Capillary Gel Electrophoresis
Capillary Gel Electrophoresis (CGE)
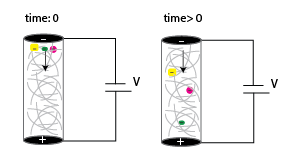
Capillary Gel Electrophoresis (CGE) is an analytical separation method where charged molecules are separated in capillaries filled with porous gel matrix. CGE is basically an adaptation of the traditional slab gel electrophoresis to the capillary electrophoresis (CE) method for its advantageous features. CGE is used to separate large biological molecules like protein, DNA, and RNA. In free solution, these molecules have similar electrophoretic migration rates due to similar charge-to-mass ratios. However, in CGE, the non-convective medium allows them to separate based only on their size. The general protocol follows four steps: 1) The start and end vials and the connecting capillaries are filled with electrolyte solution, 2) The sample to be separated is introduced in the capillary, 3) Electric field is applied and the analytes migrate in the opposite charge direction, 4) Samples and separations are detected via various modes depending on the experimental setup. The sieving matrix could be selected from a diverse array of gel materials based on the experimental requirements. At this scale, there is a smaller sample volume and reagent requirement, the analysis is done in a shorter time and offers higher resolution, the separation is more efficient, and the analyte concentration is quantified more reliably through UV absorbance, fluorescence, or mass spectrometry. The disadvantages come from the difficulty in building the device, and the restriction in the sample size. Figure 1 shows the migration of negatively charged macromolecules through the porous gel via an applied voltage. The smallest macromolecule migrates the fastest while the largest one migrates the slowest.
Basis of Electrophoretic Separations
Electrophoresis is the migration of ions in a system where an electric field is applied. Force applied by the Electric Field (FEf) is proportional to its effective charge (q) and the electric field strength (E)
- FEf = qE (1)
The movement of the ion is faced by a retarding frictional force (Ff), which is proportional to the velocity of the charged molecule (Vep), and the friction coefficient (f). In Gel Electrophoresis, friction is correlated with the sieving matrix and its pore size.
- Ff = f Vep (2)
This means that the charged molecule reaches a steady state velocity in its movement where the accelerating force equals to the frictional force.
- FEf = Ff (3)
Therefore, we can write:
- qE = f Vep (4)
If we rearrange:
- Vep = qE / f = µepE (5)
In this equation, µep (electrophoretic mobility) refers to a constant of proportionality between the velocity of the molecule and the applied electric field strength. µep is proportional to the charge of the ion and inversely proportional to the friction coefficient.
The friction coefficient f is correlated with the hydrodynamic radius of the charged molecule (r) and the viscosity of the medium (η) and can be written as:
- f = 6πηr (6)
This shows that a larger radius of the ion has a lower electrophoretic mobility. In summary, larger charged molecules migrate slower whereas the smaller molecules migrate faster.
Gel Types
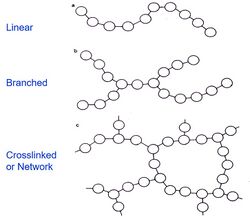
The sieving matrix in the capillaries can be selected, based on separation needs, between permanent (chemical) gels and reversible (physical) gels. The polymer network reduces the adsorption of the solute to the capillary wall and the solute diffusion rate; resulting in the suppressed electroosmotic flow of water. These features increase efficiency; a short column is sufficient to achieve efficient separation.
Physical Gels
Physical gels are networks of molecular entanglements or secondary forces including ionic, hydrogen bonding or hydrophobic interactions. All the physical interactions that take place prevent dissolution; but are reversible and can be disrupted by changes in physical conditions. (Figure 2).
Chemical Gels
Chemical gels are prepared by covalent crosslinking, have defined and controllable pore sizes, and produce high-resolution separations. Poly-acrylamide and PEG gels are commonly used with pore sizes that go as low as 1 nm, depending on the experimental requirements. Poly-acrylamide gels are widely used and preferred due to their electroneutrality.
Advantages of Capillary Gel Electrophoresis
Capillary Gel Electrophoresis (CGE) follows the theoretical principles of slab gel electrophoresis (SGE). In typical slab gel electrophoresis, an electric field is applied through a porous gel matrix and molecules (DNA, RNA, protein) are separated based on their size: larger molecules move slowly through the sieving matrix, while smaller molecules migrate faster. The main disadvantage of the SGE is joule heating as a result of the solution resistance to the applied current. Using capillaries instead of a slab gel provides controlled heat dissipation.
- H=VIt (7)
The heat produced is proportional to the voltage (V), electric current (I), and time (t) it takes to run a gel. In SGE, the heat build-up in a 10*15 cm gel cassette cannot be effectively dissipated from the system. As a result, the porous gel matrix may melt or the density gradients in the gel can alter the efficiency of separation. Whereas in CGE, the capillaries have diameters around 20-100 µm (large ratio of surface area to volume); therefore the capillaries can dissipate heat more efficiently. When potentials go up to ~200V for a typical slab gel, current technology uses CGE to apply high potentials (up to 30 kV) for fast and efficient separations.
Capillaries
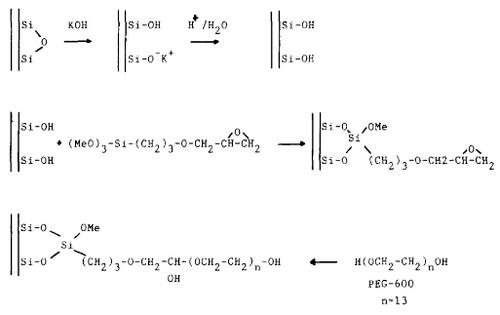
The inner coating of the capillaries should be selected so that electroosmosis is prevented and the analytes will not interact with the capillary wall. Naked fused capillaries have disadvantages since they show electroosmosis above a pH of 3 and conclude in the buffer migration towards the cathode. To avoid this, the inner silica surface is covalently coated for stability. Figure 3 shows the preparation steps of a typical Si-C bond coating. The capillary is first etched with KOH solution and rinsed with water. After, it’s flushed with hydrochloric acid to remove K+ ions to expose free silanol groups on the wall surface. After drying, a solution of trimethoxysilne is pumped in and the unbound region is flushed away. By another reaction carried out, an epoxide group is opened with a solution of PEG and boron trifluoride. Then the capilleries are rinsed with DI water. This process will help decrease a significant amount of adsorption.
Applications
CGE is mainly used for separation of molecules with similar mass/size ratio, but different sizes. Therefore, the separation of nucleic acids and proteins has been very common through this method. Some of the groundbreaking research in molecular biology in the past decades took advantage of the capillary gel electrophoresis method.
CGE was used in the human genome project, employing a physical gel, after a long period of frustrations with chemical hydrogels. The stability problems of the cross-linked polyacrylamide gels were halting the application of this separation method to DNA sequencing. If CGE were to be used in sequencing the human genome, the polymer matrix in the separation capillary had to be replenished in an automated system. In the early 90’s, it was shown that through the use of non-crosslinked polymer matrices and high pressure, the polymer could be replaced entirely. Thus, the human genome project used replaceable linear polymer matrices for automated large-scale DNA sequencing.[5]
For the separation of proteins, their properties must be taken into account. While different proteins vary in size; they also vary in charge. In order to separate proteins based on size in CGE, proteins are treated with a detergent SDS that allows their native charge to be masked. Since SDS molecules are highly negatively charged, the native charge of the proteins can be masked. This technique is analogous to the SDS-PAGE method.
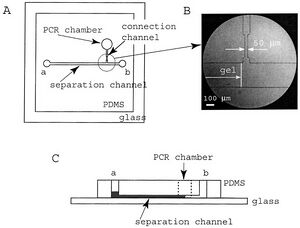
The biotechnology industry uses CGE method for separation analysis of biomaterials on a routine basis with microchips. As other processes must be run prior to CGE for protein and nucleic acid analysis, microchips provide the opportunity to run these assays concomitantly on the same platform. This allows the cost of the experiment to be low during the analytical trials. In addition, CGE provides an alternative to liquid chromatography with higher resolution quality and ability to be incorporated into a microchip. Figure 4 shows a classic PDMS microchip that incorporates two parts: a PCR chamber to amplify the DNA, and a separation chamber to separate the DNA strands.
References
1. Xu, Y. Tutorial: Capillary Electrophoresis. Chem. Educator 1996, 1, 1-14. http://dx.doi.org/10.1007/s00897960023a
2. Camilleri, P. Capillary Electrophoresis: Theory and Practice. CRC Press: Boca Raton, 1993.
3. Landers, J. P. Handbook of Capillary Electrophoresis. CRC Press: Boca Raton, 1994.
4. Ewing, A. G.; Wallingford, R. A.; Olefirowicz, T. M. Capillary Electrophoresis. Anal. Chem. 1989, 61(4), 292A–303A. https://doi.org/10.1021/ac00179a722
6. Whatley H. Basic Principles and Modes of Capillary Electrophoresis. In: Petersen J.R., Mohammad A.A. (eds) Clinical and Forensic Applications of Capillary Electrophoresis. Pathology and Laboratory Medicine. Humana Press: Totowa, NJ, 2001. https://doi.org/10.1007/978-1-59259-120-6_2
7. S. F. Y. Li. Capillary Electrophoresis—Principles, Practice and Application. Elsevier: Amsterdam, 1992. https://doi.org/10.1002/bmc.1130070215
8. Karger B. L.; Guttman A. DNA sequencing by CE. Electrophoresis 2009, 30(S1), S196–S202. http://dx.doi.org/10.1002/elps.200900218.
9. Hebenbrock K.; Schügerl K.; Freitag R. Analysis of plasmid-DNA and cell protein of recombinant Escherichia coli using capillary gel electrophoresis. Electrophoresis 1993, 14(8), 753-8. doi: 10.1002/elps.11501401118.
