Combining Body on a Chip - Recreating the Tumor Microenvironment - Organ on a Chip - Michele Caggioni
Introduction
Body-on-a-chip, Organ-on-a-chip.png, and Tumor-on-a-chip all represent a consolidation of microfluidic technologies and biological practices in efforts to model body processes on a higher level of detail and complexity. Through the development of Lab-on-a-chip devices, these microfluidic devices have opened a door that could allow for more human-specific research to occur.
Microfluidic technology offers a number of engineering advantages that make organ-on-a-chip systems versatile and useful. These devices often have external dimensions less than an inch and generally contain channels that have at least one dimension that is less than 1 mm. The small size of microfluidic devices results in a need for less reagents and experiment time than traditional systems.[1] Additionally, these lengths scales typically produce fluidic systems with Reynold’s numbers in the 1-100 range. This results in the creation of extremely controllable laminar flow, which can be used to impart precise physical forces, such as wall shear stress, cyclic strain, tensile and compressive forces on cells when used in organ-on-a-chip applications.[2] This is advantageous when trying to mimic the function of an organ by producing physiologically relevant mechanical forces that trigger a biological response. Microfluidic organ-on-a-chip systems can be designed with detailed control of experimental parameters, especially through the construction of mechanical and chemical gradients.[2] Additionally, microfluidics can be used to model specific diseases such as cancer. Using microfluidics, the effect of biochemical gradients and other non-cancerous components of the tumor microenvironment on cancer proliferation can be studied. Microfluidics also promises to be a powerful platform for in vitro pharmacokinetic studies of cancer therapeutics by allowing for drug transport properties to be studied in a 3D setting more akin to in vivo conditions.
Limitations of Current Preclinical Techniques
Modern advancements in molecular biology and genetics have allowed for the identification of an abundance of novel drug targets. However, many of these targets fail phase II and III clinical trials due to a lack of clinical efficacy. One of the major contributions to the ever-increasing failure rates is the dependency on in vitro (cell culture) and in vivo (animal experiments) as a means to effectively judge early efficacy, toxicity, and pharmacokinetics.[3]Animal models that are executed well provide a reasonable means to test efficacy, but less than 8% of all cancer drug trials in animals end up having a successful transition to human trials. [3] Animals, predominantly rodents such as mice, are commonly used as a means to simulate human disease. However, mice models of cancer (amongst many other diseases) fail on several levels, with issues ranging from a genetic level, where 41% to 81% of over 4,000 shared genes had differing binding sites [4] , to only being able to replicate a set of disease pathologies without including the entire spectrum of physiological changes that occur in humans. Thus, there was a need to introduce a method of replicating human processes that was ethical, replicable, and precise.
Engineering Advantages and Limitations
Conventional Systems
The development of traditional three-dimensional (3D) cell culture systems has expanded the capabilities to model physiological environments in vitro, elucidating the function of organs, tissues, and diseases. Conventional systems utilize hydrogels, tissue-engineered constructs, and macroscale bioreactors to perform mechanical, biological, and chemical processes.[2] Many studies are conducted using organoids, miniature and simplified organs, which model the mammary gland,[5]intestines,[6] and brain,[7] among others. Organoids can be constructed by inducing clustering of polarized organ-specific cells which interact with each other in the ECM of a hydrogel, for example. The impact of contractile modifiers on micro-cardiac muscle tissue has been analyzed [7],[8],[9] as well as models of heart failure and cardiomyopathy.[8],[9] The 3D nature of conventional 3D culture systems has permitted these developments and offers several other advantages.
The physiological structure of various organs partially facilitates their function, which often can be only be modeled by 3D systems. Examples include the cognitive ability of the brain and the mechanical responses of bones, ligaments, and tendons.[2] The use of traditional organoids and tissue sections allows for recreation of the spatial heterogeneity which is present in the body, including the diverse tissue interactions in the lungs.[2] Drug performance can also be elucidated using macro culture chambers or bioreactors, which can be fluidically connected to model interactions between organoids.[2] Bioreactors and tissue-engineered constructs can also be advantageous when large samples are desired for analytical protocols such as mass spectroscopy.[2] However, the simplicity of many conventional systems limits their potential to understand the complex function of physiological milieus.

Organ on a Chip Advantages and Disadvantages
Organ-on-a-chip Advantages
Organ-on-a-chip is said to be promising as it attempts to imitate human organs that the traditional 2-D cell culturing methods and animal models fail to accurately predict. Organ-on-a-chip opens to many possibilities of combining drug testing, cell interaction visualization, micro sensing of cell behaviors, that the other two traditional methods cannot achieve. The 3-dimensional shape, microstructures, and flexibility of the chips can mimic the human organ environment better. Organ-on-a-chip can also be combined with other techniques such as confocal microscopy, fluidic system that can optimize microchannels shapes that can enhance the nutrients and oxygen delivery to the cells. Another advantage that microfluidic chips hold is that crucial factors which are parts of the cell nurturing environment can be controlled and changed for testing for different purposes. Simple designs of microfluidics chips for simple simulations are easily made, the materials to make the chips are inexpensive (except human bones and human cells but those can be replaced with mice’s femur bones and cells or other animals’ bones and cells). Further cancer chemotherapy can be tested with microfluidics chips as drugs can also be inserted into the niches along with nutrients inside the culturing medium.[10]
Organ-on-a-chip Disadvantages
Alongside with the promising advantages, microfluidic chips may not reconstitute entire human organs as there are many other biomolecules inside the human body that are not available to insert in the microfluidics chips. The sizes of the microfluidics chips might not be to scale with the human organs and sizing matters because cells might behave differently with different volumes of fluids. Plus, microfluidics alone cannot sufficiently imitate every human organ. For example, heart-on-a-chip or nerves-on-a-chip are required to have some sort of electrochemical environment for precise simulations.
Body-on-a-chip
Body-on-a-chip devices are the integration of multiple/different organ-on-a-chips that provide a unique opportunity to replicate complex, human-level organization. These microfluidic devices allow for the mimicking of organ-specific 3D architectures and exquisitely precise control of enabling fluid flow, nutrient and growth factor supply, carbon dioxide and oxygen concentrations, as well as adequate waste removal, in a human-level. Body on a chips are best suited to study pathophysiologies that are largely related to 3D microarchitecture and perfusion, as well as modeling acute disease (<1month timeframe).[11]This control of environmental factors allow for cells to not only survive in culture for longer, but also allow for an ability to introduce experimental factors such as drugs and disease in an environment that more closely mimics human organ function. Currently, these devices are best suited to study pathophysiologies that are largely related to 3D microarchitecture and perfusion, as well as modeling acute disease (<1 month timeframe). [11] By utilizing biomimetics, biotechnologists and chemical engineers have been able to replicate the cell architectures of: liver, kidney, intestine, lung, heart, smooth and striated muscle, fat, bone, marrow, cornea, skin, blood vessels, nerves, and the blood-brain barrier. [11] One of the major drawbacks of organ-on-a-chip devices is that they are generally not a good example of overall organ function, as there is a limited variety of the types of cells included. However, the relative heterogeneity of cell types allow for a more specific analysis of the physical effects of flow and shear, as well as cell-type-specific interactions with organ function, disease, and drug toxicity. This allows for more accurate in vitro predictions of the pathophysiological mechanisms of organ function, disease, and drug efficacy and toxicity screening before human trials.

Liver-on-a-Chip

One of the best examples of utilizing organ-on-a-chip devices to study drug efficacy and toxicity in a human body is the liver-on-a-Chip. The liver is especially sensitive to drug-induced injury, and liver toxicity is one of the leading reasons for discontinuing clinical trials, and is the cause of a majority of costly late stage drug trial failures[12], [13]]. Previous in vitro studies struggled to adequately replicate the organ environment, as the general strata of the liver is described as areas of high homotypic cell density with a complex apico-basal polarity that allows for the rise of the bile canaliculi (BC) network,[14] as well as a prolific sinusoidal blood vessel network. While the BC vessel network development is mainly supported by the 3D configuration of hepatocytes, the basic structure of the sinusoidal vessels consist of endothelial cells and occasional Kupffer (immune) cells. Thus, when creating the Liver-on-a-Chip, one of the highest priorities is to address and support the 3D structure of hepatocytes and their intimate relationship with sinusoidal endothelial cells.[15] The iPSC-HPs (induced pluripotent stem cells - hepatocytes) are separated from the medium by an endothelium-like barrier, and cell-cell interactions allowed for this particular device to maintain viability for over four weeks.[14] Physiological markers of cell function (e.g.- Albumin, AST, ALT, etc.) were used as a benchmarks to measure health and survivability of the cells,[13] and the deviations in these results were used as a method to determine drug uptake and subsequent cell death in studies investigating drug-specific hepatotoxicity.
Lung-on-a-Chip
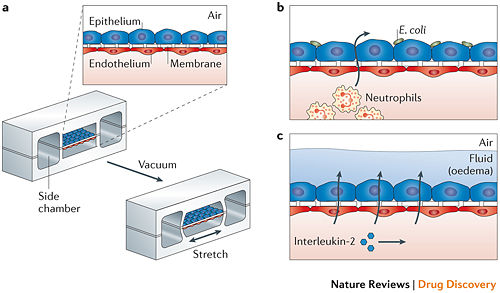
While the Liver-on-a-Chip was an excellent device to test drug toxicity using human hepatocytes, the 3D structure remains relatively straightforward. An example of using organ-on-a-chip to more effectively mimic and study 3D structure in a way that was previously inaccessible is the Lung-on-a-Chip. The fundamental functional units of the lung are the alveolar cells and the complex and intimately involved capillary network, but efforts to replicate this process in vitro struggle to capture the mechanical movement implicit in lung function.[16] In vitro studies also struggled to replicate the alveolar-capillary interface, as it required growth of cells on both sides of a membrane. With the use of a microfluidic device that utilized a flexible membrane and a vacuum with flanking chambers was able to replicate movement and structure that closely resembled that of a human lung. The device, like many others like it, consisted of a porous, flexible membrane that had alveolar epithelial cells grown on one side and pulmonary microvascular endothelial cells on the opposite side.[17] Using a device that replicated the flex-relax cycle of breathing lead to a number of kinetic-influenced discoveries, such as in one trial there was a several-fold increase in the uptake of silica nanoparticles than the conventional methods of cell culture described. In another Lung-on-a-Chip experiment, this time studying the cancer drug interleukin-2 (IL-2) and the pulmonary edema that occurs when high levels of IL-2 are administered, it was discovered that purely mechanical forces contributed significantly to the development of vascular leakage and subsequent pulmonary edema.[18] Through creating a device that was able to replicate the structure and movement, developments surrounding previously-unknown factors are now coming to light, which will further influence the way medicine will approach the development of lung disease and the role movement plays.
Kidney-on-a-Chip

Kidney-on-a-Chip effectively combines the leading motivations behind the developments of Lung- and Liver-on-a-Chip, as the kidney faces both structural 3D microenvironments and issues with drug-induced renal toxicity and injury. Unfortunately, advancements in creating an effective and relevant Kidney-on-a-Chip are hindered by the cellular composition and 3D architecture of the kidney, as it presents a fairly significant translational barrier. There are more than ten renal cell types[18] arranged in an increasingly complex 3D structure, which is further compounded by a complex vasculature system. Even the main function of the organ presents as a complex issue, as there is filtration by the glomeruli and a two-way active secretion and reabsorption process in the tubular apparatus, all occurring in tandem. However, these uptake processes are central in understanding the mechanism of drug-induced renal toxicity and quantifying sensitivity to toxic agents.[19] Another issue that plagues kidneys-on-a-chip is the dependence on nonhuman cell lines such as the Madin-Darby canine kidney (MDCK) cell line as cellular basis for the chips, as immortal human cell lines struggle to express central transporter families and are transitioning away from epithelial cells to mesenchymal cells.[20] Freshly derived human renal cells show promise, but there are issues with donor variability and maintaining their specificity as time goes on. Nonetheless, chips have been developed using a multi-level flow through design. The basic structure of the proximal tubule is replicated by sandwiching a layer of human proximal tubular epithelial cells on an ECM-coated porous membrane, then exposing both sides of the membrane to constant flow.
Kidney-on-a-Chip is a great example of how much room for growth the organ-on-a-chip field has. While the liver and lungs have a fairly homotypic cell type, the kidney has a significant increase in complexity and works as a great example for one of the major barriers that these microfluidic devices face.
Body-on-a-chip (integration of multiple organ-on-a-chips) Advantages & Limitations
Body-on-a-chip Advantages
The introduction of organs-on-a-chip and combining multiple organ-on-a-chips have revolutionized the field of physiological modeling in vitro, predominantly because these microsystems allow for precise control over several experimental variables simultaneously. For example, the combination of specific cell types and their location relative to other cell types elucidates how different kinds of tissue interact.[2] Not only this, but controlling the position of experimental elements, such as cell cultures, makes it much easier to integrate visualization technology, including fluorescence microscopy, microflourimetry, and analytical assays than traditional systems.[2] Precise control over the organ-on-a-chip microstructure is also beneficial when developing computational fluid dynamic (CFD) models, which may be used to study the function of cells, metabolites, and gases based on their CFD behaviors.[2]
One crucial advantage to using body-on-a-chips arises from the ability to control fluid flow in these systems. This is important because fluid interactions are prevalent in the body and affect the function and endurance of many cell types. By using organs-on-a-chips, cell cultures can be sustained for extended periods of time; human lung cells have been sustained in culture on a microfluidic chip system for over a month.[21] Cells respond to mechanical and chemical stimuli. Flow in microsystems engineering allows for easy establishment of mechanical and chemical gradients to determine how cells respond to fluid forces, cytokines, hormones, and gases.[2] If the cells being analyzed circulate physiologically, such as circulating tumor cells or bacteria, flow allows for a more accurate model of what is expected of these cells’ functions in vivo.[12] Not only do microsystems help accurately replicate cell, tissue, and organ function through the precise control of various parameters, but they can also produce larger sample sizes than conventional systems, allowing for more statistically significant experiments.[2] This is especially useful for performing drug testing on organs-on-a-chip.

Body-on-a-chip Limitations
Although the benefits of microsystems are prominent and still expanding, there are a number of technical challenges that must be overcome to advance the technology. Fabricating microfluidic chambers is a well-understood process. However, it usually requires advanced microfabrication technology, which can be expensive. Common fabrication issues, such as bubble formation within microstructures, can be detrimental to the survival of cells and the function of the microsystem.[2] When culturing cells in microchannels, seeding consistency is an issue, as is contamination100. Furthermore, once seeded, it is often difficult to ensure that cells will interact with each other and with a synthesized matrix in the same way that they would in vivo.[2]
Given these challenges, microfluidic technology still offers enormous potential toward elucidating the role of physiological environments in vitro. The next step in evolving this technology will require the junction of functional units of organs to recreate entire organ systems, which can then be joined by synthetic micro vessels coated with endothelial monolayers to form a more complete replica of the human body, which has coined the phrase “body-on-a-chip” 100. Commercially, this will have a huge effect on drug testing trials. By simulating the physiological environment and providing statistically significant experiments, organ-on-a-chip technology may be used instead of animal or human trials to evaluate drug performance.
Comparing Transport Properties of Doxorubicin and Doxil on Microfluidic Device Such a device was created by the Forbes lab at UMass Amherst. The dimensions of the device were 1000 mm x 300 mm x 150 mm, and a microfluidic channel, 250 mm wide and 150 mm tall. Experiments were conducted using human colon carcinoma spheroids. The device was designed to mimic the interface between a tumor and surrounding vasculature.[22]



Tumor spheroids in the device were treated using Doxirubicin and Doxil. Each drug was delivered with a green fluorescent stain to help track uptake. The accumulation of the drug within the spheroids positively correlated with apoptosis (cell death). In the case of doxorubicin, it was observed that the drug has a slow clearance rate (20 hrs) and a corresponding higher extent of apoptosis than doxil, which cleared in only 7hrs.[22]
Future Work and Ultimate Goals
Microfluidic devices have the potential to serve as analogs for organs in vitro. As the technology matures further and human in vitro tissue culture becomes easier, full organ functions could be modeled in devices. This would create a platform for an in depth analysis of disease mechanisms as well as the power of new therapies. By integrating these microfluidic organs, entire organ systems or even a complete body could be modeled in vitro on a microscale. This would allow for an unprecedented ability to examine the effects of diseases and drugs across the entire human body.
Fabrication of an organ-on-a-chip
Different microfluidic devices that imitate lungs, kidneys, hearts or guts have been broadly researched. The general steps to create a specific organ on a chip are principally the same. The first steps are to design, mold and sterilize the device. Specific types of cells for the organ chips are perfused to the device with culture media through small inlets that are connected to small tubings. Cells are continuously grown inside the chip in a sterile environment. The model is often screened under the microscope for cell population check and once the cells divide enough, the chips will go through chemical tests such as cancer drug testing and drug screening.[23]
Integration of multiple organ-on-a-chip - Body-on-a-chip
Once multiple organs-on-a-chips are extensively tested, and integration of multiple organ-on-a-chips would be the final goal to work on. Human-on-a-chip is one of the most important and most promising alternatives to animal testing as a human-on-a-chip is expected to perform as an entire human organ system (such as digestive system, respiratory system, cardiovascular system and so on) or even further, an entire living human. However, there are many challenges for the integration of all organ-on-a-chips because we have to ensure that once different chips are connected, they must support others like how human tissues support each other. Supporting computational models are needed to design an entire integration of organ-on-a-chips. In addition, developing a human-on-a-chip requires much more donated human cells and human materials (such as human bones) which are very expensive to get for research. Despite the challenges, human-on-a-chip is still a brilliant idea to be considered as once those human-on-a-chips become true, medical researches such as cancer treatment or drug testing no longer have to depend on animal testing.

Introduction-Modeling Tumor Death: Microfluidics and Biomimetic Tumor Spheroids

Cancer accounts for the highest cause of death in economically developed countries.[24] As a result, a substantial amount of effort has been focused on developing novel therapies to treat the disease. Unfortunately, drug efficacy is only tested with high throughput monolayer cell death assays, which do not account for transport barriers encountered in vivo.[25][22][26] Microfluidic devices can model in vivo tumor biology while maintaining the high throughput capacity of monolayer cell death assays.
Overview of Cancer Pathology
Cancer follows the following distinct path: Benign growth, Infiltrative growth, vascularization, and finally metastasis. Initially, premalignant masses grow in a benign manner and are confined to a distinct area of the respective organ. After this phase of growth, many tumors cells remodel the extracellular matrix and become increasingly invasive. These tumor cells commonly enter lymphatic vessels and colonize near and distant lymph nodes. In the later stages of the disease, primary tumor cells break off from the main mass, enter blood vessels and colonize distant organs. This metastasis of cancer cells leads to organ failure and eventual death.[27][28]
Modeling Cancer: Tumor-on-a-Chip
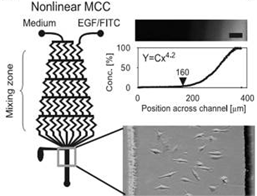
Modeling Biochemical Regulation of Cancer Proliferation The growth, differentiation, and migration of tumor cells are regulated by the biochemical and biophysical properties of the tumor microenvironment. Many of these processes are controlled through various biochemical gradients which can be modeled using microfluidics. There are two main methods by which this is accomplished: flow-based methods and diffusion based methods. The former utilizes convective forces with in laminar flow regimes to produce chemical gradients, while the latter uses passive diffusion of molecules to form gradients201. For example, multiple studies have been conducted on the effects of Epidermal Growth Factor (EGF) gradients on cancer cell chemotaxis[20] (Fig. 1)
Modeling Non-Cancerous Cell Components of the Tumor Microenvironment
Fibroblasts are connective tissue cells found to be associated with all stages of cancer proliferation. Fibroblasts are recruited by tumor cells to produce specific growth factors. Microfluidics have been used to simulate these interactions in vitro. For example, Qin et al. fabricated a multichambered device for the co-culture of fibroblasts and tumor cells while allowing for diffusional interactions between parallel channels.[29]

Modeling Drug Delivery: Overview and Motivation
Microfluidics can be used to model pharmacokinetics in a 3D environment. Microfluidics provide a promising alternative to monolayer or mouse experiments for predicting drug efficacy in vivo. Traditional monolayer experiments only examine cytotoxicity in a 2D environment. This ignores the transport properties of the drug, which can significantly impact drug efficacy. Upon intravenous delivery, the blood plasma concentration of a drug is at its highest value. The plasma concentration then decreases over time via absorption by cells and metabolism. The drug spreads through tumors via diffusion, and may bind to target receptors. Once the plasma concentration of a drug falls below the cellular concentration, the drug is cleared from the system. The cytotoxicity of a drug is effected by the combination of these transport processes.[22] Drugs that demonstrate cytotoxicity in monolayer are generally then tested in mice. However, there is generally no correlation between efficacy in monolayer and efficacy in mice. Moreover, experiments in mice are expensive and time consuming. Using microfluidics, the transport of a therapy from the blood vessel into tumors can be simulated.[22]
Current High Throughput Drug Testing Methods

Cancer cells have been taken from patients, immortalized or conditionally reprogrammed and cultured in monolayer reliably.[31] This is the main reason why monolayer cell culture assays are commonly used to test cancer therapies in high throughput. Out of all the drugs that are screened, less than 5% of drugs make it through pre-clinical trials and of those, only 10% make it through clinical trials.[32][33]Monolayer assays do not account for important transport limitations imposed on in vivo tumors.[22][26] This poses a problem because larger molecules have difficulty penetrating and treating tumor cores, which are the most drug resistant regions. Unlike microfluidic devices, monolayer assays cannot properly assess drug efficacy within the drug resistant quiescent and necrotic regions of tumors which contributes to cancer relapse.[26] The heterogeneous nature of tumors and significant transport barriers demonstrates a need for improving high throughput drug testing that mimics in vivo conditions.[30]
Role of Microfluidics in Cancer Research Abroad
With the numerous advantages of microfluidics becoming fully realized, researcher labs have embraced microfluidics for a diverse range of applications specific to cancer research. The applications of microfluidics for cancer research can be categorized into four general groups: cell sorting (isolation of cancer cells), controlled generation (from seed to harvest), drug screening, and modeling. In some capacity, there is overlap between the groups as is shown is later examples. With advent of tumor spheroids as a means for improved in vitro modeling, microfluidic applications have shifted towards production and screening of these spheroids.
Cell Sorting

Cell sorting through microfluidics allows for single cell analysis using traditional methods, most notably flow cytometry. Being able to separate different cells is critical for accurate drug screening, medical diagnostics, and cancer cell assays.(With permission from Wlodkowik, Copyright 2010, MDPI)[34] Because micro-sorters possess the capability to sort by a wide range of characteristics (size, charge, shape, binding affinity, etc), they can be installed into virtual any microfluidic device that requires sorting of particles, in this case cancer cells.[34] Once cells are sorted out, fluorescently activated cell sorting (FACS) has proven to be a powerful tool for analyzing cancer cells. Depending on the fluorescent labels being used, FACS can show cell death (stages of apoptosis or necrosis), drug diffusion and permeation, and protein location.[34] Other modes of cell sorting exist such as Raman activated cell sorting (RACS) and fluorescence activated droplet sorting (FADS), which carry their own benefits in terms of chemistry and throughput efficiency.[34]
Cell sorting has also used to isolate circulating tumors cells in large blood samples.[35] The ability to isolate, harvest, and analyze circulating tumors cells creates opportunity to better understand metastasis and other cancer mechanisms. In 2007, a collaborative research team successfully isolating circulating tumors cells by precisely washing blood through a filter of posts that were treated with antibodies specific to the target cancer cells.[35] By precisely flow blood through the micro post filter under laminar flow conditions, the device was able to isolate tumors cells from the sample medium.[35]
Tumor Spheroid Production

Precise 4D(Space-time) laminar flow control, biocompability, and relatively low cost-effectiveness have made microfluidics an ideal application in the culturing, forming, and harvesting of 3D tumor spheroids.[34] In recent years, the methods for tumor formation in a microfluidic device has increased with the advent of tumor spheroids as a viable means for modeling.[34] The two primary modes of tumor production depend on sufficient mixing of a cell suspension to generate natural clustering or forced spheroid growth through geometric confinements (concave interfacial boundaries) as found in poly-hydroxyethyl methacrylate or hanging droplets.[34] (Tumor on Chips)
Very recently, a collaborative research based in Taiwan was able to seed, culture, form, and remove tumor spheroids from a two-layer device through simple laminar flow of seeded medium (in the top layer) over micro wells(in the bottom layer) .[36] To temper the parabolic nature of flow through the channel, seeded medium was manually pumped at flow rates over 100 microliters per minute.[36] The wells were filled with seeded due to gravity.[36] Geometry of the wells was proven to generate consistency is tumor spheroid size and shape.[36]

Research has been done to successfully form tumor spheroids by hanging drop method, a process of allowing gravity and liquid-air tension to form droplets containing cancer cells.[34] These cells grow and aggregate within the liquid droplets to spherical shapes. The uniformity of the spheroids proves inadequate for reliable testing, as spheroid size has shown great effect on drug efficacy.[37] The precise spatiotemporal control on fluid at the microscale has been used to produce droplets of reliably uniform size and shape on the nanometer and micron scale.[34][36][37] This capability was employed using an intersection of cross-flow streams.[37] As the bisecting aqueous stream enters the intersection, the faster continuous stream applies a shearing force on the stream head, essentially cutting off a plug of fluid.[37] These plugs are spherically in shape and consistent in diameter because of the flow control within the device.[37]
As mentioned, the common materials used for microfluidic devices are often compatible with living cells. Polyethylene glycol (PEG) and polydimethylsiloxane (PDMS) are the primary materials being used due to their aversion for protein and cell adhesion.[34] Through the use of photolithography, a well-established method of generating silicon masks for PDMS molding allows for low cost, rapid manufacturing of devices for testing. Within this vein, certain considerations must be made in terms of treating the microfluidic device to improve cell viability. In cases where tumor spheroids are being readily cultured and formed within the device, devices are often sterilized with deionized water, UV light, or ethanol to avoid contamination.[34][37]
Modeling
While three-dimensional tumor spheroids have been shown to provide improved in vitro modeling, work has been done to design microfluidic channels to better replicate tissue vascularity and also to mimic important interfacial cell-to-cell or cell-to-intracellular matrix regions. In the pursuit of understanding cancer metastasis mechanisms, microfluidic devices are designed to replicate these regions, which allows for analysis of biological activity occurring at the microscale that couldn’t be observed in vitro otherwise.
Zervantonakis et al, and researchers from departments within the Massachusetts Institute of Technology replicated the tumor tissue – endothelium interface to analyze and model how tumors cells successfully enter the blood stream.[27] The device consists of a three-channel composition with the channels running parallel and connected at the sides1.[27] In the other channels, endothelial cells and cancer cells are seeded respectively with a hydrogel extracellular matrix in between.[38] As such, experiments can be done to observe how tumor cells respond in real time to changes in microenvironment – adjustments to gradients of nutrients and oxygen or stimulation from spikes in growth factors.[38] Additionally, reasoning for the dependence of a damaged blood or lymphatic vessel to promote cell entry into the blood stream can be made due to the design of the endothelial channel.[38]

Drug Screening
In many cases, the three previously described applications are paired with drug screening to test drug efficacy, cell death, cell diameter reduction, cell membrane integrity, diffusion properties of drugs, among many other important results of drug testing. [34][37][38][39][40] In projects previously mentioned, a common theme of experimentation was to analyze tumor spheroids and monolayer cell cultures after introducing anti-cancer drugs into the systems. The Tawain group that used gravity-packed wells for tumor spheroid production tested three drugs and observed cell diameter and membrane integrity through microscope imagery and fluorescence activated cell sorting respectively.[36] Sabhachandani et al ran two drugs through continuously flowing medium past packed microdroplet-formed spheroids.[37] As proof-of-concept, both groups were able to confirm a significant difference in cell death between two-dimensional monolayer cell cultures and three-dimensional tumor spheroids.[36][37] Microfluidic devices can be designed to recreate the tumor microenvironment in vitro. The devices can accurately mimic transport barriers and the heterogeneous nature of tumors.[30] Fluid flow in both tumor microvasculature and microfluidic devices is laminar. In addition, tumor spheroids can be positioned within the devices to form actively dividing, quiescent and necrotic tissue which resembles in vivo conditions.[26] Due to nutrient gradients present in tumor-on-a-chip devices, cells residing on the tumor interior are nutrient deprived and undergo necrosis while media exposed cells actively divide. This spatial heterogeneity within in vitro tumors improves drug efficacy and transport modelling due to the system’s in vivo resemblance.[30]
Using fluorescent modifications, like FITC, and microscopy, chemotherapeutic treatment can be spatiotemporally modelled for a variety of drugs with a microfluidic device. This experimental data can be used in Fick’s law of diffusion to calculate the effective diffusion constant which describes a drugs transport efficacy. The simplified Fick’s law used for tumor transport modelling is as follows:
[math]\displaystyle{ \frac{\partial C_A}{\partial t}+v\bigtriangledown C_A=D_A \bigtriangledown^2 C_A+R_A }[/math] (1)
With the following boundary conditions:
[math]\displaystyle{ \frac{\partial C_A}{\partial z}|_{z=1}=0 }[/math] (2)
[math]\displaystyle{ \frac{\partial C_A}{\partial z}|_{z=0}=e^{-kt} }[/math] (3)
[math]\displaystyle{ \frac{\partial C_A}{\partial r}|_{r=0}=0 }[/math] (4)
[math]\displaystyle{ \frac{\partial C_A}{\partial r}|_{r=1}=0 }[/math] (5)
And the following initial condition:
[math]\displaystyle{ \frac{\partial C_A}{\partial t}|_{z=0}=C_A0 }[/math] (6)
The boundary conditions (2), (4) and (5) are all no flux boundary conditions due to either cylindrical symmetry or because the tumor is in contact with an impermeable wall and boundary condition (3) pertains to the half life of drug in the media that is contact with the tumor surface. The reaction term of the drug accounts for drug binding and metabolism within a tumor cell and will usually be a negative term.
Experimental data acquired from a tumor-on-a-chip model can be fit to a discretized version of (1) and an effective diffusion constant can be calculated. The calculated diffusion constant will give an indication of how easily a drug penetrates into a tumor. For large drugs, like antibodies, examining drug diffusion into tumors is useful because it provides answers about how well quiescent and necrotic tissue are treated when dosed with macromolecular therapies. These studies have not been performed to date and can provide valuable information about biologic dosing regimens.
Tumor-on-a-chip drug testing must be expandable to allow for high throughput screening. In order to make microfluidic drug testing scalable, the technology must be automated and allow for stringent flow control within a multichannel device. This can be achieved through the use of multilayered, valved microfluidics. To make this device compatible with high throughput screening, automation is required. This can be achieved through the use of programmable pneumatic actuators coupled with valve control channel networks.
Scalability of Tumor-On-A-Chip Based Drug Screening

Scaling out can be achieved by creating a binary, multiplexed system which can selectively target single flow channels. The purpose of scaling out is to create a high throughput device, which can handle many tumor spheroids, as well as distribute them into specific channels. Building a high throughput microfluidic system can be a difficult task if the particular device requires selective valve control, but multiplexed systems offer a structured method to accomplish selective valve control. A multiplexed The multiplexed valving system operates using a binary pattern, where n fluid channels are controlled using 2 log2 n control channels.[41] This limits the specific number of controllable channels within a device to be within a binary domain, however, achieving a multiplexed system allows a more organized microfluidic device. Can be developed to release tumor spheroids into specific channels. A MUX Quake Valve System will be implemented and operated via code written in Python. Devices will be made with 128 flow channels and different 14 valve layers, this will allow handling of tumor spheroids to be conducted in a high throughput manner.[41]
The scalability of a multiplexed system allows for a robust level of design capabilities, and devices could be designed to handle more or less tumors, depending on the experiment. This method is more beneficial to currently used methods because the micro scale nature of these devices result in reduced materials cost and smaller volumes used. Scaling these experiments out will also allow for a much more efficient and less time consuming experiment to be conducted.
- ↑ Amin, R. et al. 3-D Printed Microfluidic devices, Biofabrication, (2016). DOI: 10.1088/1758-5090/8/2/022001
- ↑ 2.00 2.01 2.02 2.03 2.04 2.05 2.06 2.07 2.08 2.09 2.10 2.11 2.12 2.13 Bhatia S. N. & Ingber D. E. Microfluidic organs-on-chips. Nature biotechnology 32, 760–772,(2014). DOI: 10.1038/nbt.2989
- ↑ 3.0 3.1 Mak IW, Evaniew N, Ghert M. Lost in translation: animal models and clinical trials in cancer treatment. American Journal of Translational Research. 2014;6(2):114-118. PMID: 24489990
- ↑ Gawrylewski A. The Trouble with Animal Models. The Scientist. 2007 DOI:10.1017/S0963180115000079
- ↑ Fingleton B. Matrix metalloproteinases as valid clinical targets. Curr Pharm Des. 2007;13:333–46. DOI: 10.2174/138161207779313551
- ↑ Mroue, R. & Bissell, M.J. Three-dimensional cultures of mouse mammary epithelial cells. Methods Mol. Biol. 945, 221–250 (2013). DOI:10.1007/978-1-62703-125-7_14
- ↑ 7.0 7.1 Sato, T. & Clevers, H. Growing self-organizing mini-guts from a single intestinal stem cell: mechanisms and applications. Science 340, 1190–1194 (2013). DOI:10.1126/science.1234852
- ↑ 8.0 8.1 Grosberg, A., Alford, P.W., McCain, M.L. & Parker, K.K. Ensembles of engineered cardiac tissues for physiological and pharmacological study: heart on a chip. Lab Chip 11, 4165–4173 (2011). DOI:10.1039/c1lc20557a
- ↑ 9.0 9.1 Lancaster, M.A. et al. Cerebral organoids model human brain development and microcephaly. Nature 501, 373–379 (2013). DOI:10.1038/nature12517
- ↑ Marturano-Kruik, A., Nava, M. M., Yeager, K., Chramiec, A., Hao, L., Robinson, S., . . . Vunjak-Novakovic, G. (2018). Human bone perivascular niche-on-a-chip for studying metastatic colonization, 115(6), 1256-1261. DOI:10.1073/pnas.1714282115
- ↑ 11.0 11.1 11.2 McCain, M.L., Sheehy, S.P., Grosberg, A., Goss, J.A. & Parker, K.K. Recapitulating maladaptive, multiscale remodeling of failing myocardium on a chip. Proc. Natl. Acad. Sci. USA 110, 9770–9775 (2013). DOI:10.1073/pnas.1309820110
- ↑ 12.0 12.1 Wang, G. et al. Modeling the mitochondrial cardiomyopathy of Barth syndrome with induced pluripotent stem cell and heart-on-chip technologies. Nat. Med. 20, 616–623 (2014). DOI:10.1038/nm.3545
- ↑ 13.0 13.1 H.Y. Tiong, et al. Drug-induced nephrotoxicity: clinical impact and preclinical in vitro models. Mol. Pharm., 11 (2014), pp. 1933–1948 DOI:10.1021/mp400720w
- ↑ 14.0 14.1 Kim, H.J., Huh, D., Hamilton, G. & Ingber, D.E. Human gut-on-a-chip inhabited by microbial flora that experiences intestinal peristalsis-like motions and flow. Lab Chip 12, 2165–2174 (2012). DOI:10.1039/C2LC40074J
- ↑ Huh, D. et al. Reconstituting organ-level lung functions on a chip. Science 328, 1662–1668 (2010). DOI:10.1126/science.1188302
- ↑ Wang S. J., Saadi W., Lin F., Nguyen C. M. C., and Jeon N. L., Exp. Cell Res. 300, 180 (2004) DOI:10.1016/j.yexcr.2004.07.014
- ↑ Hare, P. (2010). Lung on a chip. Nature Biotechnology, 28(8), 816-817. DOI:10.1038/nbt0810-816
- ↑ 18.0 18.1 Esch, E. W., Bahinski, A., & Huh, D. (2015). Organs-on-chips at the frontiers of drug discovery. Nature reviews Drug discovery, 14(4), 248-260. DOI:10.1038/nrd4539
- ↑ Huh, D., Leslie, D. C., Matthews, B. D., Fraser, J. P., Jurek, S., Hamilton, G. A., ... & Ingber, D. E. (2012). A human disease model of drug toxicity–induced pulmonary edema in a lung-on-a-chip microdevice. Science translational medicine, 4(159), 159ra147-159ra147. DOI:10.1126/scitranslmed.3004249
- ↑ 20.0 20.1 Ma H., Xu H. & Qin J. Biomimetic tumor microenvironment on a microfluidic platform. Biomicrofluidics. 7, 11501 (2013). DOI:10.1063/1.4819565
- ↑ Boudou, T. et al. A microfabricated platform to measure and manipulate the mechanics of engineered cardiac microtissues. Tissue Eng. Part A 18, 910–919 (2012). DOI:10.1089/ten.tea.2011.0341
- ↑ 22.0 22.1 22.2 22.3 22.4 22.5 Toley, B. J., Ganz, D. E., Walsh, C. L. & Forbes, N. S. Microfluidic device for recreating a tumor microenvironment in vitro. J. Vis. Exp. 5–9 (2011). DOI:10.3791/2807
- ↑ Huh, D. (2015). A Human Breathing Lung-on-a-Chip, 12(1), S42-S44. DOI:10.1513/annalsats.201410-442mg
- ↑ Jemal, A., Bray, F. & Ferlay, J. Global Cancer Statistics: 2011. CA Cancer J Clin 49, 1, 33–64 (1999). DOI:10.3322/caac.20107
- ↑ Neuži, P., Giselbrecht, S., Länge, K., Huang, T. J. & Manz, A. Revisiting lab-on-a-chip technology for drug discovery. *Nat. Rev. Drug Discov.* **11**, 620–32 (2012). DOI:10.1038/nrd3799
- ↑ 26.0 26.1 26.2 26.3 Walsh, C. L. et al. A multipurpose microfluidic device designed to mimic microenvironment gradients and develop targeted cancer therapeutics. *Lab Chip* **9**, 545–554 (2009). DOI:10.1039/b810571e
- ↑ 27.0 27.1 27.2 Hanahan, D. The Hallmarks of Cancer. *Cell* **100**, 57–70 (2000). DOI:10.1016/S0092-8674(00)81683-9
- ↑ Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: The next generation. *Cell* **144**, 646–674 (2011). DOI: 10.1016/j.cell.2011.02.013
- ↑ Liu T., Lin B., and Qin J., Lab Chip 10, 1671 (2010). DOI:10.1039/C004084H
- ↑ 30.0 30.1 30.2 30.3 Toley, B. J., Tropeano Lovatt, Z. G., Harrington, J. L. & Forbes, N. S. Microfluidic technique to measure intratumoral transport and calculate drug efficacy shows that binding is essential for doxorubicin and release hampers Doxil. *Integr. Biol. (Camb).5, 1184–96 (2013). DOI: 10.1039/c3ib40021b
- ↑ Liu, X. et al. ROCK inhibitor and feeder cells induce the conditional reprogramming of epithelial cells. *Am. J. Pathol.* **180**, 599–607 (2012). DOI: 10.1016/j.ajpath.2011.10.036
- ↑ DiMasi, J. a, Reichert, J. M., Feldman, L. & Malins, a. Clinical approval success rates for investigational cancer drugs. *Clin. Pharmacol. Ther.* **94**, 329–35 (2013). DOI:10.1038/clpt.2013.117
- ↑ Hay, M., Thomas, D. W., Craighead, J. L., Economides, C. & Rosenthal, J. Clinical development success rates for investigational drugs. *Nat. Biotechnol.32, 40–51 (2014). DOI:10.1038/nbt.2786
- ↑ 34.00 34.01 34.02 34.03 34.04 34.05 34.06 34.07 34.08 34.09 34.10 34.11 Wlodkowic, D. & Cooper, J. M. Tumors on chips: Oncology meets microfluidics. *Curr. Opin. Chem. Biol.* **14**, 556–567 (2010). DOI: 10.1016/j.cbpa.2010.07.005
- ↑ 35.0 35.1 35.2 Nagrath, S. et al. Isolation of rare circulating tumour cells in cancer patients by microchip technology. *Nature* **450**, 1235–9 (2007). DOI: 10.1038/nature06385
- ↑ 36.0 36.1 36.2 36.3 36.4 36.5 36.6 Patra, B., Peng, C.-C., Liao, W.-H., Lee, C.-H. & Tung, Y.-C. Drug testing and flow cytometry analysis on a large number of uniform sized tumor spheroids using a microfluidic device. *Sci. Rep.* **6**, 21061 (2016). DOI: 10.1038/srep21061
- ↑ 37.0 37.1 37.2 37.3 37.4 37.5 37.6 37.7 37.8 Sabhachandani, P. et al. Generation and Functional Assessment of 3D Multicellular Spheroids in Droplet Based Microfluidics Platform. *Lab Chip* **16**, 497–505 (2015). DOI: 10.1039/C5LC01139F
- ↑ 38.0 38.1 38.2 38.3 Zervantonakis, I. K. et al. Three-dimensional microfluidic model for tumor cell intravasation and endothelial barrier function. *Proc. Natl. Acad. Sci.* **109**, 13515–13520 (2012). DOI: 10.1073/pnas.1210182109
- ↑ Hirschhaeuser, F. et al. Multicellular tumor spheroids: An underestimated tool is catching up again. *J. Biotechnol.* **148**, 3–15 (2010). DOI: 10.1016/j.jbiotec.2010.05.008
- ↑ Miller, O. J. et al. High-resolution dose – response screening using droplet-based microfluidics. *Proc. Natl. Acad. Sci.* **109**, 378–83 (2012). DOI: 10.1073/pnas.1113324109
- ↑ 41.0 41.1 Thorsen, T. Microfluidic Large-Scale Integration. Science (80-. ). 298, 580–584 (2002). DOI: 10.1126/science.1076996
