CH391L/S12/Restriction enzymes and BioBricks assembly standards
Overview of restriction enzymes
What is a restriction enzyme?
A restriction enzyme, or restriction endonuclease, cleaves DNA at a restriction site.
Restriction enzymes are split into four types. Type I have multiple subunits and are not very useful because they randomly cleave the DNA distant from the recognition site, making the fragments produced unpredictable.

Type II are much more useful in the lab, because they cut DNA close to or within the recognition sequence (4-8 base pairs that exhibit inverted-repeat symmetry--see figure). In addition, type II endonucleases do not require energy from ATP hydrolysis[2].
An example type II restriction enzyme is EcoRI (E.C.3.1.21.4). The E.C. number specifies that this enzyme is a type II site-specific endodeoxyribonuclease producing 5'-phosphomonoesters.
For a step-by-step guide to cloning using restriction enzymes, see the Barrick Lab's wiki.
Digestion
After the digest reaction, the recognition site will be cleaved, with either blunt (no single-stranded DNA) or sticky (single-stranded overhang) ends. Isoschizomers recognize the same restriction site, but they can cleave in a different pattern—both blunt and sticky, with varying amounts of overhang[3]. The 5' phosphates remain attached, potentially allowing self-ligation, so a phosphatase step may be necessary following digestion.
Cleavage can be prevented by methylating the recognition sequence, on A or C nucleotides[3]. The host's DNA is methylated by methyltransferases, so that foreign DNA gets digested[2]. In this way, restriction enzymes can be thought of as a basic immune system.
Partial digestion occurs when the concentration of restriction enzymes is too low. This can be desirable, such as when there is more than one recognition sequence[3].
Nicking endonucleases
In contrast to restriction endonucleases, which cleave both strands of a DNA duplex, nicking endonucleases cleave only one strand, introducing a gap into the hydrolyzed strand. New England BioLabs supplies 10 different nicking enzymes and has information on 552 nicking enzymes in their REBASE database. They also provide the NEBcutter tool, which tests a sequence for endonucleases that only cut once.
More information on nicking endonucleases can be found in NEB's article.
Advancements in restriction enzymes
Type IIS endonucleases
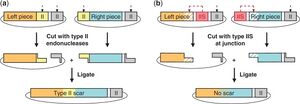
Type II restriction enzymes result in a scar when two fragments are ligated together. Type IIS endonucleases solve this problem by causing the sticky ends to be part of the gene itself, not the recognition sequence (see figure).
Zinc finger nucleases
Zinc finger (ZF) nucleases are artificial restriction enzymes.
The restriction enzyme FokI has two distinct domains: a DNA-binding domain, and a nuclease domain. The DNA-binding domain can be replaced with an engineered domain recognizing new sequences, allowing cleavage at a desired site. Additional site specificity comes from the fact that the ZF nuclease must dimerize in order to cleave the DNA, so two different binding sites must be specified that flank the site to be cleaved[5].
TAL nucleases
TAL (transcription activator-like) effectors allow targeted DNA binding with predictable specificity. By fusing a TAL effector with the nuclease domain of FokI, specific sequences can be cleaved[6].
Commercial sector
New England BioLabs
New England BioLabs (NEB) is a prominent supplier of reagents for the biological sector. It places most of its focus on restriction endonucleases, offering over 260 different varieties. They also provide free use of REBASE, a restriction enzyme and methyltransferase database, and Polbase, a DNA polymerase database.
BioBrick/BglBrick assembly standards
Purpose of RFCs
RFCs (requests for comments) have multiple purposes, including describing standards/protocols, how-to descriptions, and referencing previous RFCs to extend them or make them obsolete.
Assembly 10: the original BioBricks assembly standard
Assembly standard 10 was introduced by BBF RFC 10 in 2003.
Making BioBrick parts
BioBrick parts can be constructed by adding the prefix and suffix to a part with PCR.
The prefix is prepended to a primer sequence:
5' GTTTCTTC GAATTC GCGGCCGC T TCTAGA G 3'<EcoRI> <NotI> <XbaI>
Note that if the part codes for a protein, the prefix must instead be:
5' GTTTCTTC GAATTC GCGGCCGC T TCTAG 3'<EcoRI> <NotI> <XbaI>
The suffix is:
5' T ACTAGT A GCGGCCG CTGCAG 3'<SpeI> <NotI> <PstI>
So the other primer would include the reverse complement of this sequence:
5' GTTTCTTC CTGCAG CGGCCGC T ACTAGT A 3'<PstI> <NotI> <SpeI>
After PCR, these parts will have the standard triplet of restriction sites that make up the BioBricks prefixes and suffixes.
Putting two BioBrick parts together
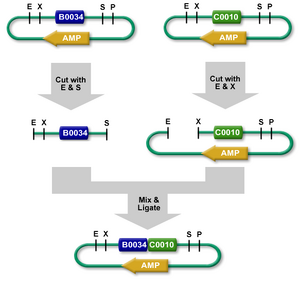
Ligating two BioBrick parts together results in a scar between the two parts: because the suffix restriction site does not match the prefix restriction site, upon ligation the palindrome is broken and will no longer comprise either recognition sequence. The scar codes for a tyrosine residue and a stop codon, plus two additional nucleotides (5' AG 3') that cause a frame shift. This means that fusing two proteins together cannot be done under Assembly 10, a significant disadvantage, driving the development of new RFCs that allow protein fusion. These standards include BBF RFC 23 ("Biofusion", which makes the scar into a 6-bp sequence that codes for tyrosine and arginine, with no extra nucleotides to cause a frame shift), BBF RFC 25 ("Fusion parts", which also eliminates the frame shift but changes the second residue from the problematic arginine to a glycine), and BBF RFC 21 ("Berkeley format" AKA BBb AKA BglBricks).
Assembly 21: BglBricks
Assembly 21 is described by BBF RFC 21. The main difference is its use of a different set of isocaudomers (restriction enzymes that recognize slightly different sequences but result in the same sticky ends after digestion), BamHI and BglII instead of XbaI and SpeI. As a result, the scar produced from fusion of two parts is not recognizable by BamHI/BglII because the palindromic sequence no longer exists, so more parts can be iteratively added. The parts can be added in any desired order by changing which restriction enzymes are used.
BglBricks also uses XhoI instead of PstI, and NotI is not included.
Advantages of BglBricks
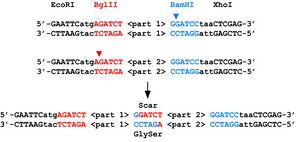
Parts adhering to RFC 21 can be fused together without a frame shift. In addition, the 6-bp scar between parts after fusion is a relatively safe glycine-serine sequence (in fact, this sequence is commonly used as a linker between fused proteins), and without the stop codon present in RFC 10. The restriction enzymes also have a higher efficiency than those used in RFC 10[8].
Disadvantages of BglBricks
BglBricks are incompatible with RFC 10.
References
- Kim YC, Grable JC, Love R, Greene PJ, and Rosenberg JM. Refinement of Eco RI endonuclease crystal structure: a revised protein chain tracing. Science. 1990 Sep 14;249(4974):1307-9. DOI:10.1126/science.2399465 |
Refinement of Eco RI endonuclease crystal structure: a revised protein chain tracing.
- Pingoud A and Jeltsch A. Structure and function of type II restriction endonucleases. Nucleic Acids Res. 2001 Sep 15;29(18):3705-27. DOI:10.1093/nar/29.18.3705 |
Structure and function of type II restriction endonucleases.
-
Bloch, Kenneth and Grossmann, Barbara. "Restriction Endonucleases." Current Protocols in Molecular Biology (1995) 3.1.1-31.21
- Lippow SM, Aha PM, Parker MH, Blake WJ, Baynes BM, and Lipovsek D. Creation of a type IIS restriction endonuclease with a long recognition sequence. Nucleic Acids Res. 2009 May;37(9):3061-73. DOI:10.1093/nar/gkp182 |
Creation of a type IIS restriction endonuclease with a long recognition sequence.
- Kim YG, Cha J, and Chandrasegaran S. Hybrid restriction enzymes: zinc finger fusions to Fok I cleavage domain. Proc Natl Acad Sci U S A. 1996 Feb 6;93(3):1156-60. DOI:10.1073/pnas.93.3.1156 |
Hybrid restriction enzymes: zinc finger fusions to FokI cleavage domain.
- Christian M, Cermak T, Doyle EL, Schmidt C, Zhang F, Hummel A, Bogdanove AJ, and Voytas DF. Targeting DNA double-strand breaks with TAL effector nucleases. Genetics. 2010 Oct;186(2):757-61. DOI:10.1534/genetics.110.120717 |
Targeting DNA double-strand breaks with TAL effector nucleases.
- Anderson JC, Dueber JE, Leguia M, Wu GC, Goler JA, Arkin AP, and Keasling JD. BglBricks: A flexible standard for biological part assembly. J Biol Eng. 2010 Jan 20;4(1):1. DOI:10.1186/1754-1611-4-1 |
BglBricks: A flexible standard for biological part assembly.