Christiaen:Research
page in perpetual construction
background information
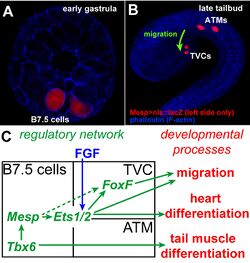
In ascidians, the so-called trunk ventral cells (TVCs) migrate from the anterior part of the tail to the ventral part of the trunk in tailbud embryos and constitute the precardiac mesoderm. They originate from a single pair of blastomeres in the early embryo, called B7.5 cells (Fig.1A). The B7.5 blastomeres give birth to the TVCs and their sister cells, the anterior tail muscles (ATMs), which differentiate into skeletal muscle and do not migrate (Fig.1B). Previous studies showed that TVC-specific gene expression and migration require transcriptional inputs from the bHLH transcription factor Mesp, the FGF signaling pathway and the forkhead transcription factor FoxF (Fig.1C). Because of these functional evidences, TVC migration constitutes a suitable model system to investigate the relationship between transcription regulation and directed cell migration.
To this aim, I developed a method utilizing fluorescence activated cell sorting (FACS) and microarray analysis to obtain TVC-specific genome transcription profiles. These data, together with an analysis of the function and regulation of the Rho GTPase RhoDF, indicated that 1) the gene regulatory network impinges on most cellular processes underlying cell migration, including actin dynamics, cell-matrix adhesion, polarity and vesicle trafficking; 2) for each cellular process, only a subset of the effector genes are subjected to transcription regulation and 3) specific cell behaviors result from the modular association of individual cellular processes that can be experimentally uncoupled from each others. This study led to the identification of additional candidate regulators and effectors of TVC migration and to the definition of a conceptual and experimental framework for future studies.
transcriptional control of TVC migration
A systems-level analysis is required to understand the structure and function of the interface between the precardiac GRN and TVC migration. Towards this goal, we are conducting functional analysis of candidate transcription factors and signaling molecules. A special attention is devoted to the forkhead transcription factor FoxF and its downstream candidate transcription regulators. These projects employ tissue-specific interference with transcription regulators function, cis-regulatory DNA analysis by electroporation of reporter constructs, knock-down by microinjection of morpholinos oligonucleotides and selected experimental manipulations will be subjected to subsequent analysis using FACS and whole genome microarrays.
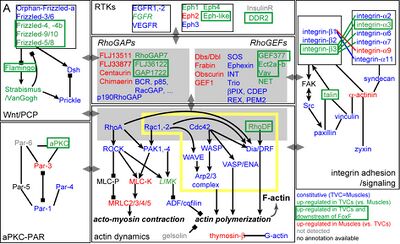
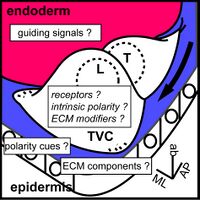
Regulation of heart vs atrial siphon muscle (ASM) precusors specification and collective cell migration
Once the TVCs stop migrating, they undergo stereotyped rounds of asymmetric cell divisions, which distinguish three types of muscle precursors: the first heart precursors first segregate from secondary TVCs, which divide again to produce second heart precursors and the atrial siphon muscles (ASM) precursors. The ASM precursors undergo a second collective migration towards the dorsal atrial siphon placode. Shortly after they are born, the ASM start expressing genes encoding the DNA binding transcription factors Collier/OLF/EBF (COE) and Islet. We found that COE function is necessary and sufficient to inhibit heart specification and impose the ASM fate, including migration towards the atrial placode. We use a combination of cis-regulatory analyses, candidate gene approach, cell sorting and whole genome microarrays to understand the mechanisms of ASM vs. heart fate specification and ASM migration. We recently identified a fundamental cross-regulatory antagonism between the NKX2-5 ortholog NK4 and the conserved cardiopharyngeal determinant Tbx1/10 that acts upstream of GATAa and COE to specify heart versus ASM fate in distinct second heart precursors and atrial siphon muscle precursors. These results were published in PLOS Biology.
We are currently dissecting signaling, regulatory and cellular mechanisms governing heart vs ASM fate specification upstream of COE and GATAa.
