IGEM:IMPERIAL/2009/M0/Modelling
M0
What we aim to do
Using characteristics of the CRP promoter and diauxic growth,
- Modelling how glucose consumption is linked to cell growth and using the drop in glucose concentration for indirect timing of encapsulation (via the CRP promoter)
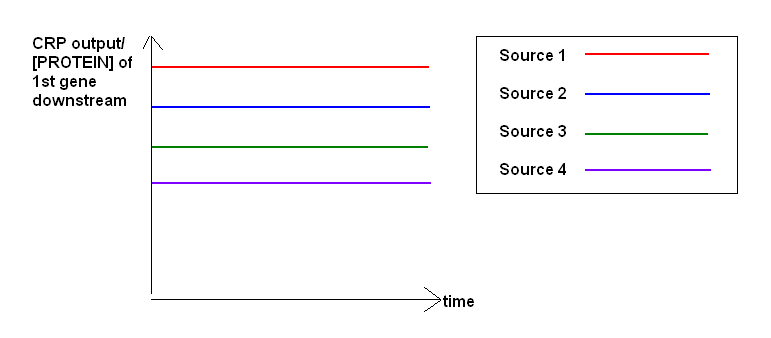
- Modelling how the CRP promoter activity varies with different secondary carbon sources
Literature background
A good overview of catabolite repression modelling(comprehensive citation), followed by comparison of 2 different models
Catabolite repression in Escherichia coli– a comparison of modelling approaches
CRP Promoter
Edinburgh 08 on CRP promoter
1) There is a dramatic drop in promoter activity between glucose concentrations of 0mM and 10mM (prior to overnight LB cultures)
2)PcstA is sensitive to glucose, arabinose and maltose. Arabinose and maltose actually repress PcstA action also.(ie. secondary carbon sources repress CRP as well)
Diauxic growth curve
What all biologists will generally know about diauxic growth
See Diauxic growth curves
Monod classified sugars into class "A" or "B". Only class B sugars (arabinose, maltose, rhamnose, lactose, xylose, galactose) give rise to diauxic growth when present with glucose
[http://mic.sgmjournals.org/cgi/reprint/141/1/71.pdf Is Escherichia coli growing in glucose-limited chemostat culture able to utilize other sugars without lag?] [http://www.pubmedcentral.nih.gov/picrender.fcgi?artid=167924&blobtype=pdf Kinetics of the Simultaneous Utilization of Sugar Mixtures by Escherichia coli in Continuous Culture]
Other Bacterial growth/consumption models
Possible models
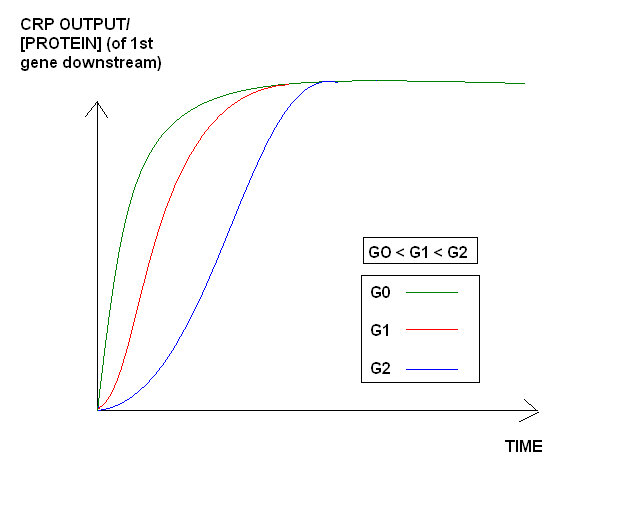
Varying glucose affect time delay and output yield
Expected outcome: Increases in CRP PoPs output will be related to low levels of [glucose]. Similarly, decreases in CRP output will be related to high levels of glucose in the medium. The plot below shows a rough idea of what we expect to "see" for different initial concentrations of glucose in the medium. Hopefully we would like a transfer function that relates PoPs = f([glucose])
- Goal: To quantify the relationship between glucose present in the medium and time to reach steady state GFP activity/CRP PoPs output.
- Input: Different initial concentrations of glucose
- Output: Fluorescence over time, relate to CRP expression. (Working at promoter "inducible phase")
Growth curve
- See IC igem 2008 dry labs: Growth Curve
Current models
Bacterial Growth-Consumption model: Glucose consumption
What are we doing?:
We are characterizing the glucose medium and growth conditions in the population. The intention of using this model is that we will link glucose consumption to CRP promoter characterization. In this phase we are characterizing the "primary source", in the pre-protein synthesis phase. Once we obtain optimal conditions for our medium, we will couple this to the secondary source and eventually, full characterization of the switching mechanism of the MO module.
What are the hypotheses and justifications?:
Bacterial growth is dependent on the amounts of glucose (carbon source) present in the medium. Once resources are used up, the population stops growing. Furthermore, glucose consumption not only depends on the amounts of glucose initially present in the medium. Amounts of glucose decrease faster if we have more bacteria present in our medium. This mutual dependence model shows that once glucose is used up to a certain amount, bacterial growth will saturate.
The model:
Table 1. Meaning of modelling terms
| Term | Variable/Parameter/function? | Meaning/description |
| [Glu] | Variable | Concentration of glucose in our medium (as a function of time). |
| [Popul] | Variable | Population of bacteria in our medium (as a function of time). |
| k1 | Parameter | Glucose consumption rate |
| k2 | Parameter | Bacterial growth rate |
| fh([Glu]) | Function | Activating hill function of glucose. The higher the amount of glucose, the faster the population grows. When glucose falls below the threshold of activation (Th) the population stops growing. |
| Th | Parameter | Activating threshold for glucose hill function. |
| n | Parameter | Hill exponent. |
Typical simulations:
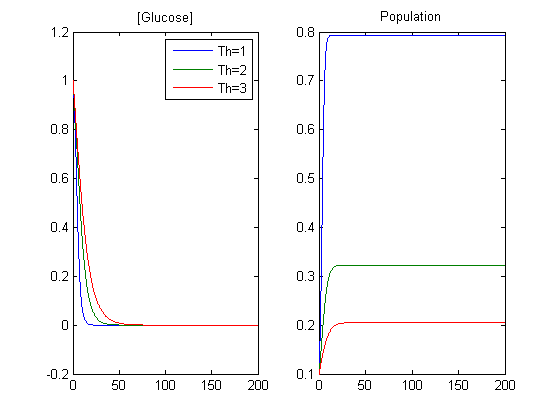
1) Variation of threshold (Th) in glucose hill function fh([Glu])
What do the simulations mean?
1) Variation of threshold (Th) in glucose hill function fh([Glu])
- The Th parameter sets the minimal amount of glucose needed for the population to grow. *Nuri Purswani 13:12, 18 August 2009 (EDT): Insert a sketch for illustration
- The higher the value of Th, the slower the consumption of glucose over time. (As glucose limits population growth).
- The higher the value of Th, the more glucose needed for the population to grow. Therefore, growth saturates earlier (i.e. our bacteria grow less)
Data fitting
How to test data against simulations?
How to test simulations against data?
Matlab code
- ODE file
- Call function file: Vary parameters here
Media:II09_call_glu_growth.ogg
To be tidied up
Overview
Test 1) Use the Jason Kelly technique[1] to characterize basal levels of expression of CRP using different secondary carbon sources.
Test 2) Quantify the time it takes to reach steady state expression of the CRP promoter using different initial amounts of glucose in our medium.
Aims
- How varying glucose input affects output yield:
The paper by Kelly provides a description of how to relate GFP output to PoPs in a constituitive promoter. We can also relate this to inducible promoters. [2] [1] Similar ideas can be used for the characterization of our CRP promoter.
- Relate GFP to POPs: Parts of pdf file from supplementary material in [2][1]. Here they are characterizing BBa_E0240, which we will be using as the testing construct for this section and also other bits of the project.
Modelling
Test 1)
- Goal: To characterize the secondary Carbon source that provides a suitable level of basal expression for the CRP promoter.
- Input: Different secondary carbon sources
- Output: Fluorescence over time, relate to CRP expression. (Working at promoter "constituitive phase")
References
- Canton B, Labno A, and Endy D. Refinement and standardization of synthetic biological parts and devices. Nat Biotechnol. 2008 Jul;26(7):787-93. DOI:10.1038/nbt1413 |
-
Measuring the activity of BioBrick promoters using an in vivo reference standard Jason R Kelly, Adam J Rubin, Joseph H Davis, Caroline M Ajo-Franklin, John Cumbers, Michael J Czar, Kim de Mora, Aaron L Glieberman, Dileep D Monie, and Drew Endy J Biol Eng. 2009; 3: 4. Published online 2009 March 20. doi: 10.1186/1754-1611-3-4