IGEM:IMPERIAL/2009/M1 VERSION
Module 1
Overview
This module is aimed at synthesising the protein of interest that will be subsequently encapsulated. The aim of our project is to create a generic platform for drug production.
Phenylalanine Hydroxylase (PAH)
Mutation in this enzyme has been associated with occurence of Phenylketonuria. Because PAH cannot break-down phenylalanine (an amino acid that cannot be synthesised by the body, it has to come from the diet) into tyrosine, phenylketonuriacs accumulate phenylalanine in the blood and in the brain which in term causes mental retardation if the diet is not tightly controlled. More information on PKU here, here and here
Cellulase
We propose to use a cellulase enzyme able to catalyse all three enzymatic reactions (crystal to cellulose to cellobiose/cellotetrose and to glucose) involved in the degradation of cellulose crystals into glucose. The particular enzyme we will be using is from Clostridium thermocellum Gene/Protein information. Another advantage of using Clostridium's cellulase is that it is protease resistant, therefore will be protease resistant in the target environment of the small intestine. More information can be found here.
Description of genetic circuit
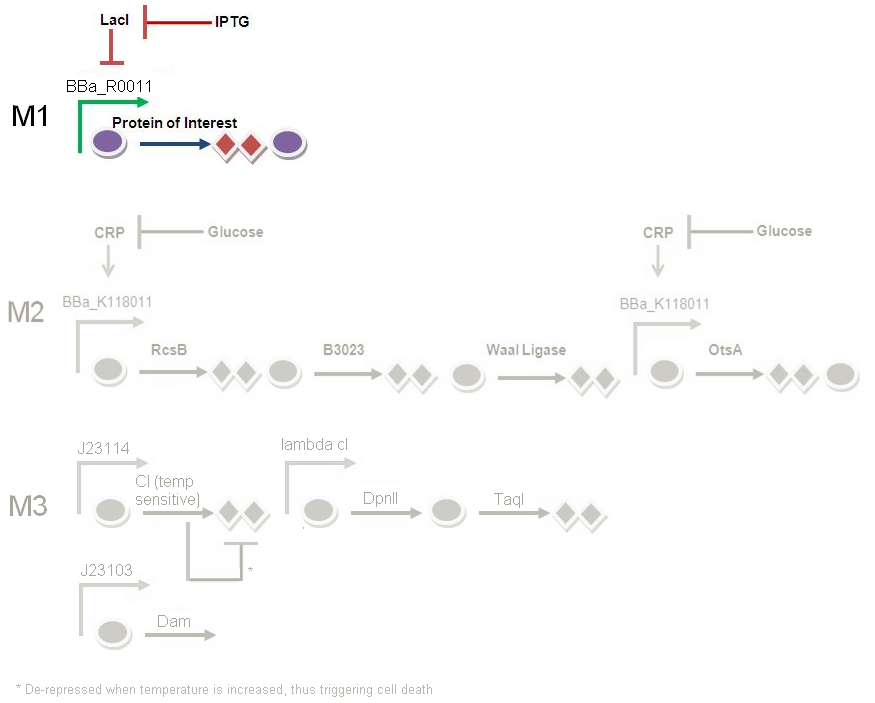
In this genetic circuit, LacI is being produced constitutively all the time. LacI represses the production of PAH / cellulase, and hence repressed M1. This is when there is no IPTG input.
When we want to kick start M1, we pipette in IPTG. An increase in IPTG levels in the media is sensed by e. coli. This will repress the LacI promoter, and LacI will not be produced. As a result, production of our enzyme of interest will not be repressed.
In conclusion, the addition of IPTG represents the beginning of protein production.