Citizen Science/Open Spectrophotometer Project/Application
<html> <body>
<script type="text/javascript"> var sc_project=4270054; var sc_invisible=1; var sc_partition=46; var sc_click_stat=1; var sc_security="d09c9961"; </script>
<script type="text/javascript" src="http://www.statcounter.com/counter/counter.js"></script><noscript>
</noscript>
</body> </html>
INTERACTIVOS?09: Garage Science Workshop-Seminar Application Form
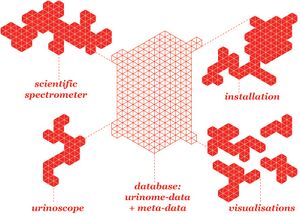
Illustrations
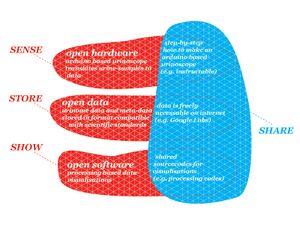
- Generic platform (sensing, storing, showing and sharing)
- Spectrophotometer hardware design (schematics) + arduino pictures
- Database infrastructure + data flow and interfaces
- Workshop installation sketch
- Visualisation output sketch
Project title
- Vincent 08:27, 1 December 2008 (EST): suggestion: "Body Rainbow"
- --SxE00 10:12, 3 December 2008 (EST) Depending on the output we want to go for:
- body shapes
- body colors
- body clock
- body talk
- body language...
- *Tuur Van Balen 19:33, 13 December 2008 (EST):
- spectro body
- you're in [data]
- urination
- you're in body
- body2body
Project summary
We propose 'Urinomics' as the science of piss; the study of urine and the information it can reveal. We'll call this information the 'Urinome'. A 'urinoscope' is the device that translates urine to data using spectrophotoscopy, and we've built a first prototype. In this project, we want to Sense, Store, Share and Show both our individual and collective genome. The result will be a collection of urinoscopes, a data-set accessible to everyone, and an installation that visualises this data.
Project description
Our own vocabulary how do we call:
- the open software spectrophotometer: ???
- the urine spectral data: ???
- the open data set database: ???
Overview

Urine spectral profiling
- personal but not analytical
- track your own profile/signature in time
- compare your own profile/signature with others.
- Urine portrait, urine profile/signature,
Generic platform:
- Sensing / Sharing:
- Arduino-based spectrophotometer
- Open hardware
- Storing / Sharing
- Compatible with scientific data format
- Open XML standard
- Open access, online database
- Showing:
- Open software Processing
- experiments in visualising scientific data
> Spectrophotometer Design
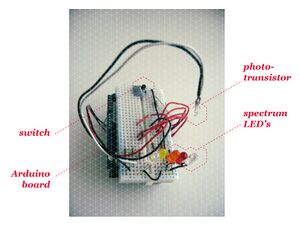

Spectrophotometer, first prototype on YouTube
We've build the first prototype of a simple spectrophotometer using an Arduino, some LED's and other cheap, accessible components. The spectrometer measures the absorption of light at various wavelengths, in our case the 'colour' of the 5 LED's. We will use it to create a data-portrait out of people's urine.
UV/visible spectrophotometry is an essential tool in the identification and quantification of a very broad range of chemical and biological substances. It relies on the measurement of radiation intensity absorption at various spectral wavelength. It has proved particularly useful in biochemical analysis, and is of vital importance in the clinical laboratory where various components of blood and/or urine samples, are determined and monitored on a 24h basis. Building a cheap and accessible (simple) spectrometer can empower and open up possibilities for Citizen Science.
> Open Spectra Data Set
After collection (take a urine sample and run a spectrography), the data will be stored in an open database on the internet. The data differs for each urine sample but isn't analytical, it won't tell anything about the urine or person. The database will contain the spectrophotometer-data and whatever data we want to add to it: the time and place of urination, ... . We call this meta-data.
We'll build an (open) dataplatform using the Google App framework (similar to Django). It provides an easy to develop SDK as well as database resources. We'll make sure the platform and formats we use are compatible with those used within the scientific community. Crossing these boundaries can provide benefit for scientists and citizens as both can engage with the same data: experimenting with new way of using, aggregating or visualising ...
> Visualisation
Because the data from the spectrophotometer feeds into Processing, it opens up a huge amount of 'interactive' visualisation potential. We want to make use of this potential in the design/art community to experiment with new ways of visualising this scientific data. On the other hand, by complying with scientific formats for this data, we want to offer the opportunity to engage with the kind of data that normally isn't accessible.
Visualising this data will result in a shadow-play with spectrometer-data. The data isn't analytic but it is personal and will differ between people and over time, opening up space for interpretation. We imagine it will be like cloud-watching: comparing your urine-clouds with that one from the day before or those of others.
> Installation
1.4.4.1 The installation influences at its turn the outcome of the spectrometry (Body - Feedback loop // Pataphysics machines for the body = patamedicines?)
- a food vending machine activated by urine hands out food based on a spectrograph: vitamin c, beetroot, ...
- a hacked portocabin locks up the visitor and gives sun-bath / UV light (with a countdown showing how long)
1.4.4.2 The installation creates a shape from the data that gets made physically
- data translated into stain (you're in / rorschach inkblot) in processing, gets lasercut out of cardboard/wood and put up in the installation space
- data translated to (2D) barcode that gets printed on a Tshirt so you can walk around with it
1.4.4.3 The installation is biological
- spectrography data translated to 1 or 0 for every point of a 10x10 matrix, installation is a dripping installation on 100 plants (10x10)
- spectrography data switches on and off a bunch of fast-growing lights on top of plants and thus slowly directs the pattern the plants grow
Direction 1.4.4.1 is nice because of the feedback loop to the body, the body being influenced by the installation again, 1.4.4.2 is probably slightly less complicated and doesn't have to involve peeing on the spot. (easier yet more boring, depends on condition in the exhibition), 1.4.4.3 is easy to and nicer because of the installation being a living organism, needs a bit longer to actually see something happen (depends on time). Also possible is a combination of 1.4.4.1 and 1.4.4.3, where the plants in a room have influence on your body: with the oxygen the produce or maybe if you eat them ...
(I'm really not used to this WIKI thing yet but it's interesting!!!) So for now, I think we need to propose these different directions and see what the possibilities are: time, manpower, means ... After all, we need to build the sensors and gather some data first.
Project technical requirements
> Spectrophotometer Design
- Hardware-related:
- NX Arduino
- NX LEDs
- NX ...
- Software-related:
- Processing
- XML
> Open Spectra Data Set
- Google App framework
- Django / python / SQL
> Installation
- ...
Other project requirements, with estimated budget (optional)
- Urinoscope-related
- LED-based
- Arduino-based
- estimated budget per prototype: 50 euros
- 5 prototypes to be built: 250 euros (5x 50 euros)
- Collective Urinome OpenDataset
- Installation-related
- estimated budget: 300 euros
- Let's also define what kind of people we're looking for, roughly for every task
Project schedule
Over the 2 weeks that last the workshop, we plan to develop 3 distinct and interconnected parts to support this Urinomics project: Urinoscope, Collective Urinome OpenDatabase, Visualisation and installation.
Week 1 deliverables
- Urinoscope-related:
- Finalised Urinoscope hardware design + prototype testing/calibration.
- Finalised Arduino-software to enable the Urinoscope to be 'SpectroML' format compatible.
- Urinoscope documentation (an 'Instructables' for example)
- Collective Urinome OpenDataset-related:
- Implementation of a 'SpectroML' + Metadata compatible database
- Implementation of a data entry interface to submit data to Urinome OpenDataset.
- Urinome Dataset documentation on OpenWetWare
- Installation-related:
- Hardware:
- Construction of a water dripping system controlled by an Arduino
- 'SpectroML' file format to Processing software code
- Hardware:
Week 2 deliverables
- Urinoscope-related:
- Production and testing of 4 additional Urinoscope
- Data collection
- Collective Urinome OpenDataset-related:
- Implementation of a output interface to query/retreive data from the Urinome OpenDataset
- Urinome Dataset documentation on OpenWetWare
- Installation-related:
- Integration of the different Urinomics parts
- Fabrication of the water-dripping/plant watering system
Project documentation (optional)
- To be completed
Attachments (max 3 Files)
- To be completed
