IGEM:IMPERIAL/2009/M1/Assays/Cellulase
Cellulase assay
Aims
- To monitor the activity of cellulase produced using a fluorescent substrate. The cellulase degrades the fluorophore-containing precursor into glucose and fluorescent molecule. The amount of fluorescence measurement over time is therefore directly proportional to the activity of the enzyme.
Description
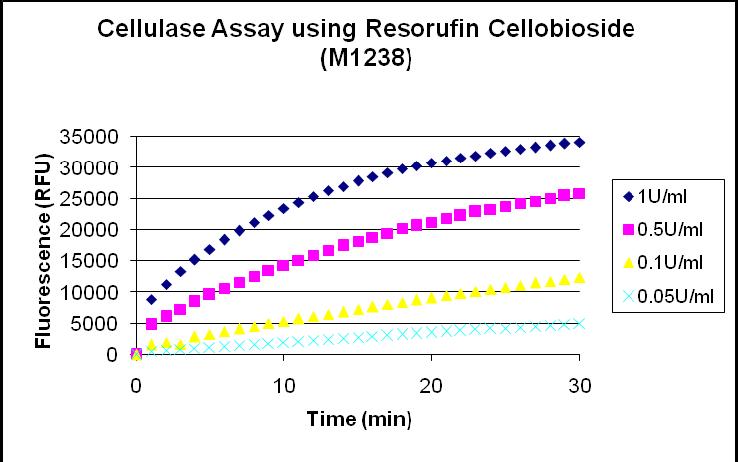
Activity of most cellulases is monitored by the fluorescent substrate, Resorufin Cellobioside, contained in the kit. Upon cleavage, the fluorescent compound, Resorufin, is released and fluorescent activity measurements can be taken. The Fluorescent Cellulase Assay Kit allows fast and easy detection of most cellulases in a microtiter plate based assay format.
2005 assay (Preferred one)
Description
The assay can be completed in 30 minutes or less. It is done by mixing the enzyme with the EnzChek® cellulase substrate working solution, incubate, and read the results.
This fluorescence substrate is highly sensitive, with a detection limit as low as 40 μU/mL cellulase. It is also possible to use this substrate in a colorimetric assay, albeit with reduced sensitivity.
Assay kit is from invitrogen.
Printable protocol is here
Calculations
Activity of the enzyme is measured in mU/mL and worked out from the assays' calibration curve:
2009 assay (2nd best)
COMMERCIALLY AVAILABLE ASSAY KITS
The kit works for 200 assays.
The MarkerGene™ Fluorescent Cellulase Assay Kit from Axxoraor from Marker Gene itself
MATERIALS
A.) Substrate Reagent: 5mM Resorufin Cellobioside in Dimethyl sulfoxide (DMSO). Dilute to 0.5mM for use in the assay protocol below.
B.) Reference Standard: 5mM Resorufin in Dimethyl sulfoxide (DMSO). Dilute to desired concentration using supplied DMSO and reaction buffer.
C.) Reaction Buffer: This buffer contains 100mM Sodium Acetate, pH 6.0.
D.) Stop Buffer: This buffer contains 165mM Tris Base, pH 10.0.
E.) Dimethyl Sulfoxide
Storage and Handling: The substrate reagent and reference standard included in this kit should be kept cold when not in use and stored at -20˚C. Protect solutions of the substrate reagent and reference standard from light.
ENZYME ASSAY
Date should be normalised by subtracting the enzyme reactions from a blank sample.
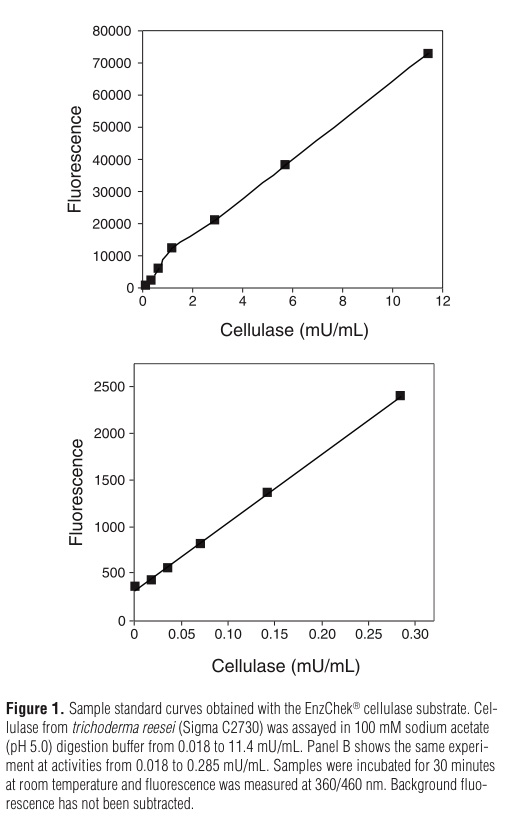
Enzyme can be extracted from solid material (such as plant tissue) by freezing in liquid nitrogen and grinding in a mortar, then suspending the resulting powder in reaction buffer (see Figure 1 below). Presence of solid material in sample may affect fluorescence readings. Solid materials should be removed by centrifugation prior to measurement if possible.
1.) It is recommended that samples to be assayed are diluted at least 1:1 in reaction buffer (Product M1245-001). Enzyme can be extracted from solid material (such as plant tissue) by freezing in liquid nitrogen and grinding in a mortar, then suspending the resulting powder in reaction buffer (see Figure 1 below). Presence of solid material in sample may affect fluorescence readings. Solid materials should be removed by centrifugation prior to measurement if possible. Keep samples in an ice bath until needed.
2.) Purified enzyme concentrations should be prepared fresh by diluting enzyme in reaction buffer (Product M1245-001). Keep samples in an ice bath until needed. *Nuri Purswani 11:15, 13 August 2009 (EDT): What volume of reaction buffer to dilute in?
3.) To a 96-well microtiter plate, add samples and purified enzyme solutions to wells in triplicate (50 μL/well). Also include in triplicate wells for blanks and reference standards (50 μL reaction buffer/well).*Nuri Purswani 11:15, 13 August 2009 (EDT): Ratio of sample to enzyme solution?
4.) Prepare reference standard solutions by diluting 5mM reference standard solution (Product No. M1245-003) to 10X desired concentration in DMSO (Product No. M1245-005). Dilute 10X solution 1:10 in reaction buffer (Product No. M1245-001) to reach final concentration. Mix solution and let stand for 5 minutes prior to use. To wells to be used as reference standards, add prepared reference standard solution. (100 μL) (see note (2) below). If a standard curve is desired to produce kinetic data, reference standard solutions should be prepared at several concentrations and added to wells. Concentration of standard may vary depending on enzyme concentration or sensitivity of reader.
5.) Prepare a 0.5mM substrate reagent solution by diluting 5mM substrate reagent (100 μL) (Product No. M1245-002) in reaction buffer (900 μL) (Product No. M1245-001). Prepare this solution just prior to performing the assay (see note (1) below). Add 0.5mM substrate reagent (50 μL/well) to all wells, except those containing reference standard.
6.) Read fluorescence (Ex/Em = 571/585 nm) in a microtiter plate reader, using appropriate filters. Use the wells containing reference standard to optimize reading conditions. If kinetic data is desired, readings may be taken beginning immediately after addition of the substrate reagent, and at several defined time points thereafter. If only an endpoint reading is needed, plate should be incubated at room temperature for the desired time, and fluorescence recorded after the addition of stop buffer (30μL/well) (Product M1245-004). *Nuri Purswani 11:15, 13 August 2009 (EDT): What filters?: Does the lab have this one?
7.) Average the readings of duplicate samples. Subtract fluorescence of blanks from that of each sample in order to normalize data. (See Note (1) below).
8.) Generate a calibration curve using purified enzyme samples by plotting normalized fluorescence vs. time (log-log).
9.) Using the calibration curve generated in step 8, determine the activity (concentration) of the enzyme in the sample, and extrapolate this data to determine the original enzyme concentration.
Note (1.) 0.5mM substrate reagent should be prepared just prior to performing the assay to prevent nonspecific hydrolysis of substrate. A small amount of nonspecific hydrolysis may occur over time if assays are performed over a long period (several hours). Potential hydrolysis may necessitate the need for blank wells as recommended in the assay protocol.
Note (2.) In addition to generating kinetic data, use of reference standard wells may be useful in optimizing reading conditions of the microtiter plate reader.
CALIBRATION CURVE The curve below tells us the enzymatic activity as a function of relative fluorescence units over time(RFU). By quantifying our RFU, we can directly relate our data to the standard curves below to obtain a measure of enzymatic activity.
1995 assay (we're not using this!!!)
PRINCIPLE
Cellulose + H2O ---Cellulase--> D-Glucose
D-Glucose + ATP ---Hexokinase--> D-Glucose 6-Phosphate + ADP
D-Glucose 6-Phosphate + ß-NAD ---G-6PDH--> 6-PG + ß-NADH
Abbreviations used:
ATP = Adenosine 5'-Triphosphate
ADP = Adenosine 5'-Diphosphate
ß-NAD = ß-Nicotinamide Adenine Dinucleotide, Oxidized Form
ß-NADH = ß-Nicotinamide Adenine Dinucleotide, Reduced Form
G-6PDH = Glucose 6-Phosphate Dehydrogenase
G-PG = 6-Phospho-D-Gluconate
CONDITIONS:
T = 37°C
pH = 5.0
A340nm
Light path = 1 cm
METHOD: Spectrophotometric Stop Rate Determination
REAGENTS:
A. 50 mM Sodium Acetate Buffer, pH 5.0 at 37°C (Prepare 200 ml in deionized water using Sodium Acetate, Trihydrate, Sigma Prod. No. S-8625. Adjust to pH 5.0 at 37°C with 1 M HCl.)
B. 5% (w/v) Sigmacell Solution (Sigmacell)(Prepare 100 ml in Reagent A using Cellulose (Sigmacell), Type 20, Sigma Prod. No. S-3504. Mix and heat gently to make a uniform suspension.)
C. Cellulase Enzyme Solution (Cellulase)(Immediately before use, prepare a solution containing 2 - 6 units/ml of Cellulase in cold deionized water.)
D. Glucose (HK) Determination Vial (16-10)(Use Glucose (HK) 10, Sigma Stock No. 16-10. Dissolve the contents in 10 ml of deionized water.)
PROCEDURE:
Step 1:
Pipette (in milliliters) the following reagents into suitable tubes:
| Reagent | Test | Control |
| Reagent B (Sigmacell) | 4.00 | 4.00 |
Equilibrate to 37°C. Then add:
| Reagent | Test | Control |
| Reagent C (Cellulase) | 1.00 | - |
| Deionised Water | - | 1.00 |
Immediately mix by swirling and incubate at 37°C for exactly 120 minutes with moderate shaking.
Pipette (in milliliters) the following reagents into suitable cuvettes:
| Reagent | Test | Control |
| Reagent D (16-10) | 3.00 | 3.00 |
Equilibrate to 25°C.
Monitor the A340nm until constant, using a suitably thermostatted spectrophotometer.
Record the initial A340nm for both the Test and Blank.
Then add:
| Reagent | Test | Control |
| Test Supernatant (Step 1) | 0.10 | - |
| Blank Supernatant (Step 1) | - | 0.10 |
Immediately mix by inversion and record the increase in A340nm until complete (for approximately 5 minutes).
Obtain the final A340nm for both the Test and Blank.
CALCULATIONS:
A340nm Test = A340nm Test Final - A340nm Test Initial
A340nm Blank = A340nm Blank Final - A340nm Blank Initial
Units/ml enzyme =
( (A340nm Test - A340nm Blank)(3.1)(5)(df) )/ ( (6.22)(2)(1)(0.1) )
3.1 = Final volume (in milliliters) of Step 2
5 = Total volume (in milliliters) of reaction mix (Step 1)
df = Dilution factor
6.22 = Millimolar extinction coefficient of ß-NADH at 340nm
2 = Conversion factor from 2 hours to 1 hour as per the Unit
Definition
1 = Volume (in milliliter) of cellulase used in Step 1
0.1 = Volume (in milliliter) from Step 1 used in Step 2
Units/mg solid = units/ml enzyme / mg solid/ml enzyme
UNIT DEFINITION:
One unit will liberate 1.0 μmole of glucose from cellulose in one hour at pH 5.0 at 37°C (2 hour incubation time).
FINAL ASSAY CONCENTRATION:
In a 5.00 ml reaction mix, the final concentrations are 40 mM sodium acetate, 4% (w/v) Sigmacell and 2 - 6 units of cellulase.
REFERENCE:
Worthington, C.E. (1988) Worthington Enzyme Manual, pp. 76-79, Worthington Biochemical Corporation, Freehold, NJ
NOTES:
1. This assay is based on the cited reference.
2. Where Sigma Product or Stock numbers are specified equivalent reagents may be substituted.
original page here
Relating activity to concentration
1. Purchase cellulase powder from sigma
2. Make cellulase solutions of concentrations (around the concentration of cellulase produced from M1).
3. Perform the same assay on all the concentrations, to obtain activity from each concentration.
4. Generate a calibration curve, matching concentration to activity. A calibration curve of cellulase concentration against activity is determined.
5. Match the activity of cellulase produced from M1, to its concentration, by read off this calibration curve. We now have the concentration of cellulase produced from M1.