CH391L/S12/Pigments
Concepts and physical basis of pigment
Pigment is the substance showing the color. Specifically, "substance" means a certain kind/kinds of molecule(s), for pigment, it is usually the chemical bond of a conjugating system; "showing" means selectively absorbing certain wavelength out of the background light. And the color of the pigment we see is the net effect of all others remaining on the background. This is different and often compared to fluorescence or phosphorescence. Whereas pigment shows color by simple subtraction, these two adds new light for emission.
→"absorbing"
Electrons in molecules exist on certain energy levels. They try to exist at the lowest possible energy level as possible as they can. However, they can be boosted to higher level by "absorbing" the energy that fits the level difference. This is exactly what pigment does. They absorb the light that has right amount of energy that fits such energy level difference. The remaining nonabsorbed ones are perceived by us to "see" the color of pigment.
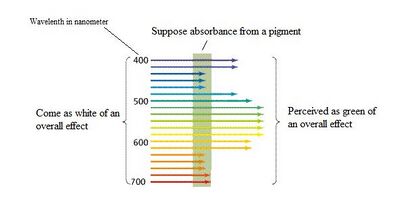
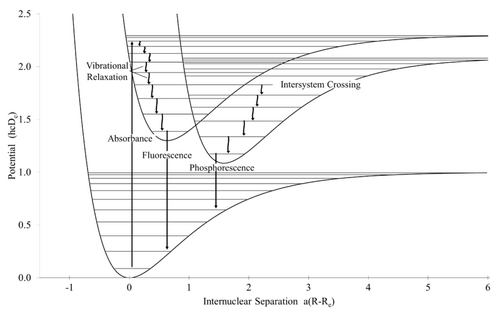
→"conjugating system"
Conjugating system generally means the delocalized electrons within overlapping p-orbitals. Since the electron orbitals are overlapping with each other, so single electron that presumably assigned to one atom can be shared to some extent by other atoms, which lowers the systemic/molecular energy. To make a figurative comparison (as how I understand it), suppose you have a balloon in hand. That is the molecule we have, and the air in it are the "electrons". And suppose you have a big balloon, that in another way means it gets high tension on its rubber surface---even a squeeze may cause it to burst---which means high "energy" of your balloon "molecule". However, if there is another space, or simply another small bag (overlapping p-orbital) that connects with this "molecule", you would see that the air (electrons) in this big balloon (molecule) is transferred to the small bag, and the big balloon now shrinks to a smaller size (lower energy). Since conjugating system usually possess lower energy, it is commonly seen as the "receiver" for the light energy, and thus performs as pigment.
→"fluorescence and phosphorescence"
Fluorescence can be generally described as an electron of a molecule relaxes to its ground state by emitting a photon of light after its excitation from a higher energy quantum state. The most commonly seen example of fluorescence occurs when ultraviolet region of energy is absorbed, which is invisible to human eye, and the emitted light is in the visible region, and shows the color (As can be seen of a TLC plate before staining under short UV). The process begins with excitation, relaxation (dropping to a relatively lower energy state), and shines fluorescence from that "relatively lower energy state". There are several kinds of relaxations: 1) non-radioactive: which means non-fluore/phosphorescnece, non-pigment, pure heat is released. 2) vibrational: returned to Maxwell–Boltzmann distribution. As in infrared/Raman process (relates to structure biology/chemistry). 3) Intersystem crossing: which means relaxation time is consumed upon "intersystem crossing", and emission of light is slow and delayed. Thus you see relatively long lasting phosphorescence. Accurately, it means total spin angular momentum of a molecule equals to one (triplet state), and thus it only takes a forbidden or non-preferred route to relax, since it is non-preferred, so there is the probability that one out of a million molecules would like to take this route, so that the overall "decaying" is very slowly. I personally see this from this example: if you get each of the four wheels of your car run (each with singlet state), it runs (fast relaxation: fluorescence); if your get a boot on three of them (triplet state, all to one), it still runs by only one wheel, but very very slowly (decaying: phosphorescence).
Biochromes and their uses
For a short history, pigments such as iron oxides have been used since prehistoric times. This is one of the naturally occurring pigment (Besides synthetic one after Industrial Revolution). The other one is the biological pigments or simply, biochromes. There are two kinds for this: plant pigment and animal pigment.
Plant pigment
Needless to say, pigment exists in plant for some reasons. Often, pigments help or implement certain tasks required for survival of plants. Rose is not red for heart recognises, but for attracting insects to flowers to encourage pollination. The most cited example for plant is green pigment chlorophyll used for photosynthesis:

→"Chlorophyll"
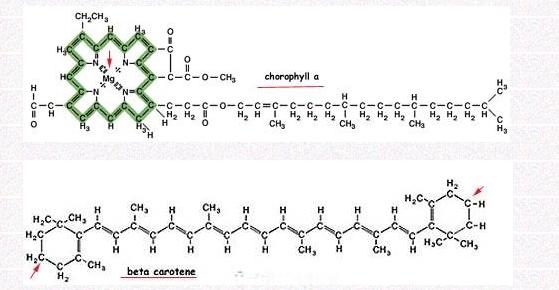
It is the primary pigment in plants; it is a chlorin that absorbs 430nm(blue) and 662nm(red)of light while reflecting green. It is the presence and abundance of chlorophyll that gives plants the green color. It contains a hydrophobic phytol chain that allow it to be embedded in a lipid membrane. And tetrapyrrolic ring rests outside of the membrane performs the light absorbance function. There are two forms of this pigment: chlorophyll a and chlorophyll b. "b" absorbs slightly different range of wavelengths and thus increase the light absorbance range of "a".
An interesting synthetic biological design on this is a so called de-greening reporting system[1]. The initial goal of this Colorado group was to develop a simple and inexpensive biological and chemical monitor that have following traits: 1) it can be used for large area detection. 2) signals are easily detectable. 3) remote monitoring. 4) re-set or repeatable capacity. After searching through candidates such as green fluorescent protein (GFP) and its variants, β-glucuronidase (GUS, GUS assay), luciferase or anthocyanin, they finally chose chlorophyll because of its fulfillment of all four traits.
The basic idea is assembling the chlorophyll production control parts under a steroid transcriptional inducible system. Their choice is called 10XN1P. It works in the presence of a synthetic steroid (4-hydroxytamoxifen, 4-OHT), with a transcriptional regulator relocating to the nucleus and inducing the expression of a promoter made up of specific response elements −46 region upstream of the cauliflower mosaic virus (CaMV) 35S promoter.
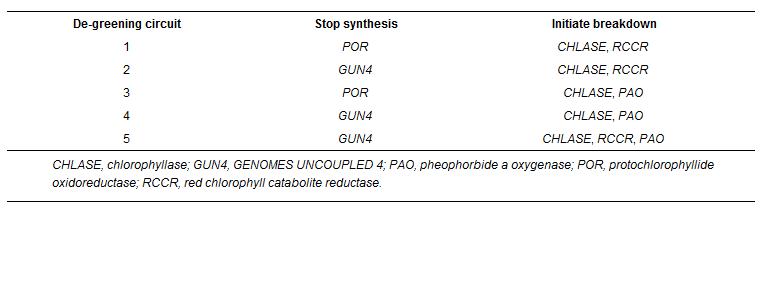
Based on previous knowledge and preliminary data, greening or chlorophyll production is to some extent auto-regulated by the plant itself. This means that mere cutoff biosynthesis or boosting degradation may not show desired phenotypical change, which was further confirmed in their study. To overcome this, they included in this reporting system two circuits: 1) stop synthesis 2)Initiate breakdown. In the stop part, they uses diRNA (double-stranded interfering RNA) construct to target several key enzymes in chlorophyll biosynthesis: 1) NADPH:protochlorophyllide oxidoreductase, which is an important rate-limiting enzyme. 2) GUN4 (GENOMES UNCOUPLED 4), encoded by GUN4, functions in positive regulating chlorophyll biosynthesis, precursor trafficking and activation of chelatase, which is important enzyme for making one chlorophyll precusor: protoporphyrin IX. In the initiate breakdown part, they aimed at three indispensable ones: 1) chlorophyllase: reponsible for hydrophobic tail removal. 2) pheophorbide a oxygenase: responsible for porphyrin ring cleavage (adding two oxygens). And 3) red chlorophyll catabolite reductase: also responsible for porphyrin ring cleavage (adding four hydrogens).
They made different combinations of the stop parts and initiate breakdown parts. After constructing on a T-DNA ( a plasmid transfers DNA fragment into the host plant's nuclear DNA genome), they tested these and selected the most prominent ones. Luckily they got the desired change, and furthermore, the change can be re-set through removal of the inducer. Remote imgaing measuring maximum quantum efficiency of photosystem II initially acknowledged its application significance.
→"Carotenoids"
Another important pigment in plant is called carotenoids, which is a category of tetraterpenoid (means 8 isoprene, hence 40 carbons in skeleton) organic pigments. They naturally occurs in the chloroplasts and chromoplasts of plants, but also are found in some other photosynthetic organisms like algae, some bacteria, and some types of fungus. The unoxygenated (oxygen free) carotenoids such as α-carotene, β-carotene and lycopene are known as carotenes, which are famous because of name-giving and bright orange color. Though they have been shown to act as antioxidants and to promote healthy eyesight in humans, their main function is to help fueling photosynthesis by collecting wavelengths of light not readily absorbed by chlorophyll.
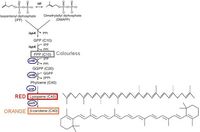
The Cambridge 2009 iGEM team aiming to "report a response using clear, user-friendly outputs" relates to this by designing "colour generator". The idea was simple: building biobricks that modulates enzymes involved in β-carotene biosynthesis pathway, tranformation into E.coli, and measure the yield by absorbance. The enzymes they focused on are Crt E, B, I, Y, which are phosphatase reponsible for PPi removal in the β-carotene biosynthesis pathway, the end-stage pathway following Non-mevalonate Pathway in E.coli producing the needed starting materials: isopentenyl pyrophosphate (IPP) and dimethylallyl pyrophosphate (DMAPP).
The biobricks they used were mostly from the previous iGEM teams, and they assembled them under a constituitive hybrid R0011 promoter and arabinose inducible I0500 pBad promoter so that they can measure the shift from the one of the starting material lycopene to β-carotene (Though they failed partially due to single wavelength measurement and hight cost of pure lycopene as a standard working curve). After fixing a stable strain problem (changing from TOP10 to MG1655), they succeeded in observing the desired result qualitatively and quantitatively.
→"anthocyanins"
Anthocyanins literally means blue flower, which is a water-soluble flavonoid (the old Vitamine P) pigments that appear red to blue, according to pH. They occur nearly in all parts of a plant but are most visible in the petals of flowers. Thus, it acquires great attention in cultivar field of making a blue flower, or more precisely, a blue rose.
A blue rose has long been synonymous with the unattainable. And it is often related to a never-heard (at least by me, because we traditionally do not use rose for love symbol) Chinese folktale. Because of this undesirability, Australian company - Florigene, and a Japanese company - Suntory, spent nearly 20 years creating a blue rose which was succeeded in 2004, and had displayed to public in 2008 World's first blue roses after 20 years of research.

Among the pigments influencing flower color, flavonoid/anthocyanin biosynthesis has been the most extensively studied[2]. Each plant species usually accumulates limited kinds of anthocyanins in its cell on certain parts and thus limited flower color. This is obtained through the expression of specific sets of biosynthetic genes, selections on key substrates from key enzymes in these pathways. Therefore, it is rare for a single species such as rose to have the entire spectrum of flower color, though it has been bred to yellow, white, pink and red. Besides rose, carnations and chrysanthemum, do not accumulate delphinidin-based anthocyanins either. And thus, all these important floricultural crops lack violet/blue varieties naturally (you can color a white rose blue though).
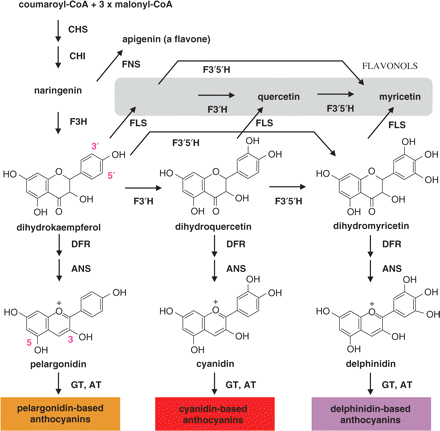
So the idea is simple: modulating the pathways of anthocyanin to change it to more "blue" anthocyanin containing direction. This is done through modulating the amount of key enzymes in the anthocynain biosynthesis pathway in rose. As can be seen, in this pathway, the "blue" anthocyanin is synthesized through delphinidin, which is a pivotal part rendering the blue color in the final anthocyanin molecule. Delphinidin itself is the product from dihydromyricetin, which is an important precursor for "blue" pathway. And it is converted largely by the help of flavonoid 3′,5′-hydoxylase (F3′5′H), which is absent in rose. So naturally, one way to build blue is to insert this F3′5′H from other species such as petunia into rose, so the dihydroflavonol 4-reductases (DFRs) of rose could enjoy more substrate. This is the basis for the early blue rose engineering.
However, to pursue a pure and exact blue color, pumping more blue making parts is not enough. As can be seen from the graph, there is also other contaminating pigments such as pelargonidin or cyanidin-based anthocyanins, which give orange and red colors. Thus these two needs to be eliminated. To achieve this, a Japanese group[3] added several modifications to the pathway: 1) constitutive overexpression of the viola F3′5′H BP40 gene and the torenia 5AT gene in rose by pSPB130 vector. Torenia 5AT catalyzes the transfer of the coumaroyl or caffeoyl moiety to 5-glucose of anthocyanidin 3,5-diglucosides, shifting its color more toward blue. This is based on the fact that the aromatic acylation of anthocyanin shifts its color toward blue by 3–4 nm[4]. 2) Overexpression of iris DFR genes by pSFL236 vector. As can be seen from the pathway, the substrate specificity of the DFR is a key in determining which anthocyanin is to be accumulated. The DFR of iris mainly accumulates delphinidin and lacks pelargonidin[5], is expected to have an efficient consumption on dihydromyricetin. 3) down-regulation of the endogenous rose DFR gene using RNA interference (RNAi). Though Rose DFR can utilize dihydromyricetin as a substrate on the basis of feeding of dihydromyricetin to rose petals[6], its crossing substrate affinity poses huge difficulty in getting to pure blue. Contaminating pelargonidin or cyanidin often overrides delphinidin production.
Non-plant pigment
→"Melanin"
Melanin is a dark biological pigment found in various parts on animals' skin, hair, feathers, scales, eyes, and even peritoneum. Specifically, it is a substance characterized as being brown, stiff, and granular in shape with individual granules having a diameter of less than 800 nanometers. This reveals its polymer nature. The most common form of biological melanin is eumelanin. It is formed by crosslinking of 5,6-dihydroxyindole (DHI) and 5,6-dihydroxyindole-2-carboxylic acid (DHICA). These two are generated in the tyrosine metabolism by two steps: Tyrosine → DOPA → dopaquinone. There is a plant melanin called catechol melanin.

Multiple functions for melanin in various organisms have been suggested: 1)it can be used as a defensive mechanism against predator, such as ink component in cephalopod. 2) protection against UV radiation, reactive oxygen species, heavy metals. 3) Used in innate immune defense system against invading pathogens by forming the bubble encapsulating the pathogen (melanization) and generating free radicals inside to kill[7]. 4) Radiotrophic fungi use melanin as a photosynthetic pigment to capture gamma rays and thus the energy for growth[8].
Melanin relates much to genetic disorders and diseases. One famous example is an autosomal recessive disorder called oculocutaneous albinism. And the first type of this relates directly to tyrosinase gene alteration, which encodes the enzyme tyrosinases in the two steps above producing the melanin monomer. Another example is Parkinson's disease. It is attributable to loss of neuromelanin in pigment-bearing neurons of four deep brain nuclei, especially substantia nigra and locus coeruleus. Neuromelanin is made from oxyradical metabolites of monoamine neurotransmitters including dopamine and norepinephrine, and is suggested of neurotoxicity prevention function by participation in autophagic process[9].
Cambridge 2009 iGEM team uses the MelA gene from Rhizobium etli , an aerobic, gram-negative, soil-living bacteria that is able to form symbiotic relationship with legumes. MelA encodes tyrosinases which involve cresolase and catecholase in the melanin production two steps mentioned above. The team made the simple biobrick with the native ribsome binding site before MelA and constructed on pTrcHis2B vector under the control of the IPTG-inducible promoter Ptrc after removal of the restriction site by fusion PCR. The expected results are obtained, however, not well characterized as using absorbance method in their paragon[10], they omitted library construction (Transposon mutagenesis for random knockout) and high-throughput screening as well.
→"Violacein"
Violacein is commonly regarded as natural antibiotic pigment generated by Chromobacterium violaceum, which is a a Gram-negative, facultative anaerobic, non-sporing coccobacillus. It has been suggested as of bactericide, trypanocide and tumoricide functions[11]. It has been also suggested used for treating tarballs pollution[12].
The biosynthetic pathways[13] for violacein include a decarboxylative fusion of two tryptophan units. As can be seen, it involves several steps catalyzed by five key enzymes: vioABCD. VioA generates an IPA imine from L-tryptophan and VioB converts the IPA imine into a dimer. VioE then transforms the dimer into protodexyviolaceinic acid (PVA), which can be converted into a green pigment called deoxychromoviridans. VioD and VioC hydroxylate PVA to form violacein that produces dark violet metallic sheen.
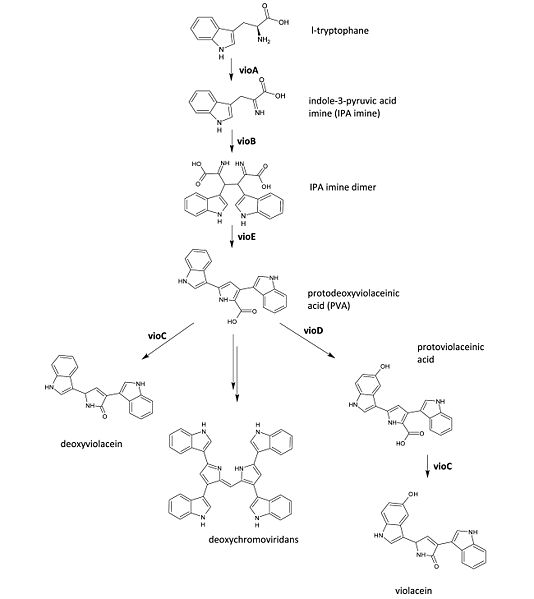
The Cambridge 2009 iGEM team made several operons of different combinations of vioABCDE enzymes from Chromobacterium voilaceum, since it has been suggested that removal of vioC or D can result in differnet color. They synthesized the entire main part of operon to remove the multiple restriction enzyme sites, and put it into pJexpress cloning cassette from DNA2.0. This cassette has promoter that is repressed by flanking copies of the lac operator. Symmetrical lac operators spaced around the promoter are to maximize cooperativity ensuring significantly tighter repression than regular lac operators[14]. The pigment results were confirmed by visual inspectiona and quantitative absorbance measurement.
Other pigment related iGEM parts
1. BBa_M11403 "Type 1 promoter of the cpcBA operon in Synechocystis sp. PCC 6803D: Phycocyanin A and B subunit promoter; cpcBA is a photosynthetic light-harvesting bile pigment-protein; absorbs orange/red light at ~620nm and emits ~650nm."
2. BBa_K322237 "In luminescent reactions, light is produced by the oxidation of a luciferin: luciferin + O2 → oxyluciferin + light. This biobrick was made from BBa_I712019. 6 bases upstream of the start codon, present in BBa_I712019 (presumably to make a proper eukaryotic context for the start codon), were removed for improved expression in E. coli. Note that the luciferase is temperature sensitive. Growing the host at 30°C will result in considerably increased brightness."
3.BBa_K317000, BBa_K317001 "Halobacteria exhibit phototaxis responses to changes in light intensity and color using the seven-transmembrane retinylidene photoreceptors sensory rhodopsins II (SRII). Light-activated SRII transmits signals to their cognate transducer, HtrII, respectively. The Htr protein contains two transmembrane helices and cytoplasmic methyl-accepting and His-kinase-activating domains homologous to those of chemotaxis transducers of eubacteria, such as the Escherichia coli and Salmonella enterica serovar Typhimurium Tar, chemotaxis transducers for serine and aspartate, respectively. Sensory rhodopsin II from Natronobacterium pharaonis (NpSRII) is very similar in spectroscopic and functional properties to the repellent receptor SRII in Halobacterium salinarum. And it has been found to be more stable in response to variation in external conditions such as pH and ionic strength. The NpSRII protein mediates a repellent response to blue-green light (maximum λ,497 nm) when it is coexpressed with its transducer, NpHtrII, in H. salinarum. Also, when expressed in E. coli, NpSRII is capable of binding all-trans retinal to form a blue-green-lightabsorbing pigment. "
4. BBa_K678035 "The gene yA from Aspergillus nidulans encodes a p-diphenol oxidase (laccase) that converts a yellow pigment precursor to the mature green form. The A. nidulans conidia (asexual spores) contain a dark green pigment in their walls. The pigment protects the spores from damage by ultraviolet irradiation. The synthesis of this pigment involves the expression of several genes (ωA, yA andfiA) and other genes can modify the amount- and chemical properties of the pigment (dilA, chaA, drkA). A yA mutant produces yellow spores."
Colorful bacteria
Pigmented bacteria or so called chromobacteria remains a huge source for the future development of bio-synthetic pigments. Their chemical components range from pyrrole, phenazine, carotenoid to xanthophylls and quinone derivatives. This variety provides with great variety in color: almost all colors in spectrum. Because molecular oxygen is essential for pigmentation, so anaerobic bacteria are usually nonpigmented. Pigment synthesis is also greatly affect by light, pH, medium conditions. Some of the representative ones of the pigmented bacteria and applications of these pigments:
References
<biblio>
- degreen pmid=17309732
//De-greening reporting system assembling key components in chlorophyll biosynthesis and degradation pathways
- mostextensive Tanaka Y, Brugliera F. Flower colour. In: Ainsworth C, editor. Flowering and its Manipulation. Oxford: Blackwell; 2006. p. 201-239.
//Flavonoid pathways
- bluerose pmid=17925311
//Blue making parts insertion together with RNAi knockdown endogenous DFR
- blue34 pmid=9881162
//Aromatic acylation of anthocyanin shifts its color toward blue
- lackpelar pmid=10417729
//Iris DFR basis
- roseDFR Holton TA, Tanaka Y. Blue roses—a pigment of our imagination? Trends Biotechnol. 1994;12:40-42.
//rose DFR
- melaninimmune pmid=15199959
//Melanin encapsulation of foreign invading pathogen
- melanintrophic pmid=17520016
//Melanin to capture light for fungi growth
- neuromelanin pmid=18384642
//Neuromelanin involved in autophagic process
- melaparagon pmid=18156325
//Melanin-Based High-Throughput Screen for l-Tyrosine Production in E.coli
- velafunc World Journal of Microbiology and Biotechnology Volume 14, Number 5, 685-688, DOI: 10.1023/A:1008809504504
//Production, extraction and purificationof violacein
- tarball Itah A. Y., Essien J. P. (2005). "Growth Profile and Hydrocarbonoclastic Potential of Microorganisms Isolated from Tarballs in the Bight of Bonny, Nigeria". World Journal of Microbiology and Biotechnology 21 (6–7): 1317–1322. doi:10.1007/s11274-004-6694-z.
//Chromobacterium violaceum in tarball treatment
- violabiosynthesis pmid=16874749
//Violacein biosynthetic pathway
- T5promoter pmid=8045263
//Quality and position of the three lac operators of E. coli define efficiency of repression

