but absent from microbial genomes that have been sequenced to date. We are also exploring the application of this approach to viral sequence data.
- Contributors: T Sharpton, S Kembel, J Green, J Eisen, K Pollard
2.1c. If fast evolving gene families are reliably identified from metagenomic data, perform detailed investigation of those families.
- Status: 100% complete - project deemed impossible with current metagenomic sequencing read lengths
The goal of this project is to apply PhyloP or similar methods to protein family phylogenies in order to identify families with unusual rates and patterns of sequence evolution in particular environmental sequencing samples. Unfortunately, our simulation studies (section 3.1b) indicate that while protein family phylogenetic trees can on average be constructed with reasonable reliability, individual branches of these trees can be drastically misplaced or distorted (i.e., estimated to have much faster or slower rates of sequence evolution than they should). This result indicates that analyses focused on the most extreme fast-evolving protein families are likely to be plagued by false positives due to limitations of working with metagenomic data. We are exploring the possibility of using our simulation approach to provide protein family specific correction factors that might enable such analyses in the future.
- Contributors: S Riesenfeld, K Pollard
2.2 Population genomics (Eisen, Pollard, Wu).
2.2a. Develop FST like measures of genomic variation within communities versus between communities.
- Status: 100% complete.
This activity was deemed complete because of developments of others including work of Steven Kembel in prior studies as well as work from the metagenomics field.
2.2b. Develop methods to quantify insertion-deletions, recombination, and rearrangement by comparison to reference genomes
- Status: 100% complete
This activity was deemed complete because of developments of others including work of Jill Banfield as well as work from others in the metagenomics field.
2.2c. Develop the genomic x spatial species concept for microbes
- Status: 100% complete
This work was modified slightly to focus on the "ecotype" concept in Martin Wu's lab.
Wu Lab, University of Virginia
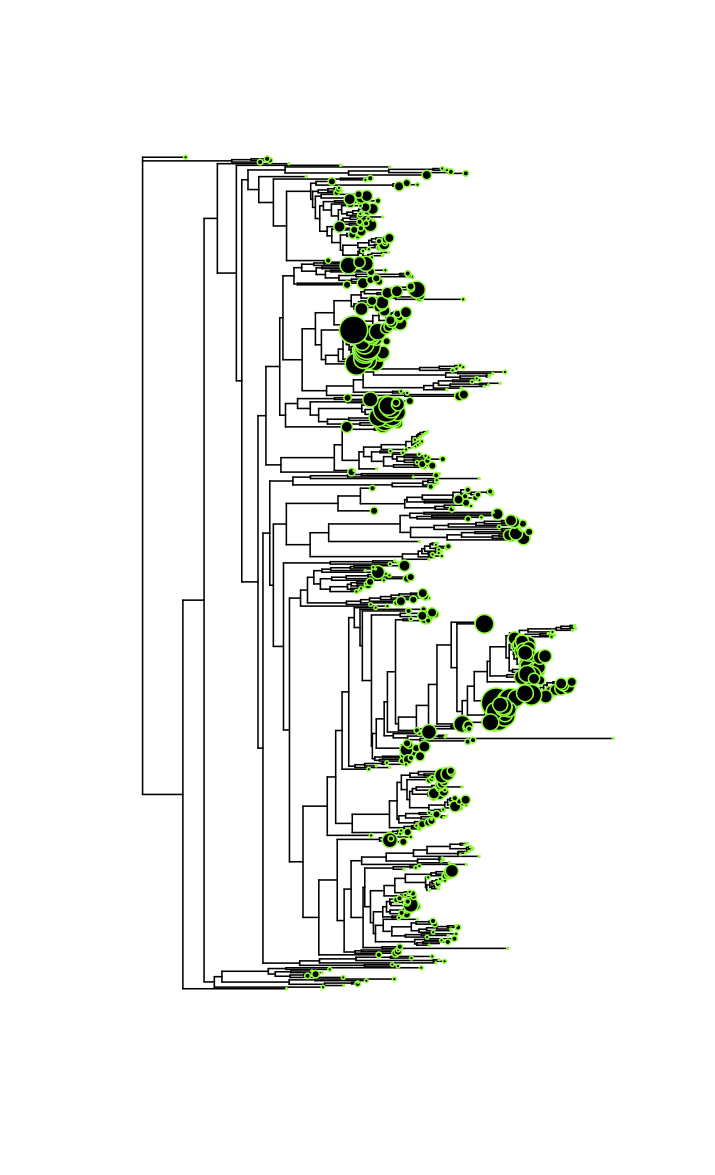
We studied the question is at what taxonomic level bacteria display ecological coherence, i.e., they share similar lifestyles or traits that distinguish them from members of other taxa. Ecological coherence exists where most members of a taxon share ecological traits and therefore exhibit similar responses to the environment. Bacterial communities are commonly characterized based on the relative abundances of higher order taxa, but the ecological coherence of these taxa has not been extensively studied.
Recent studies have shown evidence of habitat association among higher bacterial taxonomic ranks, but when studying higher order taxa in bacteria, it is not always clear whether their habitat associations reflect the ecology of most, or even many of the species contained within them. It is possible that a few highly abundant species could make it appear that the taxon as a whole was strongly associated with a particular habitat, even if the majority of the species within it had no such association. Interpreting the results of habitat association studies requires a better understanding of how uniformly associations among higher order taxa are reflected among the species that make them up. However, the ecological coherence of taxa above the species level has not been explicitly tested , leaving open the question of how best to characterize and compare the diversity of bacterial communities.
We investigated the depth of ecological coherence in the bacterial taxonomic hierarchy. We chose to analyze sequence data from the human microbiome, which we believe to be an excellent system for addressing this question for two reasons. First, examining the depth of ecological coherence in bacteria has clear implications for ongoing human microbiome research and second advantage because of the availability of deeply sequenced datasets.
Using two microbiome datasets, one from human skin, the other from human gut, we investigated the depth of ecological coherence by testing whether patterns observed at higher taxonomic ranks are maintained at lower ranks. We performed a comprehensive test by comparing the habitat associations along the entire hierarchy, from the phylum level all the way to species and subspecies (as approximated by operational taxonomic units (OTUs)).
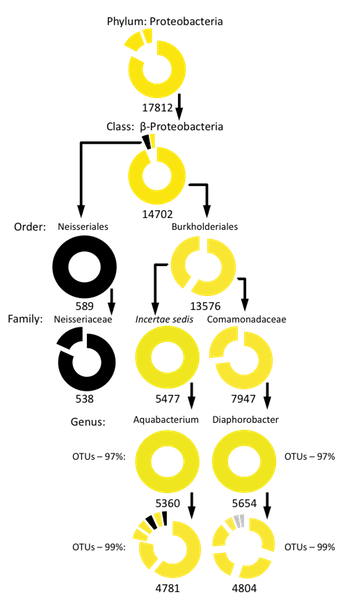
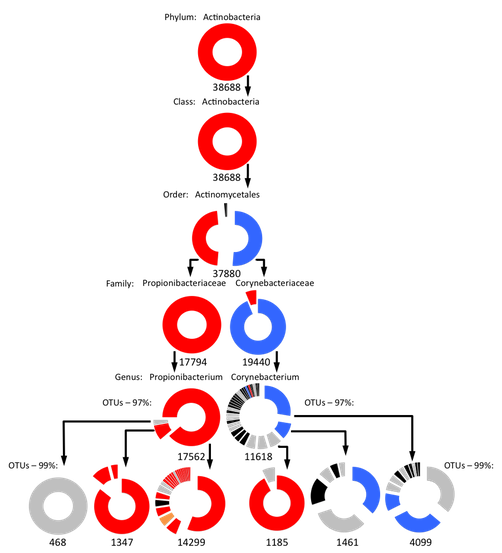
Hierarchical analysis of skin habitat associations in the phyla Actinobacteria, Firmicutes, and Proteobacteria
- Each wedge in the doughnut graphs represents a subtaxon (with at least 100 sequences) present in the labeled taxon, with the wedge size corresponding to the subtaxon’s relative abundance. Subtaxa are sorted clockwise by their relative abundance. Examples: In the first graph (Actinobacteria) the doughnut for the whole phylum (top) is an unbroken circle because all of the sequences in the phylum belong to one single class. The graph for the order Actinomycetales is split into four wedges because there are four families in that order containing at least 100 sequences. Two of those families (the Corynebacteriaceae and Propionibacteriaceae) are much more abundant than the other two and so are represented by larger wedges. The wedges of each graph are color coded to reflect the habitat associations of the taxa represented. Color-coding is as follows: Red-- Significantly associated with sebaceous skin; Yellow--Significantly associated with dry skin; Black-- ANOVA showed no significant associations; Grey-- ANOVA showed a significant association, but the Tukey-Kramer test showed no positive association to any particular skin type. Blue-- Significantly associated with only moist skin; Purple-- Significantly associated with both moist and sebaceous skin (i.e. underrepresented on dry skin); Orange—Significantly associated with both dry and sebaceous skin (i.e. underrepresented on moist skin); Green—Significantly associated with both dry and moist skin (i.e. underrepresented on sebaceous skin). Numbers below each graph show the number of sequences represented.
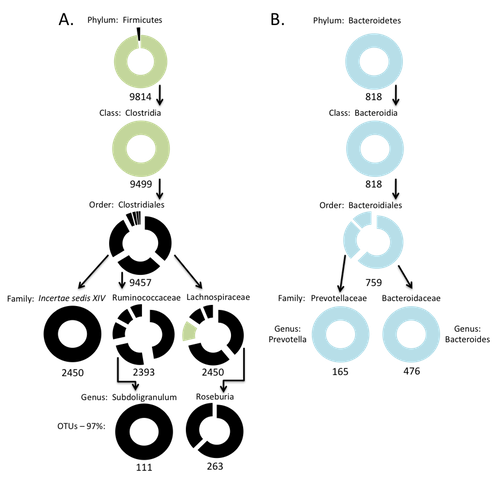
Hierarchical analysis of gut habitat associations in the phyla Firmicutes and Bacteroidetes.
- The gut dataset is coded as follows: Light Green-- Significantly associated with obese subjects; Light Blue--Significantly associated with lean subjects; Black-- ANOVA showed no significant associations.
Our hierarchical analysis of habitat associations revealed four broad patterns, described below, and in the accompanying diagram:
- Habitat association of a taxon is maintained among all or nearly all of its constituent subtaxa (examples: the associations of the phylum Bacteroidetes with lean subjects, the order Burkholderiales with dry skin and the family Propionibacteriaceae with sebaceous skin).
- Parent taxon has a significant habitat association but none of its subtaxa have any associations at all (examples: the associations of the order Clostridiales with obesity and the family incertae sedis XI with non-sebaceous skin).
- Habitat association of a taxon is shared by only a few of its constituent subtaxa. Most subtaxa have either no association or their associations are different from that of the parent taxon (example: the association of the genus Corynebacterium with moist skin).
- Parent taxon has no habitat association but many of its subtaxa do have significant habitat associations (examples: The class Bacilli, which itself has no association to any skin type, but contains numerous OTUs that are associated with either moist skin, or sebaceous skin, or both; Several species in the genera Corynebacterium and Staphylococcus who themselves have no associations but contain 99% OTUs which do).
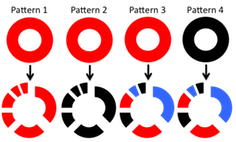
Schematic diagram of four patterns of habitat associations.
- Graphs in the top row represent higher order taxa, while the bottom row shows their division into taxa of the next lower rank. Coloring of graphs represents associations to hypothetical habitats. Red-- Associated with habitat 1; Blue—Associated with habitat 2; Black—No habitat association.
With the entire clade sharing the same habitat association, we argue that pattern 1 taxa are ecologically coherent. It is by no means our intention to suggest that members of pattern 1 taxa are homogeneous over all aspects of their ecological niche, only that they display coherence with respect to certain specific ecological dimensions relevant to the habitat with which they are associated. Pattern 1 taxa may possess traits originating deep in the history of their lineage that confer similar ecological properties, resulting in habitat filtering. We found evidence of coherence above the species rank, at the phylum (e.g., Bacteroidetes), order (e.g., Burkholderiales) and family (e.g., Propionibacteriaceae) levels.
From a community perspective, we argue that pattern 2 taxa could also possess ecological coherence. An ecological community requires the presence of a certain complement of genetic and metabolic functions in order to be stably maintained. If a certain metabolic function is a synapomorphy (a derived character inherited through vertical descent) of a clade, then any member of that clade present within the community might be sufficient to carry out that function, and which particular subtaxon it is may not be relevant. The subtaxa would be interchangeable with respect to that particular ecosystem, even if they were ecologically distinct in other respects. We hypothesize that competitive exclusion may be a more important process than habitat filtering in pattern 2 taxa. Species in competition with one another at a particular habitat could be associated with the same habitat type, yet only one or a few species could be dominant in that habitat on any particular subject. Consequently, the within-habitat variation in abundances would be high for each species, making it more difficult to detect statistically significant habitat associations among species, but easier to detect them among higher order taxa. Individual species might be perfectly capable of thriving in a given habitat if only abiotic factors were considered, but inter-specific competition can mask that association by limiting the number of subjects upon which the species in question is found in abundance at that habitat.
We would argue that taxa displaying pattern 3 or 4 are not ecologically coherent because the habitat associations of the subtaxa are heterogeneous. Within these taxa, transitions between habitats among species seem to have taken place relatively recently. This could indicate that the traits that promote or allow associations to particular skin habitats are evolutionarily labile in these taxa, allowing them to make the transition between habitat types relatively easily and frequently. In pattern 3, the parent taxon's habitat association mostly reflects that of certain highly abundant subtaxa, and not necessarily the majority of its members. Although heterogeneity was observed at different levels of organization, in both high and low taxa, it is of particular interest to note the apparent incoherence in some species. For example, species within both the Corynebacterium and Staphylococcus genera exhibiting no specific skin type associations nonetheless contained subspecies that did have significant associations.
These findings raise a challenge to the methods by which communities of human microbiota are currently characterized in association studies. Such studies generally compare communities based on the relative abundances of taxa at one level, usually at or above the genus. Our results suggest that the choice of taxonomic level used to define the operational unit is a very important factor to consider. To truly understand whether and how the composition of microbiome affects human health, we might need to dissect the biological patterns along the entire taxonomic hierarchy, including the very fine scales of microdiversity. Our results demonstrate that in order to get a complete picture of the structure of a bacterial community, more comprehensive taxonomic analyses are necessary. Demonstrating associations of higher order taxa to a specific habitat may or may not be informative depending on whether or not that taxon is an ecologically coherent unit with respect to the environment being studied.
A manuscript detailing these results is in preparation.
2.2d. Develop phylogenetic sliding window approach for determining if multiple populations are present in a single bin.
- Status: 50% complete
The Eisen lab (especially Aaron Darling) have been working on this implementation. It was deemed more complicated than we originally imagined and Darling and Eisen have now received separate funding from DHS to develop this "cobinning" approach.
- Contributors: J. Eisen and A. Darling
2.2e. Estimate effective population (Ne) sizes for different microbes and design of methods to detect community-level bottlenecks that may make communities vulnerable as seen in endangered species.
This general issue has been integrated into the ecotype analysis described in 2.2c above.
2.2f. Correlation analysis on general patterns of “evolvability” such as mutation rates, population size, and recombination patterns with community characteristics.
This general issue has been integrated into the ecotype analysis described in 2.2c above.
Outcome 1, Output 3: Develop statistical methods to correlate metagenomic sequence data with environmental metadata.
3.1 Metagenomic analysis of community phylogenetic structure (Pollard).
3.1a. Survey metagenomic data to determine scope of data types.
- Status: 100% complete
A preliminary step of the iSEEM project was to use CAMERA to assemble information about existing metagenomic data. We compiled information on the types of metadata that are available for each project on CAMERA by consulting (i) individual metadata files for each project (on each project's webpage) and (ii) the listing of metadata on the CAMERA Project Samples webpage. We also gathered information on sequence data for each project by consulting files available for download on each project's webpage, published papers for each project, and the File Server Download page. For additional details, please refer to the 2008 Annual Report.
We continue to monitor CAMERA for new data sets and to explore other sources of data.
- Contributors: J Ladau, K Pollard
3.1b. Develop a metagenomic simulation pipeline; use simulated data to evaluate methods in other sections, such as 1.1 (trees), 1.3 (diversity estimators), and 2.1 (evolutionary rates).
- Status: 100% complete
MetaPASSAGE Workflow
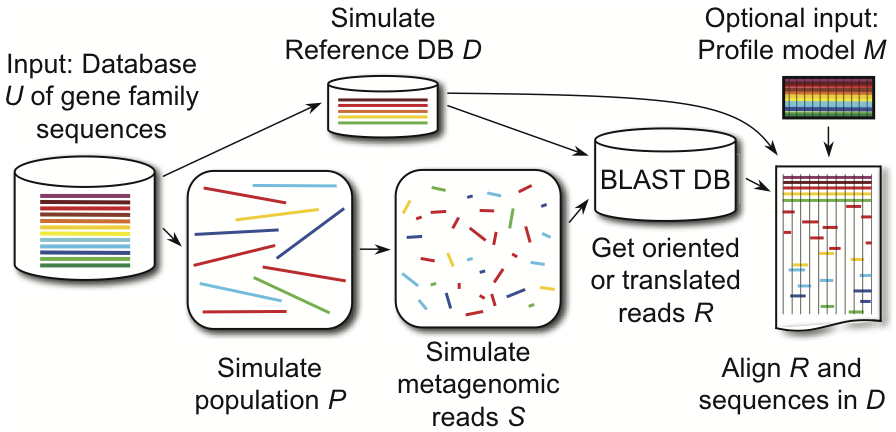
Simulation-based tools are essential for assessing the accuracy of emerging computational methods that account for the complexity of the community sampled, sampling depth, and sequencing technology. We developed MetaPASSAGE, a fully automated analysis workflow for generating simulated metagenomic libraries and producing reliable alignments of sequencing reads belonging to individual gene families, including SSU-rRNA and AMPHORA proteins (Wu and Eisen, 2008). Our workflow integrates the simulation program MetaSim (Richter et al., 2008) with packages for data processing and alignment via flexible Perl scripts and modules, enabling batch processing and bulk statistical analysis. MetaPASSAGE will have a major impact on bioinformatics tool development for metagenomics, facilitating new discoveries from the vast volume of sequencing data being generated.
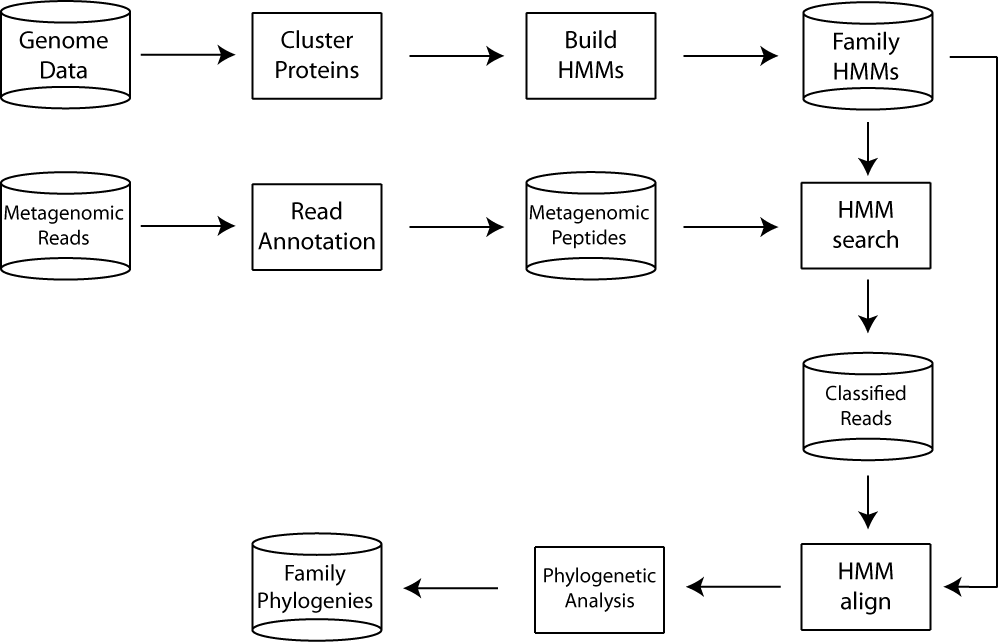
We take a gene-family-focused approach to metagenome simulation, reflecting the fact that assembly of whole chromosomes from metagenomic data is currently impractical for complex communities. Along with MetaSim, the MetaPASSAGE workflow incorporates several well-known programs, including NCBI BLAST, HMMER, and INFERNAL (Nawrocki et al., 2009), into a streamlined, completely automated tool to perform simulations in batch at the command line. The major steps in the workflow are summarized in the diagram. The workflow also interfaces with AMPHORA (Wu and Eisen, 2008), a software package for performing phylogenomic inference with metagenomic sequences belonging to housekeeping protein families.
A paper about this software was reviewed at Bioinformatics. We are current revising the manuscript for re-submission. The MetaPASSAGE software is publicly available at https://github.com/sriesenfeld/MetaPASSAGE/.
Simulations reveal the accuracy of phylogeny-based analysis of metagenomic sequences
Using simulations to evaluate phylogenetic methods on metagenomic sequence data
Phylogenetic methods are critical for many kinds of genomic analysis. Building phylogenetic trees from metagenomic sequences is a way of moving beyond simple sequence comparison to constructing models for microbial community evolution. Taxonomic trees built from the SSU-rRNA gene can be used to understand the amount of taxonomic evolution and diversity that is unique to each community. Phylogenetic analyses of protein families from different communities may reveal much more about the evolution of functional capabilities than taxonomic analyses can reveal.
Phylogenetic analyses of metagenomic data have typically been based on targeted sequencing. It is much easier to build trees from long, aligned sequences and for a well-characterized gene family, such as SSU-rRNA.
This project analyzed how different phylogenetic methods may be expected to perform on metagenomic shotgun sequence data for several different gene families. We designed a set of simulations to test the feasibility and accuracy of building metagenomic phylogenies. We specifically assessed the independent and interacting effects of (1) phylogenetic inference algorithm (i.e., tree building method), (2) sequencing read length, (3) number of reads, and (4) prior knowledge about the gene family (e.g., as encoded in the reference database used in the MetaPASSAGE workflow or to build a "guide tree" to aid phylogenetic inference).
Simulations Framework: The simulated reference database provides a way of modeling the extent that annotated full-length gene sequences represent an entire gene family. We expect the extent of this coverage to influence the ability to build an evolutionary model using those known sequences and new metagenomic sequences. For each family, two sizes of reference databases were modeled: 50 and 200 sequences.
Less phylogenetic signal is expected to be present in general among short reads, versus full-length sequences. We used MetaPASSAGE to simulate metagenomic reads of two different mean lengths: 100bp and 400bp.
More error is also expected to be present when the phylogeny includes more branches that are tips (i.e., more sequences). We simulated sets of 50 or 200 sequencing reads per protein family, to test how much the number of reads affected performance.
The phylogenetic algorithms were selected in order to represent different approaches that can feasibly be used for large-scale phylogenies: FastTree (a very fast heuristic), RAxML (a maximum-likelihood-based method), and Pplacer (a maximum-likelihood-based algorithm designed specifically for metagenomic data).
To test whether accuracy varies for different gene families, we identified five gene families that are all well-enough characterized to be useful in simulations, including SSU-rRNA, the long, well-conserved RpoB protein from the AMPHORA database, two additional shorter protein families from AMPHORA, and lolC, which is a protein family related to ABC transporters. Each gene family has at least 400 unique full-length gene sequences in current databases. As a measure of quality control, we used families for which alignments and probabilistic models had already been built.
Each simulation consisted of the following steps: (1) We used MetaPASSAGE to produce a distribution of reads across sequences and then automatically orient (in the case of SSU-rRNA) or translate (in the case of protein families) the reads with BLAST, dropping any reads for which this could not be done accurately. (2) The final set of reads was then processed using MetaPASSAGE so that there remained at most one read per original full-length sequence. This step was taken to facilitate direct comparison with phylogenies built from full-length sequences. (3) The final read set was then aligned with MetaPASSAGE to one of the simulated reference databases, and this alignment was given as input to the different phylogenetic algorithms. (4) The tree output by each algorithm was pruned to remove the tip branches corresponding to reference sequences, so that the set of tips was labeled exactly by the set of simulated reads in the input.
To evaluate the accuracy of the output tree for each simulation, we compared it to a tree built as follows: Using the same phylogenetic algorithm, a tree was constructed from the entire set of full-length gene sequences for that gene family. It was then pruned so that it contained only the tips labeled by full-length sequences that correspond to the simulated reads in the tree being evaluated. This comparison is illustrated in the accompanying figure. We used several measures of error to evaluate performance, including the normalized Robinson-Foulds, which computes the percentage of edges that occur in one or the other of the trees but not in both (a measure of topological accuracy or whether branches are in the right place), and distortion, which measured the factor by which edges in the read tree were stretched or shrunk compared to the corresponding edges edges in the tree built from full-length sequences (a measure of branch length or evolutionary rate accuracy).
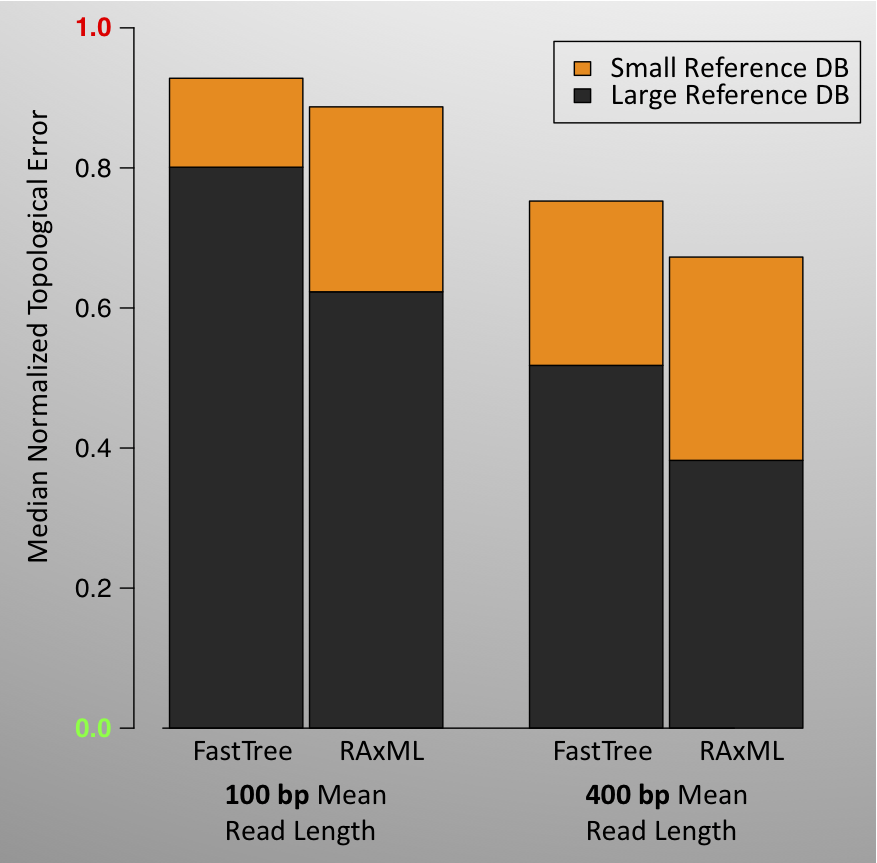
Results: Our results show that the size of the reference database (i.e., how much is already known about the gene family) can have a profound effect on our ability to build a phylogeny of metagenomic reads. This makes sense because all of the phylogenetic methods we evaluated leveraged the reference database in some way. According to all measures of error and in nearly all scenarios, the larger reference databases, which contains 43¬–48% of the sequences in a gene family, result in significantly less topological error and branch length error than do smaller reference databases, which contain 11-12% of sequences. RAxML and pplacer are particularly able to capitalize on the larger reference databases, showing proportionally greater decreases in error than does FastTree as the size of the reference database increases.
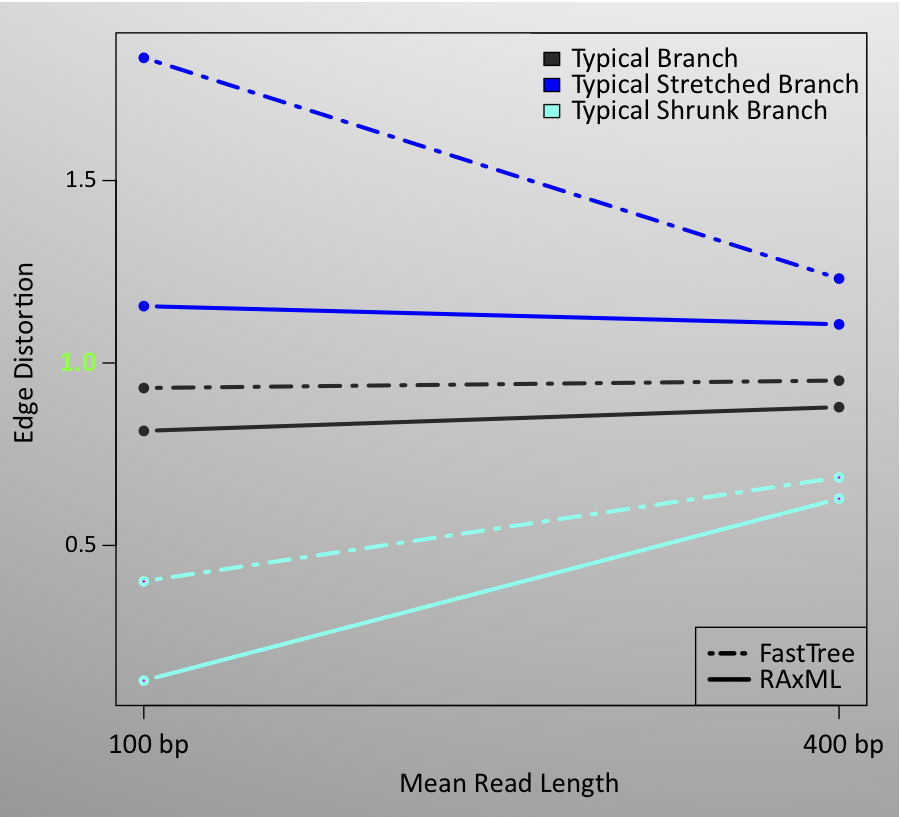
Read length also proved to have a great effect on the accuracy of topological and branch-length inference in all simulated scenarios. Shorter reads result in much more error, regardless of other simulation parameters. In many scenarios, increasing the read length cut the topological error by 25–30%. The improvement in distortion due to longer read length is particularly dramatic in the scenarios that combine a large number of reads with a small reference database. Generally, the read length appears to affect the distortion of tip branches much more than the internal branches.
The accompanying figures demonstrate some of the results for topological error and distortion, for the SSU-rRNA gene family. Even with the normalization of topological error, which puts less weight on an erroneous edge when there are more tips in the tree, we saw increased topological error and branch-length distortion error for trees with larger numbers of reads. However, the effect is only dramatic in some situations, for example, when the read length is short. Qualitatively similar results we obtained with other gene families. Median topological error is highest with lolC, especially in scenarios involving a small number of reads, but this may be due to less quality control on the data or model for lolC. Branch distortion varies somewhat across gene families.
The results of these simulations indicate that an individual branch in a metagenomics-based phylogeny should not be assumed to be accurate. However, they also indicate that a significant fraction of the edges are correct, or reasonably well-estimated, implying that downstream methods that aggregate information from the entire tree may be reasonably accurate and useful for metagenomic analysis.
Enterotype Simulations to Evaluate UniFrac: Following up on the simulation results, we designed additional studies to evaluate the performance of Fast UniFrac and weighted Fast UniFrac, which are widely used in microbial ecology to distinguish communities by assessing the amount of evolution that is represented uniquely by one community or another.
FastUnifrac can be used with metagenomic phylogenies to identify and distinguish enterotypes.
A recent study of the human gut microbiome [Arumugam et al., Nature 2011] identified three different types of gut communities, which were called enterotypes, according to the species in which they are enriched. The authors applied a clustering method to Sanger-sequenced reads sampled from gut microbiomes. When they authors applied their clustering methods to data from other studies, they found some variation in the enterotypes. They attributed this, in the first case, to a different set of reference genomes being used, and in the second case, to a different sequencing method (Illumina).
Our follow-up simulations were designed to investigate whether Unifrac could be used with shotgun-sequence-based metagenomic phylogenies to identify three enterotypes from simulated gut samples, and how the read length affected the performance.
We simulated the three types of communities using either long reads or short reads, and then built trees with the samples using either RAxML or FastTree. Then we looked to see how often Fast UniFrac could distinguish the communities, based on the input tree. As a measure of the false positive rate, we also iteratively generated three samples from a single community, and built trees from these samples. The plot here shows a ROC curve, with the true positive rate on the y-axis and the false positive rate on the x-axis. UniFrac does very well, even in the worst case of trees built with FastTree from short reads, although the performance is clearly better in other scenarios.
This result is encouraging, although the problem analyzed is not a particularly challenging identification problem, as the three enterotypes are distinguishable based on the presence or absence of a few taxa. Future work will examine the downstream effects of phylogeny estimation errors on microbial community analyses that leverage gene trees.
We are currently preparing a manuscript describing this project.
- Contributors: S Riesenfeld, K Pollard
3.1c. Develop and evaluate methods for correlation analysis of metagenomic data.
- Status: 100% complete
We modified and utilized an ecological method, called ecological niche modeling, to infer the ranges of taxa of marine microorganisms at a global scale. This approach is appropriate for relatively sparse sampling data (e.g., museum samples, marine microbial DNA sequencing surveys) where information is available about presence (but not absence) of taxa at different locations across environments. Ecological niche modeling then leverages environmental data to predict the ranges of taxa based on the types of environments in which they have been observed. Since global environmental data (e.g., mean annual sea surface temperature, nutrient concentrations, salinity) are are available at a high degree of resolution across the World Ocean, this approach enabled us to predict the ranges of marine microbes. To do so, we used the RDP classifier to identify microbial taxa (e.g., bacterial genera) at locations where sequencing surveys have been conducted (data obtained from publicly accessible databases, such as CAMERA, MICROBIS). Next, we used statistical methods (e.g., multinomial logistic regression, machine-learning methods) to model the relative abundance of each taxon as a function of environmental variables. Finally, we inferred the range of each taxon by using these models to project the ecological niche of each taxon to geographic coordinates across the World Ocean (all locations where environmental data are available). These predictions cover many locations where direct observations of the presence of taxa are unavailable, enabling us to study the global distributions of marine microbes using the relatively sparse sequencing surveys currently available.
A manuscript describing this methodology and its application is in review at PNAS.
- Contributors: J Ladau, K Pollard
3.1d. Apply correlation analysis to publicly available data.
- Status: 100% complete
Little is known about the distribution of bacterial diversity in the ocean at global scales, or about the drivers of bacterial biogeographic patterns. To meet this challenge, we compiled a global dataset of marine SSU-rDNA sequences and then use ecological niche modeling (see section 3.1c). Our results (i) predict global bacterial ranges for the first time and (ii) provide the most extensive global snapshot of bacterial biogeography to date.
Biogeographic studies of the ranges of macroorganisms in the oceans have revealed consistent patterns of biodiversity with respect to latitude and environmental conditions (e.g., Tittensor et al., 2010, Nature, 466: 1098–1101). But we have limited knowledge of the corresponding patterns in microorgisms. The major impediment to such investigations is limited data about the occurrence patterns of different microbes across the world ocean. While genomic studies have greatly accelerated the collection of such data, sampling locations are still much too sparse to enable direct measurement of microbial ranges. To address this problem, we assembled a database of marine SSU-rDNA sequencing studies and identified all known bacteria genera at each sampling location. Then, we leveraged global environmental data and niche modeling methodology to predict the global ranges of all prevalent bacteria genera at 1° resolution. Our spatially explicit range maps for hundreds of bacterial genera enabled us to quantify patterns of marine bacterial diversity across the globe. We discovered low microbial diversity in the tropics and high diversity in areas where ocean currents meet and mix (see figure below). We documented significant differences in biogeographic patterns between individual genera and across the major marine bacterial phyla. These findings contributed significantly to the ongoing discoveries in bacterial biogeography, which indicate that microbial biogeography may be driven by a fundamentally different set of rules when compared to the patterns observed for macroorganisms (e.g., Zhou et al., 2008, Proc. Natl. Acad. Sci. U.S.A. ,105: 7768 –7773).
Our carefully constructed data sets, range maps, and diversity estimates provide a solid basis for launching the emerging field of marine microbial biogeography.
A manuscript describing these findings is in review at PNAS. Data and range maps are publicly available on the Pollard lab website.
- Contributors: J Ladau, T Sharpton, S Kembel, G Jospin, A Koeppel, J O'Dwyer, J Green, K Pollard
2. Publication and Patents
Publication list
In preparation
- Kembel SW, Wu M, Eisen JA, Green JL. Phylogenetic signal in 16S copy number allows improved quantification of microbial diversity. In preparation.
- Riesenfeld S, Pollard KS. Can phylogenies be reliably constructed from metagenomic data? In preparation.
- Kembel SW, Eisen JA, Pollard KS, Green JL. Phylogenetic beta diversity measured from metagenomic data: comparison of different gene families. In preparation.
- Jospin G, Sharpton TJ, Wu D, Langille MGI, Pollard KS, Eisen JA. Automated identification and annotation of full-length gene families from bacterial genomes. In preparation.
- Sharpton TJ, Eisen JA, Pollard KS. A read classifier for characterizing protein functions in metagenomic read libraries. In preparation.
Submitted
- Ladau J, Sharpton TJ, Jospin G, Kembel SW, Koeppel A, O’Dwyer J, Green JL, Pollard KS. The Global Biogeography of Marine Bacteria. In review at PNAS. Open access.
- Ladau J, Green JL, Pollard KS. Beta Diversity Follows a Universal Model. Reviewed at Theoretical Ecology. In revision.
- Riesenfeld S, Pollard KS. MetaPASSAGE: A Metagenomic Pipeline for Automated Simulations and Analysis of Gene Families. Reviewed at Bioinformatics. In revision.
- Ayres D, Darling A, Zwickl D, Beerli P, Holder M, Lewis P, Huelsenbeck J, Ronquist F, Swofford D, Cummings M, Rambaut A, Suchard M. BEAGLE: an application programming interface and high-performance computing library for statistical phylogenetics. Accepted pending revisions, Systematic Biology.
- Ronquist F, Teslenko M, van der Mark P, Ayres D, Darling A, Hohna S, Larget B, Liu L, Suchard M, Huelsenbeck J. MrBayes 3.2: Efficient Bayesian Phylogenetic Inference and Model Choice Across a Large Model Space. Submitted to Systematic Biology.
2011
- Monier A, Welsh RM, Gentemann C, Weinstock G, Sodergren E, Armbrust EV, Eisen JA, Worden AZ. 2011. Phosphate transporters in marine phytoplankton and their viruses: cross-domain commonalities in viral-host gene exchanges. Environ Microbiol. doi: 10.1111/j.1462-2920.2011.02576.x.
- Eisen JA. 2011. Environmental shotgun sequencing: Its potential and challenges for studying the hidden world of microbes, Frans J. de Bruijn, (ed), Handbook of Molecular Microbial Ecology I: Metagenomics and Complementary Approaches, Vol. 1.. Wiley Press.
- Wu M and 'Eisen JA. 2011. Towards Automated Phylogenomic Inference, Frans J. de Bruijn, (ed), Handbook of Molecular Microbial Ecology I: Metagenomics and Complementary Approaches, Wiley Press.
- Chandler JA, Morgan Lang J, Bhatnagar S, Eisen JA, Kopp A (2011) Bacterial Communities of Diverse Drosophila Species: Ecological Context of a Host–Microbe Model System. PLoS Genet 7(9): e1002272. doi:10.1371/journal.pgen.1002272. Open access.
- Kembel SW, Eisen JA, Pollard KS, Green JL. 2011. The Phylogenetic diversity of metagenomes. PLoS ONE 6(8): e23214. Open access.
- Mindell DP, Fisher BL, Roopnarine P, Eisen JA, Mace GM, Page RDM, Pyle RL. 2011. Aggregating, tagging and integrating biodiversity research. PLoS One 6(8): e19491. Open access.
- Sharpton TJ, Riesenfeld S, Kembel SW, Ladau J, O’Dwyer J, Green JL, Eisen JA, Pollard KS. 2011. PhylOTU: A High-throughput procedure quantifies microbial community diversity and resolves novel taxa from metagenomic data. PLoS Computational Biology, 7(1): e1001061PMCID 3024254. Open access.
- Martiny JB, Eisen JA, Penn K, Allison SD, Horner-Devine MC. 2011. Drivers of bacterial beta-diversity depend on spatial scale. Proc Natl Acad Sci U S A. 2011 May 10;108(19):7850-4. Epub 2011 Apr 25. PMCID 3093525. Open access.
- Wu D, Wu M, Halpern A, Rusch DB, Yooseph S, Frazier M, Venter JC, Eisen JA. 2011. Stalking the fourth domain in metagenomic data: searching for, discovering, and interpreting novel, deep branches in marker gene phylogenetic trees. PLoS One. 2011 Mar 18;6(3):e18011. PMCID 3060911. Open access.
- Hubisz MJ, Pollard KS, Siepel A (2011). PHAST and RPHAST: phylogenetic analysis with space/time models. Briefings in Bioinformatics, (12)1: 41-51PMCID 3030812. Open Access.
2010
- Gilbert JA, Meyer F, Antonopoulos D, Balaji P, Brown CT, Brown CT, Desai N, Eisen JA, Evers D, Field D, Feng W, Huson D, Jansson J, Knight R, Knight J, Kolker E, Konstantindis K, Kostka J, Kyrpides N, Mackelprang R, McHardy A, Quince C, Raes J, Sczyrba A, Shade A, Stevens R. 2011. Meeting Report. Meeting report: the terabase metagenomics workshop and the vision of an Earth microbiome project. Standards in Genomic Sciences, 3(3): 243-248. PMCID 3035311. Open access.
- Langille MGI, Eisen JA. 2010. BioTorrents: A File Sharing Service for Scientific Data. PLoS One. 2010 Apr 14;5(4):e10071. PMCID 2854681. Open access.
- Hartman AL, Riddle S, McPhillips T, Ludäscher B, Eisen JA. 2010. Introducing W.A.T.E.R.S.: a workflow for the alignment, taxonomy, and ecology of ribosomal sequences. BMC Bioinformatics. 2010 Jun 12;11:317. PMCID 2898799. Open access.
- O'Dwyer JP, Green JL (2010). Field theory for biogeography: a spatially explicit model for predicting patterns of biodiversity. Ecol Lett 13:87 - 95. Open Access.
- Kembel SW, Cowan PD, Helmus MR, Cornwell WK, Morlon H, Ackerly DD, Blomberg SP, Webb CO. 2010. Picante: R tools for integrating phylogenies and ecology. Bioinformatics 26:1463-1464. Open access.
2009
- Wu D, Hugenholtz P, Mavromatis K, Pukall R, Dalin E, Ivanova NN, Kunin V, Goodwin L, Wu M, Tindall BJ, Hooper SD, Pati A, Lykidis A, Spring S, Anderson IJ, D'haeseleer P, Zemla A, Singer M, Lapidus A, Nolan M, Copeland A, Han C, Chen F, Cheng JF, Lucas S, Kerfeld C, Lang E, Gronow S, Chain P, Bruce D, Rubin EM, Kyrpides NC, Klenk HP, Eisen JA. 2009. A phylogeny-driven genomic encyclopaedia of Bacteria and Archaea. Nature. 2009 Dec 24;462(7276):1056-60. PMCID 3073058
- O'Dwyer JP, Lake JK, Ostling A, Savage VM, Green JL. 2009. An integrative framework for stochastic, size-structured community assembly. Proc Natl Acad Sci 106:6170-6175. Open Access.
- Pollard KS, Hubisz MJ, Rosenbloom KR, Siepel A. 2009. Detection of non-neutral substitution rates on mammalian phylogenies. Genome Research, 20: 110-121 PMCID 2798823. Open Access.
2008
- Wu M, Eisen JA. 2008. A simple, fast, and accurate method of phylogenomic inference. Genome Biol. 2008 Oct 13;9(10):R151.. PMCID 2760878. Open access.
- Wu D, Hartman A, Ward N, Eisen JA. 2008. An automated phylogenetic tree-based small subunit rRNA taxonomy and alignment pipeline (STAP). PLoS One. 2008 Jul 2;3(7):e2566. . PMCID 2432038. Open access.
- Green J, Bohannan B, Whitaker R. 2008. Microbial biogeography: from taxonomy to traits. Science. 2008 320:1039-43.
Blog Posts
Jonathan Eisen wrote many blog posts at his widely read and cited "Tree of Life" blog about work on this project. These include:
- http://phylogenomics.blogspot.com/2011/09/i-think-that-i-shall-never-see.html
- http://phylogenomics.blogspot.com/2011/03/story-behind-story-of-my-new-plosone.html
- http://phylogenomics.blogspot.com/2011/01/phylogeny-rules.html
- http://phylogenomics.blogspot.com/2010/07/summary-of-ievobio-day-1-evolution.html
- http://phylogenomics.blogspot.com/2009/12/more-coverage-of-geba-phylogeny-driven.html
- http://phylogenomics.blogspot.com/2009/12/story-behind-nature-paper-on-phylogeny.html
- http://phylogenomics.blogspot.com/2008/01/calling-all-phylogeneticists-we-need.html
- http://phylogenomics.blogspot.com/2008/04/trip-to-calit2-camera.html
Patents
None.
3. Presentations
Template for list
Please list all presentation titles and abstract citations or titles (if appropriate) that have resulted from this project since your last grant report, including those presented at scientific conferences, university seminars, etc. You may send complete abstracts as a separate file (optional). Please bold or highlight terms in the right-hand column to indicate an affirmative.
List
Green Lab
- Field Theory and Community Assembly. James O'Dwyer. Talk presented at the Moorcroft lab meeting, Department of Organismal and Evolutionary Biology, University of Harvard, September 2009. Also presented at the Plotkin lab meeting, University of Pennsylvania, September 2009.
- Community Assembly across the Tree of Life. James O'Dwyer. Talk presented at the Population Biology Seminar, University of Leeds, UK, January 2010.
- From Field Theory to Ecology. James O'Dwyer. Talk presented at the Biomathematics Seminar, University of Durham, UK, January 2010.
- Ecology without species: phylogenetic perspectives on microbial diversity. S.W. Kembel and J.L. Green. Invited speaker, Organized Oral Session on ‘Species interactions and community ecology in the context of relatedness’, Ecological Society of America Annual Meeting, August 2009.
- Measuring phylogenetic diversity. S.W. Kembel. Invited workshop instructor, ‘Ecological approaches to analyzing complex community datasets’ workshop for Fungal Environmental Sampling and Informatics Network members, Botanical/Mycological Societies of America Annual Meeting, July 2009.
- Ecology without species: phylogenetic perspectives on microbial diversity. S.W. Kembel. Invited speaker, Early Career Scientists Symposium on ‘Phylogenies and Ecology’, University of Michigan, March 2009.
- Metagenomics approaches to biodiversity and biogeography. Jessica Green. Talk presented at the University of Ioannina, March 2009.
- Theory and Metagenomics-based Biogeography. Jessica Green. ASM 109th General Meeting in Philadelphia, at the special interest session Genomics Enabled Biogeography of Planet Earth organized by Tiedje and Klugman. May 2009.
- The Rainforest Within: Biodiversity of the Human Body and its Relationship to Health and Disease. Jessica Green was the moderator of this symposium at the Ecological Society of America, which was covered by Nature News (http://www.nature.com/news/2009/090807/full/news.2009.808.html) and their blog (http://blogs.nature.com/news/blog/2009/08/esa_2009_sweater_by_grandma_gu.html).
- Exploring the Invisible. Jessica Green. TED2010, Long Beach, February 2010.
- Biodiversity Theory and Metagenomics-based Biogeography. Jessica Green. Stanford Symposium on Evolution and Genomics, April 2010.
- Ecology without species: phylogenetic diversity and microbial ecology. Steven Kembel. Invited speaker, 'Frontiers in Biodiversity: a phylogenetic perspective' Symposium, Barcelona, Spain, October 2010.
- Phylogenetic ecology and metagenomics. Steven Kembel. Invited speaker, 18th Annual Meeting on Microbial Genomics, Lake Arrowhead, September 2010.
- Field Theory, Biogeography and Metagenomics. James O'Dwyer. Talk presented at Arizona State University Biophysics colloquium, November 2010.
- Field Theory, Biogeography and Metagenomics. James O'Dwyer. Talk presented at Los Alamos Center for Nonlinear Sciences weekly seminar, May 2011.
Pollard Lab
- The iSEEM Project: Phylogenetic approaches to microbial metagenomics. Thomas J. Sharpton, Samantha J. Riesenfeld, Joshua Ladau, Steven W. Kembel, Jessica L. Green, Jonathan A. Eisen, Katherine S. Pollard. Talk presented by Katie Pollard at Cold Spring Harbor Biology of Genomes Meeting, May 2010.
- Building phylogenies with metagenomic sequence reads. Samantha J. Riesenfeld, Thomas J. Sharpton, Steven W. Kembel, Jessica L. Green, Katherine S. Pollard. Talk presented by Samantha Riesenfeld at Cold Spring Harbor Biology of Genomes Meeting, May 2010.
- PhylOTU: A high-throughput procedure that identifies Operational Taxonomic Units from metagenomic data. Thomas J. Sharpton, Samantha J. Riesenfeld, Steven W. Kembel, Joshua Ladau, James O'Dwyer, Jessica L. Green, Jonathan A. Eisen, Katherine S. Pollard. Talk presented by Thomas Sharpton at the International Society for Microbial Ecology Annual Meeting, August 2010.
- Resolving the Hidden Biosphere: A high-throughput procedure that identifies OTUs from metagenomic data. Thomas J. Sharpton, Samantha J. Riesenfeld, Steven W. Kembel, Joshua Ladau, James O'Dwyer, Jessica L. Green, Jonathan A. Eisen, Katherine S. Pollard. Talk presented by Thomas Sharpton at Evolution, June 2010.
- PhylOTU: Quantifying Microbial Diversity and Identifying Novel Taxa from Metagenomic Data. Thomas J. Sharpton, Rebecca Lamb, Samantha J. Riesenfeld, Joshua Ladau, Steven W. Kembel, James O'Dwyer, Jessica L. Green, Jonathan A. Eisen, Katherine S. Pollard. Talk presented by Thomas Sharpton at the International Human Microbiome Conference, March 2011.
- Inferring the shapes of species ranges from distance-decay relationships. Joshua Ladau, Jessica L. Green, Katherine S. Pollard. Talk presented by Joshua Ladau at the Ecological Society of America Meeting, August 2010.
- Inferring the shapes of species ranges from distance-decay relationships. Joshua Ladau, Jessica L. Green, Katherine S. Pollard. Talk presented by Joshua Ladau at the International Society for Microbial Ecology Annual Meeting, August 2010.
- Linking beta diversity to the shapes of species ranges and niches using geometric probability. Joshua Ladau, Jessica L. Green, Katherine S. Pollard. Talk presented by Joshua Ladau as an invited seminar at Stony Brook University, October 2010.
- Phylogenetic Approaches to Microbial Metagenomics - Who is out there and what are they doing? Seminar presented by Katherine S. Pollard at New York University, Department of Biology, November, 2010.
Eisen Lab
Jonathan Eisen - sampling of talks with links to slides and in some cases audio
- July 2011 Talk for MBL Microbial Diversity Course
- June 2011 Talk for Indoor Air Meeting
- March 2011 Talk on "phylogenetic analysis of metagenomic data" for Keystone Symposium on Microbial Communities
- March 2011 Talk on 'Microbial Phylogenomics' for Bodega Bay Workshop on Applied Phylogenetics
- February 2011 Talk at UCSF
- February 2011 Talk at UC Berkeley
- January 2011 Talk on "Microbial phylogenomics" at JCVI
- September 2010 Talk at Lake Arrowhead Microbial Genomes Meeting
- April 2010 Talk about GEBA at ASBMB
- April 2009 Video of talk at DOE JGI User meeting
Aaron Darling
- March 2011. Next generation metagenomics, for Environmental Microbiology sound bites series at UNSW, Sydney, Australia
- January 2011. Comparative genomics and recombination in archaeal populations, for Sydney Joint Academic Microbiology Seminars, Sydney, Australia
- October 2010. de novo Metagenome phylogeny and linkage estimation, JGI, Walnut Creek, CA
- Poster presentation: September 2010. Estimating linkage among short metagenomic read fragments with Bayesian phylogenetics. Lake Arrowhead microbial genomes meeting.
Morgan Langille
- Langille MGI and Eisen JA (2010) “Characterizing Protein Families of Unknown Function” 18th Annual International Meeting on Microbial Genomics, September 12-16, 2010, Lake Arrowhead, California, USA.
- Langille MGI and Eisen JA (2010) “BioTorrents: a file sharing service for scientific data” 110th General Meeting of the American Society for Microbiology, May 23-27, 2010, San Diego, California, USA.
- Langille MGI and Eisen JA (2009) “BioTorrents: a file sharing service for scientific data” Biology and Mathematics in the Bay Area Meeting, Nov. 14, 2009, Santa Cruz, California, USA.
- July 2010. BioTorrents: A File Sharing Service For Scientific Data. Open Science Summit, Berkeley, California, USA.
- June 2010. BioTorrents: A File Sharing Service For Scientific Data” Bioinformatics Technology Forum, UC Davis, Davis, California, USA.
Guillaume Jospin
- Poster presentation: March 2011. Building de novo protein families. Keystone Symposia Microbial Communities Meeting in Breckenridge, CO
Dongying Wu
- Poster presentation: March 2011. Identify gene markers for different taxonomic groups in Archaea and Bacteria Genomes. Keystone Symposia Microbial Communities Meeting in Breckenridge, CO
- Poster presentation: March 2010. Identify Novel Phylogenetic Markers for the Archaea and Bacteria Genomes. Genomics of Energy & Environment 5th Annual DOE Joint Genome Institute User Meeting, Walnut Creek, California
- Poster presentation: March 2009. Phylogenetic Diversity Contributions of the Genomic Encyclopedia for Bacteria and Archaea Pilot Project. Genomics of Energy & Environment 4th Annual DOE Joint Genome Institute User Meeting, Walnut Creek, California
- Poster presentation: March 2007. Tree-Based Small Subunit rRNA Taxonomy Assigning Pipeline (STAP). Genomics of Energy & Environment 2th Annual DOE Joint Genome Institute User Meeting, Walnut Creek, California
Wu Lab
Martin Wu
- September 2010, Microbial Genomics and Evolution. Department of Biochemistry and Molecular Genetics, School of Medicine, UVA.
4. Additional information
A. Exclusive of grant requirements, please describe new research collaborations with other MMI-funded grant project leads, with other marine microbiologists, and with researchers from outside the fields of marine microbiology and marine microbial ecology that resulted from this research (if applicable).
- Katherine Pollard and Jonathan Eisen have been invited to submit a proposal entitled “The Environmental Niche Atlas: Global Mapping of Microbial Functions” to MMI.
- The Pollard lab was invited to join the Human Microbiome Project analysis team.
B. Exclusive of grant requirements, have you provided data, samples, cultures, MMI-funded facilities, equipment use, etc. to other researchers? If so, please describe.
All our software has been developed openly and has been released usually even prior to publication of papers describing the programs. We have helped dozens of researchers install and run the software as well as make use of associated data sets.
C. Did you build or improve any instruments, devices, robots or software? Have the codes, designs or specs (etc.) been made available to the public? How many times has this information been downloaded or shared (if known)? Please include information here even if you listed above the related publication that describes this activity.
Software Developed
- Picante. Publicly available.
- AMPHORA. Software for automated phylogenomic analysis for whole genome phylogenies and metagenomic analyses. Publicly available.
- Amphora-2. Publicly available.
- STAP. Software for automated alignment, phylogenetic analysis, and phylogeny based classification of rRNA sequences. Publicly available.
- WATERs. Scientific workflow software for analysis of rRNA sequences. Publicly available.
- PhylOTU. Software for phylogeny-based classification of rRNA OTUs. Publicly available.
- MetaPASSAGE. Publicly available.
- PhyloP. Publicly available.
- Biotorrents. Website and software for bittorrent based sharing of biology related files. Publicly available.
- Zorro. Software for automated masking of sequence alignments. Publicly available.
D. Please describe any challenges that have been overcome and those that persist in preventing you from achieving any Grant Activities, Outputs and Outcomes.
The biggest challenge in this project was trying to integrate our tools into the CAMERA database. For reasons unclear to us, working with teams from CAMERA was very awkward and difficult and despite multiple efforts, nothing much ever came of our interactions with CAMERA.
E. Any lessons learned regarding your research and/or collaborations from a scientific, management, or other perspective?
F. Please describe any special recognition this project has received, and include information about the ways in which your scientific efforts have made a contribution beyond the field of marine microbial ecology (if any).
- Recognition
- Jonathan Eisen was the recipient of the 2011 Benjamin Franklin Award for contributions to Open Science
- Jessica Green was selected as a Senior TED2011 Fellow.
- Jessica Green was selected as a TED2010 Fellow.
- James O'Dwyer was selected as a Santa Fe Institute Omidyar fellow, beginning in August 2010.
- James O'Dwyer and Jessica Green received the Ecological Society of America Outstanding Theory paper award for 2010, for "Field theory for biogeography: A spatially explicit model for predicting patterns of biodiversity."
- In 2009, Katie Pollard was featured in the NOVA television special "What Darwin Never Knew", an Animal Planet show "Mutants", an evolution episode of the German science TV program "Quarks and Co", and a SETI radio program. She also wrote the Scientific American May 2009 cover story about comparative genomics.
- Impacts
- The tools we are developing will empower a wide range of metagenomic studies, from marine environments to the human microbiome. We have re-analyzed several published data sets and found exciting patterns (e.g. novel taxa, associations between phylogenetic diversity and environmental variables) missed in the initial studies.
- Press Coverage Examples
- Jessica Green was interviewed about biodiversity theory by the New York Times in 2011, and about microbial biodiversity by TED.com in 2011 and The Science Network in 2010.
- http://www.nytimes.com/2011/08/30/science/30species.html?_r=1
- http://www.microbemagazine.org/index.php/06-2011-current-topics/3451-metagenomic-analysis-suggests-fourth-domain-within-tree-of-life
- http://phylogenomics.blogspot.com/2011/03/story-behind-story-of-my-new-plosone.html
- http://www.economist.com/node/18437900?story_id=18437900&fsrc=rss
- http://www.genomeweb.com/informatics/morgan-langille-lured-biology-captured-challenge
- http://arstechnica.com/science/news/2010/04/bittorrenting-biology-getting-the-big-picture-in-search.ars?utm_source=microblogging&utm_medium=arstch&utm_term=Main%20Account&utm_campaign=microblogging
- http://www.nature.com/news/2010/100421/full/4641108a.html
- http://www.nature.com/news/2009/091117/full/news.2009.1093.html
G. Please list the web addresses for your lab and for any databases, resources, etc. related to this grant.
- Pollard lab: http://docpollard.org
- Pollard lab software: http://docpollard.org/software.html
- Niche mapping data and range maps: http://docpollard.org/marine_diversity/
- Green lab: http://biology.uoregon.edu/people/green/
H. Please provide a brief narrative description of expenditures to date and planned upcoming expenses, including an explanation of any budget variances and surpluses.
Budgets are all spent out.
I. Please use this space to respond to any additional questions MMI posed when sending you this form (See below). You may also use this space to provide any additional feedback to MMI. How can the Initiative do its job better?
5. Nucleic acid sequencing
No sequencing is supported by this grant.
6. Personnel
Personnel Table
Eisen Lab
- Senior Personnel: Dongying Wu, Martin Wu, and Aaron Darling
- Post docs: Morgan Langille
- PhD students: Amber Hartman
- Other staff: Srijak Bhatnagar & Guillaume Jospin
Pollard Lab
- Senior Personnel: Katherine S. Pollard
- Post docs: Joshua Ladau, Samantha Riesenfeld, Thomas J. Sharpton
- PhD students: none
- Other staff: none
Green Lab
- Senior Personnel: Jessica L. Green
- Post docs: James O'Dwyer & Steve Kembel
- Undergraduate students: Jesse Zaneveld
Wu Lab
- Senior Personnel: Martin Wu
- Post docs: Alex Koeppel
7. Additional information
Meetings
Bi-weekly PI-only conference calls
The three PIs meet bi-weekly by phone or skype conference call. These calls focused mainly on logistical issues (e.g. hiring, website, computing, reports) and strategic planning (e.g. collaborations between labs, shared resources). Notes from all calls are available on our wiki at http://iseem.private.openwetware.org/wiki/ISEEM-Conference-calls.
Bi-weekly full group conference calls
The full group (PIs, postdocs, collaborating lab members) meet bi-weekly by skype conference call. These calls focus mainly on scientific issues, in particular projects that are collaborative or impact the aims of different labs (e.g. alignments and trees for marker genes, simulation methods, diversity measures). Notes from all calls are available on our wiki at http://iseem.private.openwetware.org/wiki/ISEEM-Conference-calls.
In person meetings
Annual Meetings
- December 2008 at UC Davis: http://iseem.private.openwetware.org/wiki/Dec08_Quarterly
- December 2009 at Gladstone Institutes, UCSF: http://iseem.private.openwetware.org/wiki/Dec09_Annual
- June 2010 at University of Oregon: http://iseem.private.openwetware.org/wiki/Jun10_Annual
Ad Hoc Meetings
- The personnel of the Eisen and Pollard labs made regular visits to encourage an informal exchange of scientific questions, ideas, and techniques related to the iSEEM project.
- Joshua Ladau visited the Green lab several times per year throughout the project to discuss joint work.
- James O'Dwyer visited the Green lab in June 2011 to discuss joint work.
- James O'Dwyer visited the Levin lab, Princeton University in June 2011 to discuss work related to the iSEEM project.
CAMERA Meetings
- We held a quarterly meeting April 18, 2008 at CalIT2 on the UCSD campus as part of a meeting with the CAMERA team. The meeting consisted of a discussion of the goals CAMERA as well as the goals of the iSEEM project as well as the CAMERA subcontract to the Eisen lab. We then discussed how the iSEEM team could work with CAMERA both to get the science done that is part of the iSEEM project as well as to implement in CAMERA any tools the iSEEM project develops. Follow up discussions were held with people from CAMERA including Paul Gilna and Mark Ellisman.
- We had a joint group meeting with people from CAMERA, the Kepler Workflow Project, and our group in September 2008. The meeting was held at UC Davis in the Genome Center. In the meeting we discussed how to work with CAMERA to take methods developed from the iSEEM project and integrate them as Kepler Workflows (http://kepler-project.org/) within the CAMERA system. We came up with a plan for the next few months of work which involves first working on a rRNA analysis workflow and using it as a test to see how to take workflows from the iSEEM project and develop them into CAMERA tools. Once this is done we will move on to protein analysis workflows.