Electrophoresis
Background
Electrophoresis is a biochemical technique that separates compounds by administering an electrical current. The current runs from a power source and travels from the anode end of the electrophoretic apparatus to the cathode end. The charge gradient within the system propagates movement according to the charge of the samples. For example, the sugar backbone of DNA possesses a negative charge, and so the macromolecule is repelled by the negative charge of the anode end, and attracted towards the positive charge of the cathode end. The velocity (and distance) that a compound travels down its medium is dependent upon the compound's electrophoretic mobility, as represented by:
[math]\displaystyle{ \mu = \frac{Q}{6 \pi r \eta} }[/math]
μ = electrophoretic mobility
Q = net charge
r = ionic radius of solute
η = medium viscosity
Motivations
Electrophoresis is a versatile technique that can separate different molecules based on selective characteristics, such as net charge, molecular weight, isoelectric point (i.e. the pH at which a molecule has a neutral charge), and shape. Various electrophoretic systems and techniques have been developed to specialize in the separation of compounds based on these varying characteristics. Multiple electrophoretic assays can also be used on a single sample (see Multidimensional Analysis), providing great specificity for sample analysis. Biomarking samples is often implemented to assist in visualizing different analytes. Electrophoresis is a valuable tool in microfluidics due to the diversity of separations and methods it can perform.
Challenges

Since electrophoresis typically deals with living matter, a prevalent limitation of the technique is contamination. Contamination of either the gel or chemicals (e.g. buffer) used in an electrophoretic experiment would result in contamination of an organic sample, and thus compromise results.[1] In order to overcome this limitation, it is imperative that electrophoresis be carried out in sterile laboratory settings to produce optimal results. Compared to other microfluidic separations techniques, electrophoresis is a more time-consuming separation process, and so the working sample can degrade after extensive periods of time.
Another challenge presented by electrophoresis is regulating heat generated by the electrical current. Excess heat has the potential to damage the medium (e.g. melting the gel) or cause unwanted denaturation of a sample.[2] Non-uniform distribution of heat can distort band shapes and cause curved, "smiling" bands (Figure 1).
Electrophoretic Techniques
The development of electrophoretic techniques has given rise to specializations of electrophoresis: gel electrophoresis, capillary electrophoresis (more on capillary gel electrophoresis), and free flow electrophoresis (FFE).
Gel Electrophoresis
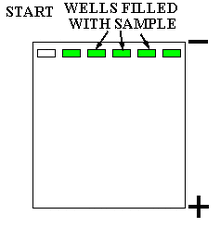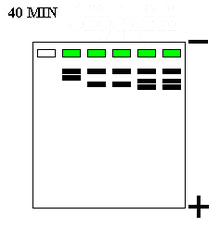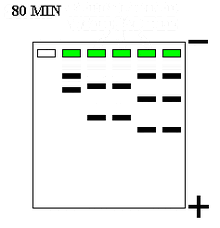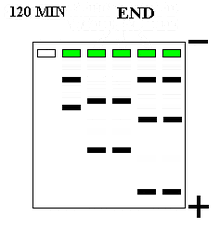
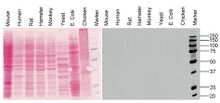
Compounds can be run through a gel medium (e.g. polyacrylamide, agarose) in electrophoresis for analysis. Polyacrylamide gel electrophoresis (PAGE) is a common separation technique used in analysis of nucleic acids and proteins within a gel. Polyacrylamide gels are used in this separation technique because it is easy to fabricate, porous, and unreactive with proteins (i.e. it will not react with samples). Gels can be run horizontally (agarose) or vertically (polyacrylamide).
Sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) is a method used to denature proteins and sort polypeptides according to molecular weight.[3] The loading buffer used in SDS-PAGE comprises of a reducing agent (e.g. β-mercaptoethanol), and a strong detergent (SDS itself). The reducing agent facilitates breaking of covalent bonds (e.g. disulfide bridges) and eliminates tertiary structure in proteins. As a detergent, SDS breaks noncovalent bonds and establishes a uniform net negative charge across the proteins, effectively eliminating discrepancies in charge between different polypeptides. Establishing a net negative charge sorts polypeptides through the gel solely based on their size (Figure 2). Smaller polypeptides will travel further down the gel because they experience less friction and are more easily able to navigate through the porous acrylamide gel, whereas larger polypeptides experience greater friction when travelling through the gel.[4] SDS-PAGE is especially useful in analysis of polypeptides and isolation of proteins based on molecular weight (Figure 3).
Native PAGE is another gel electrophoresis procedure that can sort samples through a gel. However, SDS is absent in native gels, meaning that proteins are not denatured, and instead analyzed in their original state, sorting proteins based on both net charge and molecular weight. Native gels are useful for analysis of amino acid composition in proteins and isolation of enzymes.
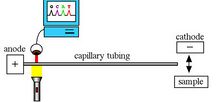
Capillary Electrophoresis
Capillary electrophoresis (CE) is a separation technique in which a sample is run through a thin capillary (Figure 4). Samples are separated on the basis of electrophoretic mobility. Heat transfer within capillaries promotes increased analysis speeds and produces more effective separations.[5] Unlike gel electrophoresis, capillaries are reusable for future analyses. CE is useful for analyzing DNA fragments and protein separations.
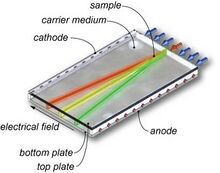
Free Flow Electrophoresis
In free flow electrophoresis (FFE), sample is inserted into a single well as a pressure-driven stream. An electrical current runs perpendicular to the stream, causing the formation of multiple streams separated by electrophoretic mobility (Figure 5). Miniaturization of FFE has improved efficiency by enabling analysis of samples in small volumes.[6]
Innovations
Miniaturization
Miniaturization of electrophoretic techniques has been developed for efficiency: maximizing productivity while minimizing area. These innovations enable experiments to run simultaneously, as well as reduce the time and cost required for running these experiments.[7] A feature of electrophoresis miniaturization is incorporation of separation channels, which work to increase channel efficiency.
Polymers, such as polydimethylsiloxane (PDMS) and poly(methyl methacrylate) (PMMA) are used in modern electrophoretic systems as they can be easily manufactured and are more robust than glass. PMMA notably is used in gel electrophoresis because it does not react with polyacrylamide gels. Different material considerations are used in electrophoresis when making the gel. For example, agarose gel is used for larger scale analysis of macromolecules, specifically larger DNA fragments.[8] Polyacrylamide has a higher resolution than agarose gels for analysis of smaller samples, such as single strands of DNA or protein analysis. The pores in PAGE are smaller and allow for greater discrimination and separation of samples, having versatility at the microscale.

Multidimensional Analysis
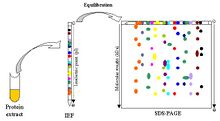
Multiple assays for a single sample can analyze between 2-4 different characteristics and improve test efficiency.[9] Multidimensional analysis makes use of analyzing different sample properties, such as combining isoelectric focusing and SDS-PAGE.[9],[10] Isoelectric focusing of the sample is run horizontally and separates proteins based on their isoelectric points (approximate pH where samples have a net neutral charge), establishing a ladder of samples based on a pH gradient. Afterwards, SDS-PAGE utilizes the same gel and runs it vertically, further separating and distinguishing proteins of similar isoelectric points by size. Multi-dimensional analysis exploits the different characteristics of samples (charge, size, etc.) order to comprehensively separate and distinguish samples (Figure 6).
Applications
Nucleic Acid and Protein Analysis
Electrophoresis sees common use in analyzing DNA, RNA, and proteins.[11] Electrophoresis is especially useful in genetic sequencing where different tissue samples from different organisms are compared to look for any corresponding protein or gene similarities. Sorting samples based on molecular weight assists in separation of nucleic acid fragments and protein polymers for research.
Medical Research
Electrophoresis is a critical tool in pharmacy; vaccine development and purification rely on electrophoresis for examination of a vaccine's contents.[12] Antibiotics are analyzed through electrophoresis to check for residue and impurity presence.[13],[14] Detection of analytes in drugs helps to maximize potency and individualize treatments for patients using drug therapy.
Environmental Monitoring
Soil chemistry in ecosystems reflects the overall health of an ecosystem. Performing electrophoresis on soil can detect harmful chemicals (e.g. allelochemicals) that can compromise the environment.[15] Electrophoresis of microbial population DNA also can reflect on conditions in the environment. Regularly monitoring environmental health discerns shifts and patterns in an ecosystem.
References
[1] C. C. Chéry; L. Moens, R. Cornelis; F. Vanhaecke. Capabilities and Limitations of Gel Electrophoresis for Elemental Speciation: A Laboratory's Experience. Pure Appl. Chem., 2006, 78, 91–103. DOI: http://dx.doi.org/10.1351/pac200678010091
[2] R. Westermeier. Gel Electrophoresis. A Guide to Theory and Practice, 1993, pp 210-212. DOI: http://dx.doi.org/10.1038/npg.els.0005335
[3] K. Weber; M. Osborn. Reliability of Molecular Weight Determination of Proteins by Polyacrylamide Gradient Gel Electrophoresis in the Presence of Sodium Dodecyl Sulfate. J. Biol. Chem., 1969, 244, 4406-4412. DOI: http://dx.doi.org/10.1016/0003-2697(78)90281-6
[4] J. I. Won; R. J. Meagher; A. E. Barron. Protein Polymer Drag-tags for DNA Separations by End-labeled Free-solution Electrophoresis. Electrophoresis, 2005, 26, 2138–2148. DOI: http://dx.doi.org/10.1002/elps.200410042
[5] J. Jorgenson; K. Lukacs. Capillary Zone Electrophoresis. Science, 1983, 222, 266–272. DOI: http://dx.doi.org/10.1126/science.6623076
[6] R. T. Turgeon; M. T. Bowser. Micro free-flow Electrophoresis: Theory and Applications. Anal. Bioanal. Chem., 2009, 394, 187–198. DOI: http://dx.doi.org/10.1007/s00216-009-2656-5
[7] A. J. Pfeiffer; T. Mukherjee; S. Hauan. Design and Optimization of Compact Microscale Electrophoretic Separation Systems. Ind. Eng. Chem. Res., 2004, 43, 3539–3553. DOI: http://dx.doi.org/10.1021/ie034071t
[8] I. Day; S. Humphries. Electrophoresis for Genotyping: Microtiter Array Diagonal Gel Electrophoresis on Horizontal Polyacrylamide Gels, Hydrolink, or Agarose. Anal. Biochem., 1994, 222, 389–395. DOI: http://dx.doi.org/10.1006/abio.1994.1507
[9] S. Tia; A. E. Herr. On-chip Technologies for Multidimensional Separations. Lab. Chip, 2009, 9, 2524–2536. DOI: http://dx.doi.org/10.1039/b900683b
[10] Y. Li; J. S. Buch; F. Rosenberger; D. L. DeVoe; C. S. Lee. Integration of Isoelectric Focusing with Parallel Sodium Dodecyl Sulfate Gel Electrophoresis for Multidimensional Protein Separations in a Plastic Microfludic Network. Anal. Chem., 2004, 76, 742–748. DOI: http://dx.doi.org/10.1021/ac034765b
[11] E. Southern. Detection of Specific Sequences Among DNA Fragments Separated by Gel Electrophoresis. J. Mol. Biol., 1975, 98, 503-517. DOI: http://dx.doi.org/10.1016/S0022-2836(75)80083-0
[12] W. Hurni; W. Miller. Electrophoresis for Genotyping: Microtiter Array Diagonal Gel Electrophoresis on Horizontal Polyacrylamide Gels, Hydrolink, or Agarose. J. Chromatogr., 1991, 559, 337–343. DOI: http://dx.doi.org/10.1006/abio.1994.1507
[13] C. L. Flurer. Analysis of Antibiotics by Capillary Electrophoresis. Electrophoresis, 1997, 18, 2427–2437. DOI: http://dx.doi.org/10.1002/elps.1150181233
[14] M. Hernandez; F. Borrull; M. Calull. Analysis of Antibiotics in Biological Samples by Capillary Electrophoresis. Trac-Trends Anal. Chem., 2003, 22, 416–427. DOI: http://dx.doi.org/10.1016/S0165-9936(03)00702-7
[15] G. Chen; Y. H. Lin; J. Wang. Microchip Capillary Electrophoresis with Electrochemical Detection for Monitoring Environmental Pollutants. Curr. Anal. Chem., 2006, 2, 43–50. DOI: http://dx.doi.org/10.2174/157341106775197439
