IGEM:IMPERIAL/2009/M0
Module 0 - Autoinduction medium
The autoinduction media will be used to direct the chassis through the various stages of pill manufacture.
PHASE 1: M1 = OFF, M2 = OFF
PHASE 2: M1 = ON, M2 = OFF
PHASE 3: M1 = ON, M2 = ON
PHASE 1: Innoculum Expansion (Kun) - Assay 0.1
Background:
The time at which we initiate protein production is dependent on culture OD.
An OD of 0.7 is generally considered to be the optimum time at which to induce protein production in a bacterial chassis. Above this value and the cells will be in the stationary phase and below this value there are too few cells producing protein.
The OD of the culture can be monitered by an automatic plate reader that informs when OD= 0.7.
There are two reasons why we wish to initiate protein production during the initial period of glucose utilisation.
1) Glucose is the more efficient energy source and cellular metabolism will be faster (therefore greater protein output per unit time)
2) During glucose utilisation, minimal energy will be diverted into colanic acid biosynthesis.
PHASE 2: IPTG addition and induction of protein production
Background:
When the OD reaches 0.7, IPTG is added to the culture, inducing protein production.
OD values are obtained to model bacterial growth rate when the cell is under protein production conditions (when IPTG is added)
The IPTG concentrations are varied to obtain maximal protein production while not being toxic to the cell.
PHASE 3: Induction of CRP promoter and time delay
Background:
The glucose concentration in the cell drops.
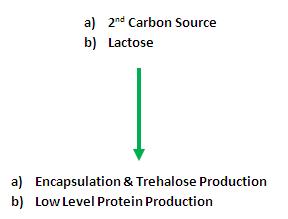
When the glucose concentration drops below a threshold concentration, the secondary carbon source will be utilised and the CRP promoter will be switched on (M2 activation)
The CRP promoter activity can be characterised by secondary carbon sources and initial glucose concentration
The time delay before glucose concentration drops below threshold is given by the following transfer function:
T = f ([glc]t=0, [IPTG])
Testing constructs
Cloning Strategy
Click on the link to see modelling directions
Shopping list
Extra details
PHASE 2: (Dineka) Characterisation of CRP Promoter's constitutive activity - on different carbon sources - Assay 0.2
Background
We aim to characterise the constitutive activity of the CRP promoter in the presence of secondary carbon sources (and in the absence of glucose). Constitutive activity will be tested using methods originally described by Jason Kelly.
By linking the promoter to the BBa_E0240 testing construct (RBS-GFP-TT), a quantitative measurement of the fluorescence over time can be directly obtained. This will then be converted to the relative transcriptional activity of the promoter (PoPS), for a given set of conditions.
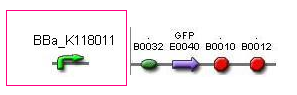
DNA construct
Promoter: BBa_K118011-BBa_E0240
Assay
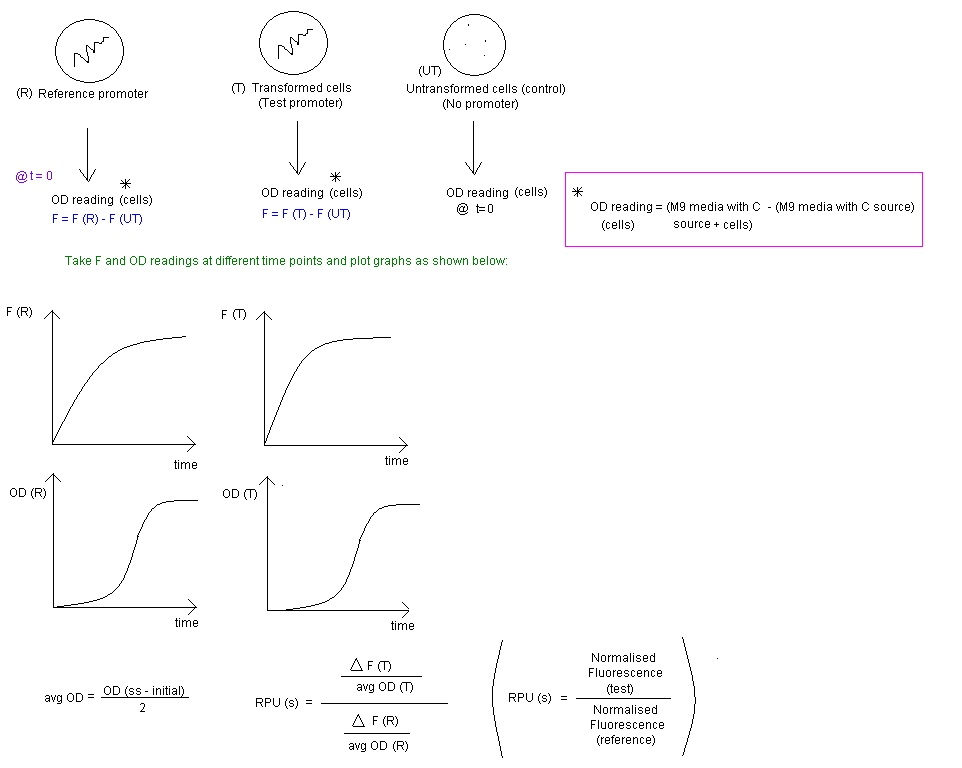
To characterise the promoters, a promoter measurement kit will be used.[1]
The contents of the M9 media will change according to the test carbon source required.
The Assembly Instructions describe a method to insert a test promoter into the promoter test construct by combining the test promoter with the GFP reporter device (BBa_E0240) and the backbone plasmid (pSB3K3).
The Measurement Instructions describe a method for measuring the activity of a test promoter relative to the reference standard promoter (BBa_J23101) so that a user of the measurement kit can report the activity of their test promoter in Standard Promoter Units (SPUs). The Registry of Standard Biological Parts includes the SPU calculator to convert the raw fluorescence and OD data into SPUs.
Data Output
1) Fluorescence output of CRP promoter in the "constituitive phase" (at steady state). (Intensity W/m2 or relative fluorescence units (RFU) depending on the type of kit we use).
2) OD measurement (CFU/mL)* Note, the conversion of OD to CFU/mL will require the prior production of a calibration curve.
- Vincent 05:48, 18 August 2009 (EDT): can you give a fictitious example of the data that would be produced by the experimental protocol ?
Note: This will be done for each carbon source. (See protocol).
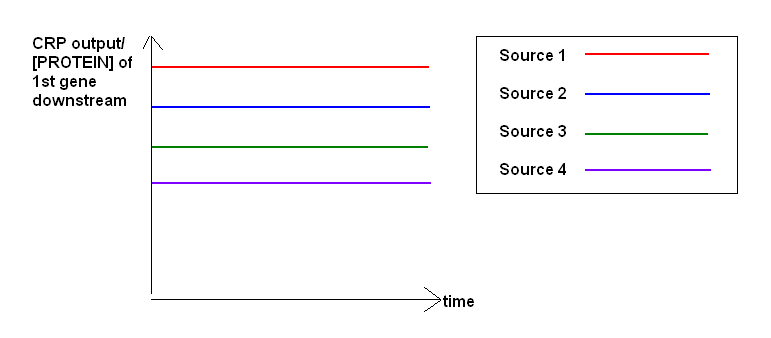
Note: For each carbon source, an RPU value will be obtained. These will be plotted on the same graph in order to find which carbon source produces the highest output.
Data Analysis & Modelling
- Goal: To characterize the secondary carbon source that provides a suitable level of basal expression for the CRP promoter.
- Input: Different secondary carbon sources.
- Output: Measure the fluorescence over time and relate this to CRP expression.
(Working at promoter "constituitive phase")
- Vincent 05:45, 18 August 2009 (EDT): if the output is GFP fluorescence, it is likely to increase over time with the growth of the population. You need to normalise fluorescence with the OD600 to have a chance to see a GFP steady state.
Protocol
http://openwetware.org/wiki/IGEM:IMPERIAL/2009/M0/Assays/1.1
PHASE 1-2 (Transition) Characterisation of glucose inhibition of the CRP Promoter (James) - Assay 0.3
Background
Relate the time it takes to reach steady state for a given input of glucose. Data generated from Assay 0.2 will indicate steady state for a given carbon source.
DNA construct
Promoter: BBa_K118011-BBa_E0240
Data Output
1) Fluorescence output from the CRP promoter (readings at 30 minute intervals). Here we are looking at its inducible action. (intensity W/m2 or relative fluorescence units (RFU) depending on what kit we use)
2) Optical density measurement (600nm)
Data Analysis & Modelling
- Goal: To quantify the relationship between glucose present in the medium and time to reach steady state GFP activity/CRP PoPs output.
- Input: Different initial concentrations of glucose
- Output: Fluorescence over time, relate to CRP expression.
- Vincent 05:42, 18 August 2009 (EDT): Where can I find the description of the model that produced these simulation outputs ? What are the assumptions that were used to build the model ?
Protocol
http://openwetware.org/wiki/IGEM:IMPERIAL/2009/M0/Assays/1.2
Editors:Nuri
The onset of encapsulation will be controlled by an autoinductive medium.
The mechanism of the autoinductive medium is the following:
In the bacterial culture, 2 carbon sources will be present -
1. Glucose: Primary carbon source (consumed first)
2. Secondary carbon source(To be decided):
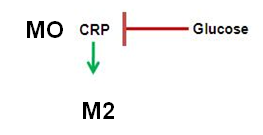
In order to induce encapsulation, the CRP promoter is inhibited by the presence of glucose. We would like to set off encapsulation once glucose levels have dropped.
Testing Rational
This part of the circuit is linked to our inducible medium idea MO, and we would like to characterize the effects of our culture medium on the output of the CRP promoter, which controls the genes that start both the encapsulation and the trehalose production. We have designed 2 tests to characterize the CRP promoter, which is the key element to kickstart our M2.
Test 1) Use the Jason Kelly technique[2] to characterize basal levels of expression of CRP using different secondary carbon sources.
Since the secondary carbon source is what bacteria will feed on once encapsulation is triggered, we would like to record their basal levels of expression of GFP, and use the source that will provide the higher state of GFP (see Modelling directions for illustration).
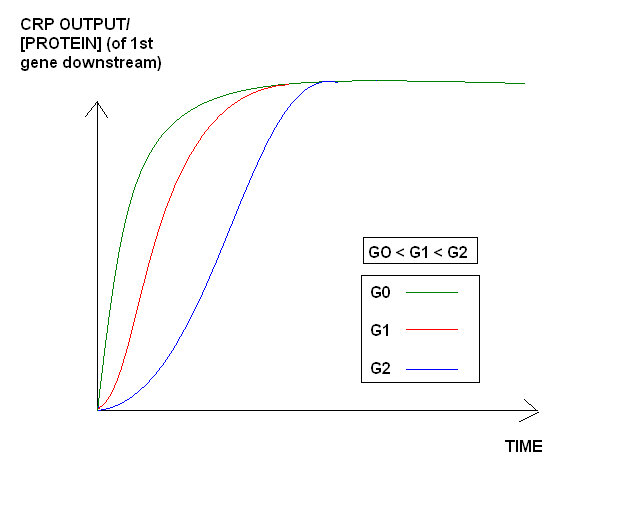
Test 2) Quantify the time it takes to reach steady state expression of the CRP promoter using different initial amounts of glucose in our medium.
In order to do this, we will have to relate initial amounts of glucose present in the culture medium to the time it takes to reach steady state with the CRP promoter. The CRP promoter is a "treshold switch"(inducible: See modelling directions for an illustration) such that when levels of glucose fall below a certain level, it switches on and triggers encapsulation. Effectively, different amounts of initial glucose in our medium make it act like a "timer", which will enable us beforehand to control the lag time before encapsulation takes place.
In order for both tests to work, we must have two sources of carbon (glucose and something else) present in the medium (diauxic growth curves) such that cells can obtain energy from an alternative carbon source once glucose has become depleted.
Cloning strategy (assembly)
No longer needed
PHASE 1: Growth of cells to correct OD
Growth Curve Modelling
To model how cell growth changes (OD) and concentration of glucose varies with time.
Glucose Sensing
This is to detect the concentration of glucose that is present.
Assays
Glucose Sensing
Glucose concentration in μmol/ml