Dahlquist:GenePix Pro Software Protocol
From OpenWetWare
Jump to navigationJump to search
Using GenePix Pro
This page shows the hardware settings and protocol for finding and analyzing the features of the GCAT and Ontario type microarrays in the Dahlquist Lab.
See also Dahlquist:DNA Microarray Protocol and Dahlquist:Microarray Data Analysis Workflow pages.
GenePix Pro 6.0 Software Manual
Scanner Settings
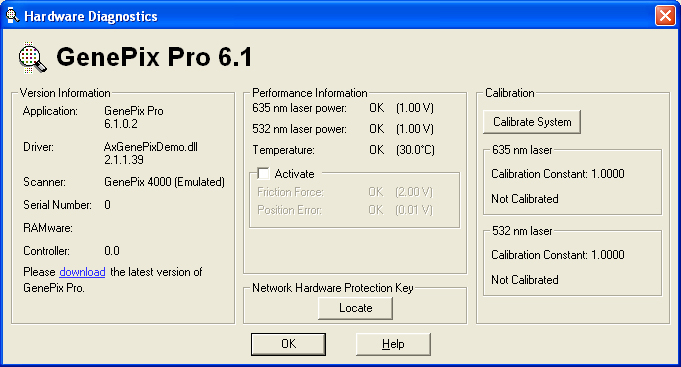
- Version information for the GenePix Pro software used to scan chips in the Dahlquist Lab.
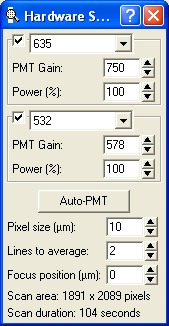
- Hardware settings:
- We use a pixel size of 10 microns.
- We use two lines to average.
- We use a focus position of 0 microns.
- We use Auto PMT settings at 100% power (see below).
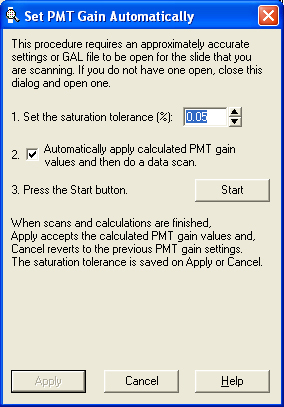
- Auto PMT settings:
- Saturation tolerance is set to the default value of 0.05%.
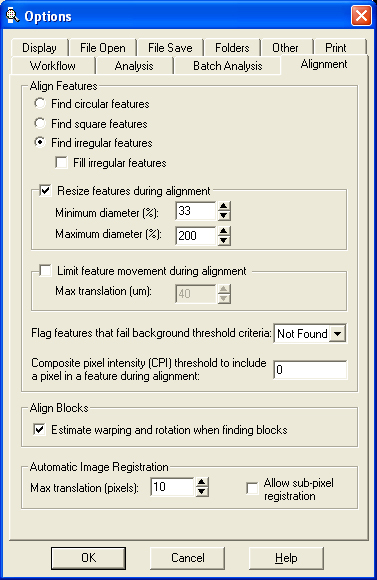
- Alignment Options:
- Find irregular features
- Resize features during alignment to minimum diameter 33% and maximum diameter 200%
- Flag features that fail background threshold criteria as Not Found
- Estimate warping and rotation when finding blocks
- Automatic Image Registration: Max translation 10 pixels.
- Saving scanned images:
- Create a folder on the C: drive called "MicroarrayData_yyyymmdd" where yyyymmdd is the current date.
- Save images to this folder as "Single-image TIFF Files (*.tif)". Do not use TIFF LZW compression (lossless).
- Name files with a Date prefix and Barcode prefix.
Gridding and Generating Intensity Data
- Launch GenPix Pro 6.1 (select Analysis Only)
- On the right hand side of the screen, select the File icon
- Select File > Open Images
- Navigate to the folder containing your .tif images
- Hold down the Control key while clicking to select both the 532 and 635 wavelength files
- Click on the Open button
- Use the brightness and contrast sliders on the upper left hand side to adjust your image to see all of the spots
- You can use the magnifying glass icon to zoom in on an area of the image, and the magnifying glass return icon to go back to the whole chip image
- Zoom in so that you can see the entire top block of grids on your screen
- On the right hand side of the screen, select the File icon
- Select Load Array List
- For the "Ontario" chips, select the file Ontario_Y6.4Kv7.GAL
- For the "GCAT" chips, select the file yeast_GCAT_gal.gal.
- To download this file, right-click on the link and select "Save link as..." from the context menu that appears.
- Click the Open button
- Using the icon that has an arrow and a square, drag the grids so that they are approximately aligned with the top block of spots
- Note that the .gal file for the GCAT chips only has grids for one half of the array, so this process will have to be performed twice, once for the top set of spots and once for the bottom set of spots on the chip.
- Click on the icon that looks like a compass, or a circle with cross-hairs, and select Find Array, Find All blocks, Align Features
- Alternately, you can do this in three steps by:
- Click on the icon that looks like a compass, or a circle with cross-hairs, and select Find Array
- Click on the icon again, and select Find All Blocks
- Click on the icon again, and select Align Features in All Blocks
- Alternately, you can do this in three steps by:
- On the right hand side of the screen, select the icon that looks like a table of data (it says B, C, R at the top of it)
- Click on the File icon on the right hand side and select, Save Results As, and save your results as type "GenePix Pro Files (*.gpr)". Save a JPEG image containing all analyzed features.
Generating an Array Quality Report
- Launch GenePix Pro 6.1 (select Analysis Only)
- On the right hand side of the screen, select the File icon
- File > Open Results
- Select the desired file
- Select Report tab at the top of the screen
- On the left side of the screen, under Navigate, select the back arrow or home icon
- Select Array Quality Control
- Adjust Vital Statistic Thresholds values to:
Quality Control Limits Median signal to background > 2.5 Mean of Median background < 500 Median Signal to Noise > 4 Median % >B + 1 StdDev > 90 Feature variation < 0.5 Background Variation < 1.2 Features with Saturated pixels < 3.3% Not Found < 18% Bad Features < 7%
- Select Start
- Select Show Printable Version
- File > Print
- Under Select Printer > Select Adobe PDF
- Select Print
- Save on the Desktop
- Name the file: ChipBarcode#_yyyymmdd
- The PDF file will either open on its own or you need to open it from the file you saved it to
Next Steps
- Go to the Microarray Data Analysis Workflow page for the next steps.