Biomod/2013/Aarhus/Results And Discussion/Origami
<html> <style> /* ul.menu li.</html>Results_And_Discussion/Origami<html> a {
color: cyan;
}
- /
- toc {
display: none; }
- mytoc {
background: none; width: 200px; }
.toc { border: 0px solid; }
- toc ul ul,.toc ul ul {
margin: 0 0 0 1em; }
table.toc { background-color: #f0f4f4; }
- mytoc a,#mytoc a:visited {
font-size: normal; color: #222222; }
- mytoc a:hover {
font-color: #009ee0; /* text-decoration: underline; */ }
- wiki-toc {
width: 200px; margin-top: 6px; float: left; }
- wiki-body {
margin-left: 200px; padding-left: 12px; padding-right: 35px; }
- toc #toctitle,.toc #toctitle,#toc .toctitle,.toc .toctitle {
text-align: left; }
- toc h2,.toc h2 {
font-weight: normal; font-size: 17px; }
/*
- wiki-contents A {
color: #00aeef; }
- wiki-contents A:HOVER {
color: #00aeef; }
- /
- toctitle span {
display: none; }
/*
- wiki-body p,#wiki-body li,#wiki-body dd,div.thumbcaption {
font-size: medium; }
- /
/* required to avoid jumping */
- tocScrolWrapper {
/* left: 450px; */ position: absolute; /* margin-left: 35px; width: 280px; */ }
- tocScrol {
position: absolute; top: 0; /* just used to show how to include the margin in the effect */ /*margin-top: 20px; */ /* border-top: 1px solid purple; */ /*padding-top: 19px;*/ }
- tocScrol.fixed {
position: fixed; top: 0; }
- editPageTxt {
text-align: left; padding-left: 15px; }
- editPageTxt P {
clear: both; }
- toPageTop {
float: left; position: relative; top: 18px; left: 13px; color: #d13f31; } </style>
<script type="text/javascript"> $(document).ready(function() { var parentTable = $("#toc").parent(); $('#mytoc').append($("#toc").first());
$('#mytoc').find("#toc").attr("id", ""); parentTable.closest('table').remove(); });
$(document).ready( function() { var top = $('#tocScrol').offset().top - parseFloat($('#tocScrol').css('marginTop').replace( /auto/, 0)); var nav = $('#tocScrol'); var max = $('#indexing').offset().top - nav.height();
$(window).scroll(function(event) { // what the y position of the scroll is var y = $(this).scrollTop();
if (y > top) { // && signs are html decoded thus this construction if (y >= max) { nav.removeClass('fixed'); nav.css({ position : 'absolute', top : max - top }); } else { nav.addClass('fixed'); nav.removeAttr('style'); } } else { nav.removeClass('fixed'); nav.removeAttr('style'); } }); }); </script> <html> <html>
<a id="toPageTop" href="#">▲</a>
[<a href="http://openwetware.org/index.php?title=Biomod/2013/Aarhus/</html>Results_And_Discussion/Origami<html>&action=edit">edit this page</a>]
</html>
Introduction to DNA origami
The design of the system was made using the DNA origami principle first described by Rothemund in 2006. [1] A DNA origami can be made using the M13mp18 single stranded genome (7249 bases long) assembled with a number of shorter DNA strands, called staple strands, in a self-assembly reaction.
The strategy was to design and create two DNA origami structures, a plate structure and a dome structure. Subsequently, the plate structure was loaded with different modified oligonucleotides at the 3' end of selected staple strands and covered by the dome structure with a DNA-modified peptide lock system.
Origami plate
The design of the origami plate was inspired by previous work with multilayered DNA origamis. The plate was designed as a double layered sheet to increase its stability and decrease its permeability. The goal was to design the plate as flat as possible to ensure that it could attach to the cell surface properly through cholesterol modifications [2, 3]. This would bring the attached photosensitizers close enough to the cell to be able to induce apoptosis. [4]
Several 3' ends of the staple strands on one side of the plate were modified with different functional groups (cholesterols, photosensitizers, peptide lock and sisiRNA).
Two different versions of the system were made. In the first version the plate was modified with photosensitizers that produce cytotoxic singlet oxygen once radiated with 635 nm light and with cholesterol modified staple strands. [5]
The second version features the plate modified with sisiRNAs [6] that were conjugated to cell penetrating peptides, which induces specific gene knockdown through the RNAi pathway. These were attached to the plate with the peptide lock that was designed to be cleaved by matrix metalloprotease 2, which is overexpressed by some metastasizing cancer cells.
Origami dome
The dome origami was designed as a lid for the plate origami. It was based on the hemisphere origami design created by Han, D.R et al. [7]
This structure allows for a cavity inside the whole origami structure which would be large enough to contain the functional modifications (cholesterol, photosensitizer and sisiRNA).
The dome was assembled in a self-assembly reaction. Afterwards, an attempt was made to connect the dome and plate.
Design
Design of the origami plate
The double layered plate design was chosen to increase the compactness of the structure and to induce further stability. Additionally, the correct shape of the structure could easily be verified by atomic force microscopy (AFM) or transmission electron microscopy (TEM). The design pattern was created in caDNAno [8][1] and subsequently sent to CanDo [2] for determination of the flexibility and bending of the 3D structures in solution (see Figure 3). [9]
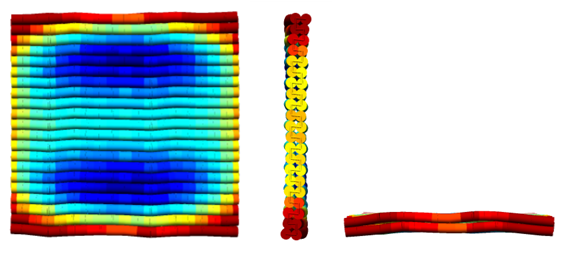
The dimensions of the plate DNA origami were limited by the length of M13mp18, and were 21 helices wide, 2 helices high and 143 base pairs long (i.e. approximately 53 nm x 5 nm x 46 nm).
The staple strands connecting the edges were omitted in the origami structure to prevent oligomerization, in which the structures align end to end and form long aggregates.[10] This leaves the scaffold strand with a single stranded region of between 16-26 nucleotides at both ends of each helix.
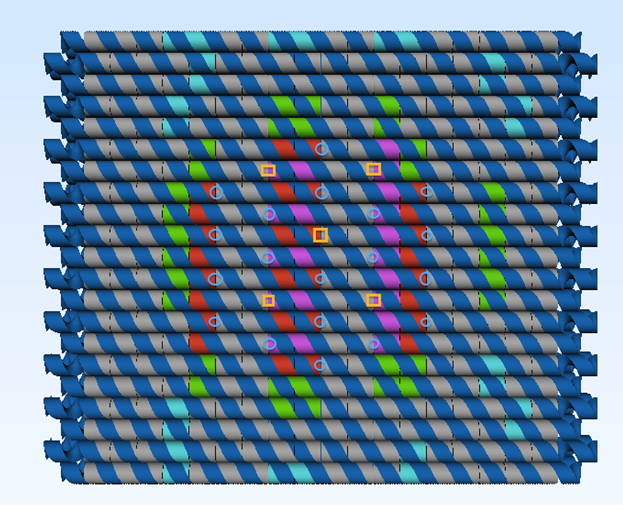
The plate was designed with the approximate crossover pattern of 32 base pairs between adjacent helices because the multilayered origamis require frequent crossovers to hold the structures tightly together (see Figure 4). The annealing domains of the staple strands are made in 8-8-8-8 motif. The strain of the structure is balanced by the deletion of one base pair per every fourth vertical crossover. An example of a such deletion is after the crossover between helix 2 and 5, where a deletion was made and then again at the third cross over between helix 2 and 5 (see Figure 4).
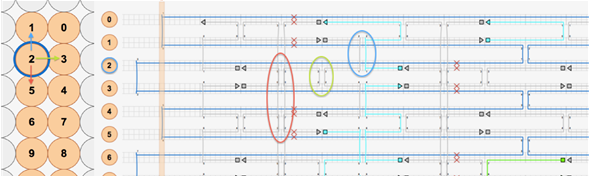
The same pattern was created between the rest of the helices in the structure to make the plate as flat as possible. The staple strands that fold the circular M13mp18 into the desired structure were divided into modules, depending on their function. These modules are shown in table S1 (supplementary data) and in Figure 5.
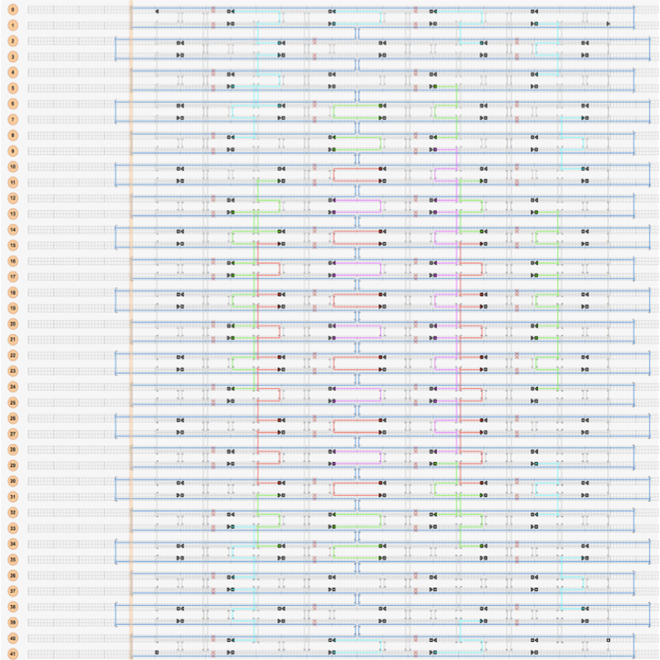


Design of the origami dome
The origami dome, designed as a lid for the plate origami, was based on the hemisphere origami design created by Dongran Han.[7] The design of the origami dome was chosen to create a hollow structure to enclose and protect the functional modifications (cholesterol, photosensitizer and sisiRNAs).
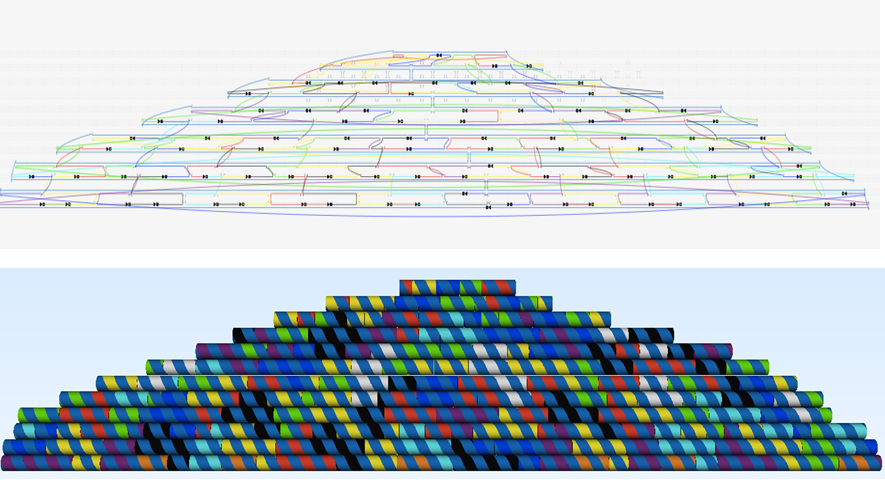
The design was made in caDNAno (see Figure 7) and the structural details of the hemisphere is consistent with table S2 in supplementary data.
The staple strands were divided into colored modules (see table S3 - supplementary data). The staple strands in the white colored module can be replaced by the peptide lock-connecting strands of the D-zip module (see Connection of the plate and the dome). The Plate XO staple module can be replaced by the Dome-Plate-Connector module (see Connection of the plate and the dome).
Design of the connection between the plate and the dome
The lock mechanism of the system, utilized different staple strand overhangs from the two structures, designed to anneal to the modified peptide-DNA lock construct (peptide lock). The 3’ overhang from the dome origami anneal in a zipper-like helix with the 5’ end of the peptide lock, while the 3’ overhang of the plate origami anneals forming an overlapping helix structure with the 3’end as toehold (see Figure 8). This lock design was chosen because it creates stability of the structure as well as easier accessibility for the enzyme to cleave at the overlapping helix region. Furthermore the zipper-like helix creates a degree of flexibility so that the lock is able to hold the structures tight, although not so tight that it will cause unwinding of the structure. The strand displacement technique was intended to test the lock and key mechanism of the system, using the 3’ toehold of the staple strand from the plate (green) to remove the WB-peptide-lock (black) holding the structures together (see Figure 8B).
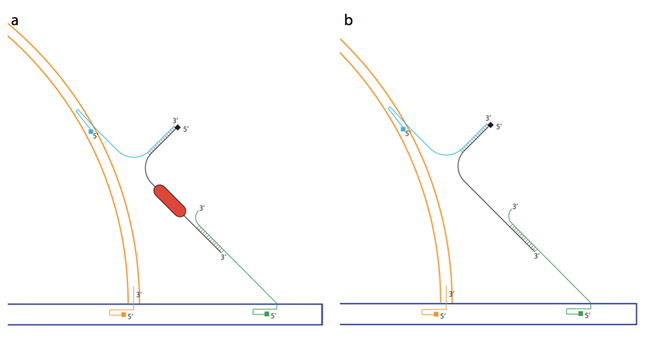
Furthermore, an aid mechanism was made in order to create a higher yield of assembly of the two structures. Eight strands from the dome connecter-module of the plate were designed to create a direct crossover between the structures. These strands anneal with the 5’ end in the plate origami and cross over into the outer ring of the dome origami, annealing at the plate-XO strands sites. These are marked in orange in Figure 8.
These staple strands can be removed again using toeholds applied to the 3’ end of the strand, followed by strand displacement. This mechanism ensures that the final system is able to separate when the peptide locks are cleaved.
Results and discussion
Origami plate
To investigate the optimal self-assembly conditions for the plate origami, reactions were performed with different ratios of staple strands to M13mp18, time ramps and self-assembly buffers. The self-assembly reactions were performed with buffers containing MgCl2 in concentrations of either 5 mM, 8 mM, 12.5 mM, 18 mM or 20 mM or MgAc in concentrations of 12.5 mM or 18 mM.
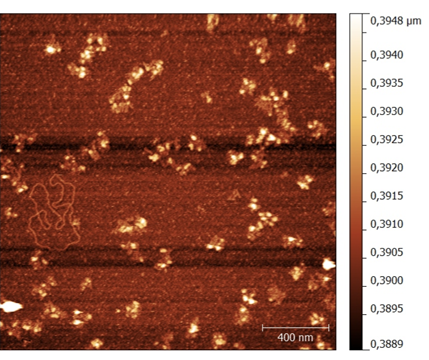
The agarose gel in Figure 9 showed clear band shifts in the lanes containing the assembled structures over both 1, 3 and 5 hour linear temperature ramps. In these reactions, a 1:10 ratio between M13mp18 and staple strands was used (see Figure 9). However, further characterization using AFM showed incorrect folding of the structure. This led to the conclusion that the structure did not assemble at a 3-hour linear ramp (see Figure 10).
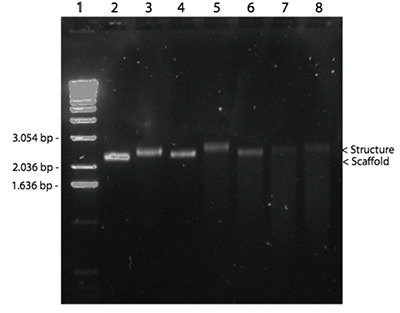

<html>
</html>
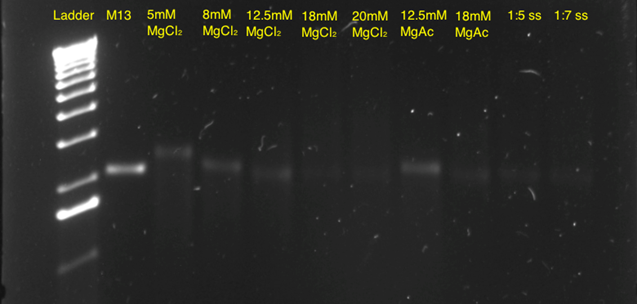

Subsequently, a 10-hour linear ramp was tested (Figure 11). In both the 5mM, 8mM and 12.5 mM MgCl2 lanes, a band was seen indicating correct self-assembly. The faint bands in the lane with 18 mM and 20 mM MgCl2 indicated that these concentrations were too high, thereby leading to aggregation of the structures. In addition, a band in the 12.5 mM MgAc lane is seen, indicating that this buffer could also be used, however working commenced with the 12.5 mM MgCl2 concentration, as this appeared to be optimal.
Several annealing ramps were tested, both non-linear and linear ramps, ranging from 1 hours to 72 hours. A longer annealing time could ensure that the DNA strands further approaches the free energy minimum, which should make the structure assemble correctly. A longer annealing time might result in aggregations due to kinetic traps, when longer strands anneal in unwanted ways. However, this is not proven and need further testing.
The reactions that showed the clearest and most intense band shifts were the 5 hours and the 13 hours non-linear temperature ramps (see Figure 12).
The gel in Figure 12 showed signs of aggregations that might be caused by enhanced stacking between the structures. However, a difference in the amount of aggregation was observed when comparing the samples with higher Mg2+ concentration. Lane 6 appears to contain more and larger aggregates than lane 5 that contain a lower Mg2+ concentration (Figure 12).
The samples in Figure 12 all seemed to be assembled, as indicated by the band shift.
AFM images of the 18 mM Mg sample that was annealed over 5 hours still showed incorrect assembly (images not shown).
Additional
AFM imaging was made on samples assembled in the 12.5, 18 and 20 mM Mg2+ with both the 17-hour non-linear ramp and the 13-hour non-linear ramp (Figure 13 shows the 12.5 mM Mg sample at a 17-hour ramp).
Based on these images, all future work was done on a 12.5 mM MgCl2 and a 17-hours non-linear time ramp at a 21 nM origami concentration, showing the best yield of non-aggregated correctly assembled plate origami. In addition to AFM images, TEM images of the sample with 12.5 mM MgCl2 and 17 hour ramp were obtained, and those images confirmed that the plate origami assembled correctly using the described conditions (Figure 14).
The size of the plate appears to be approximately 45 nm x 45 nm with only little aggregation observed using 12.5 mM MgCl2.
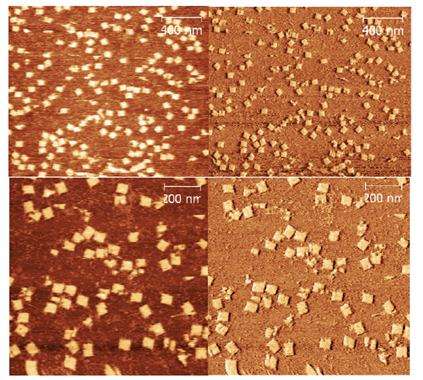
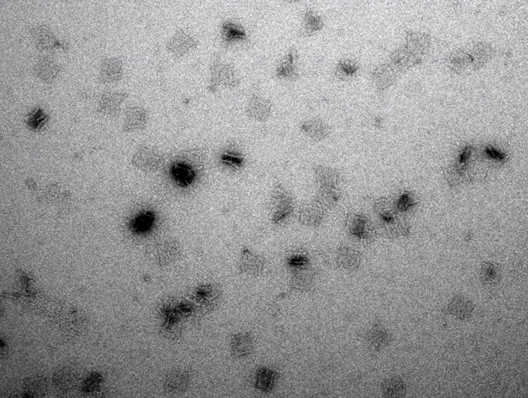
Origami dome
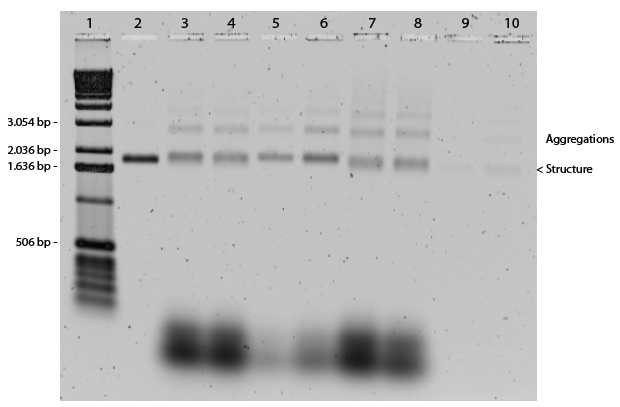
To investigate the optimal self-assembly conditions for the dome origami, reactions were performed with different self-assembly buffers. The assembled dome was expected to be localized slightly above the unfolded M13mp18 control band in a native 1 % agarose gel. The buffers containing different Mg2+ concentrations (see materialts and methods) were tested as well as the required amount of staple strands to fold the M13mp18 correctly. The different reaction mixtures were annealed over a non-linear annealing ramp constituted by rapid heating to 95°C and subsequent slow cooling.
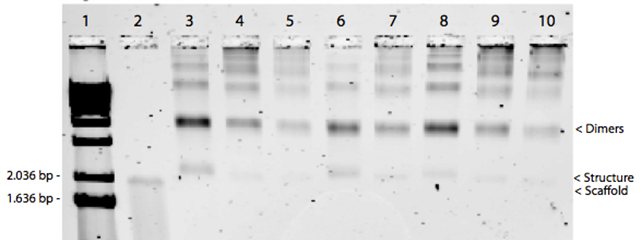
The gel in Figure 15 showed distinct band shifts between the single stranded scaffold in lane 2 and the bands in lanes 3-10. These shifts indicate structural changes of the scaffolds in the samples, but also show that the structures had aggregated. From these results it was not possible to conclude if the folded structures had assembled correctly. The structures in lane 3-5 and 8-10 were assembled with a 10x staple strand excess compared to M13mp18, while the structures in lane 6 and 7 were assembled with a 20x excess of staple strands. There was no difference between the positions and intensities of these bands, indicating that a 10x excess of staple strands would be sufficient for self-assembly to take place. The increased Mg2+ concentration in the different reactions enables the helices to pack more closely, thereby creating a tighter structure that appears smaller in the gel. Thus, when the Mg2+ concentration is increased in the self-assembly reaction, the structures migrate further in the gel. The difference when using the 13-hour non-linear ramp and 17-hour non-linear ramp was very small, and other characterization methods were therefore needed to determine if the 13-hour ramp sufficed, or if the 17- hour ramp were necessary. Furthermore, the size of the structure was hard to determine with certainty due to the large unpaired region of the scaffold strands and the asymmetrical shape of a hemisphere.
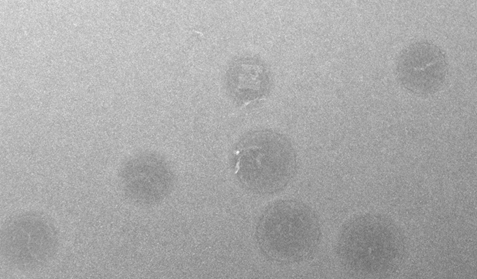
A reaction with 12.5 mM Mg2+ buffer and a 10x excess of staple strands, annealed over an even shorter non-linear temperature ramp of 11 hours was imaged using AFM ( see Figure 16). These images confirmed that the dome was able to fold into the desired structure under the tested conditions.
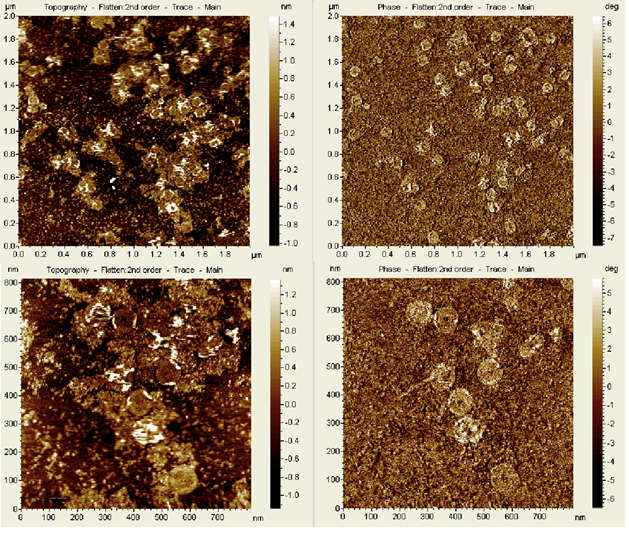

A dome self-assembly reaction with 12.5 mM MgCl2 annealed over a 17 hour non-linear time ramp imaged with TEM concur with the AFM images revealing the assembled dome structure (see Figure 17).
In the gel in Figure 15, aggregations were evident. To avoid this, adding formamide to the self-assembly buffer was tested.[11]
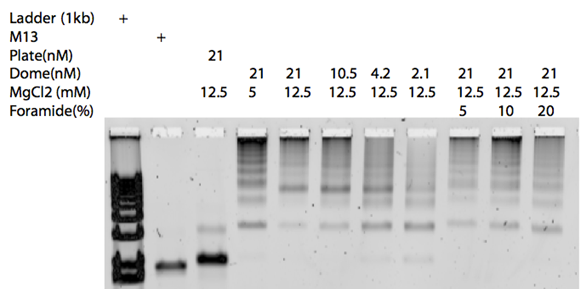
However, the gel did not show minimized aggregation upon addition of formamide to the buffer and this approach was abolished.
Connecting the plate and dome origamis
After succesfully assembling the plate and the dome separately, experiments were performed to connect the two structures. The plate and dome were originally intended to be connected through the peptide lock, but this was not finished at the time of this experiment. Instead it was attempted to show that these structures could be connected using staple strands that mimicked the peptide lock (WB-peptide lock). Before connecting the dome and plate, both origamis had to be purified to avoid unwanted aggregates using a 100kDa Amicon Spin Filter. The gel in Figure 19 indicates that the purification using spin filtration was successful in removing the excess staple strands from the plate. However, a small degree of aggregation was observed, due to sample upconcentration occurring during spin filtration.
Spin filtration proved to be less useful for purification of the dome origami, as the folded structure was unable to withstand the high speeds of centrifugation required in this method. Centrifugation at lower speeds was tested, but this approach was not successful in removing the excess staple strands.
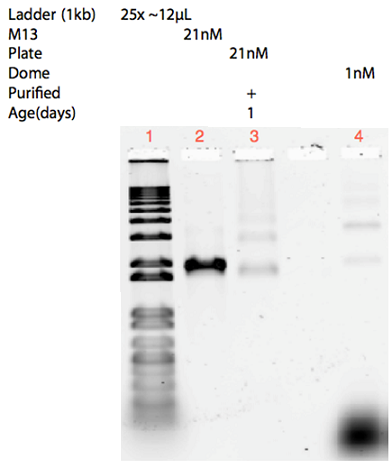
Instead, freeze n’ squeeze gel purification was employed. This method proved successful for purification of the dome, by removing both excess staple strands and aggregations visible in the unpurified sample in Figure 20.
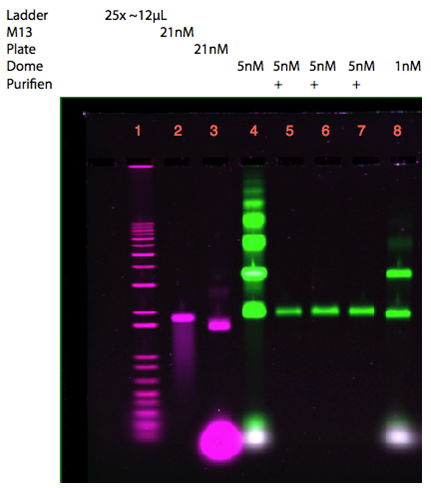
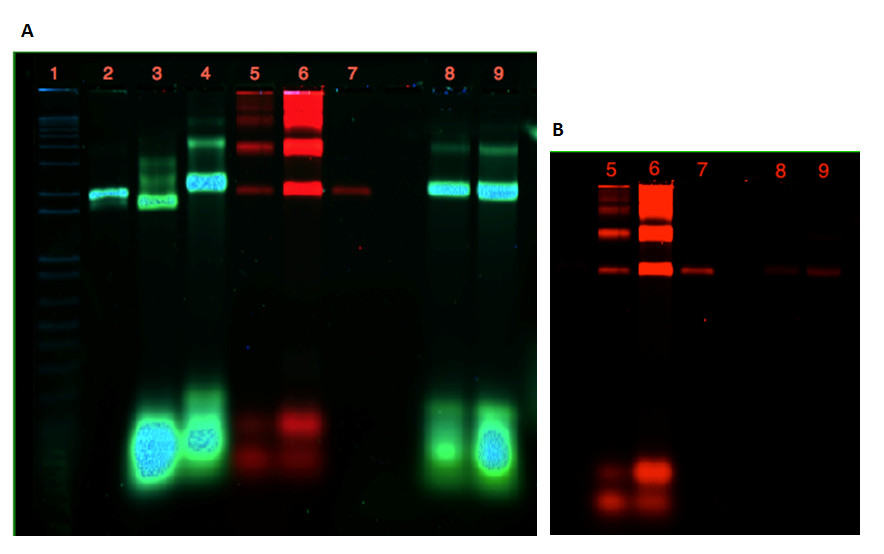
As seen from the gels in Figure 20 and 21, attachment of the dome and the plate was inconclusive. Lane 3 contains 21 nM unpurified folded plate, but instead of shifting upwards in the gel, as anticipated from previous experiments (see Figure 9, 11, 12), the band appears lower than the unfolded M13mp18 control in lane 2. In addition, the purification of the dome origami gave a low yield, even after an attempted upconcentration of the sample on a spin filter.
If a connection of the two origamis had been achieved, a merge of the two colors from the Cy-dyes would have been seen in lane 8 and 9. This is not apparent from the gel in Figure 21A, but Figure 21B show that this might be due to the low amount of dome origami seen in lane 8 and 9.
Due to time constraints it was not possible to perform further experiments to connect the two origamis.
Conclusion
Designing two complex 3D DNA origami structures was achieved using the program caDNAno in combination with the plug-in feature of Autodesk Maya to visualize the 3D structures of the origamis. CanDo was employed to provide a computational prediction of the structures, regarding additional information about the shape and flexibility in solution.
Succesful self-assembly was achieved form for both of the two origami structures. Through optimizations of the magnesium concentration in the self-assembly buffer and as well as changing the time and temperature of the annealing ramp, the structures were successfully assembled.
Characterization of the structures was achieved by both gel electrophoresis, AFM and TEM, confirming the formation of the correct structures.
Furthermore, attempts were made to connect the two structures. These results were inconclusive and further experiments will need further optimizations.
References
-
Rothemund, P. W. K. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297-302 (2006)[1]
-
Teruya, K. et al. Semisynthesis of a protein with cholesterol at the C-terminal, targeted to the cell membrane of live cells. K. Protein J. 29, 493–500 (2010). [1]
-
Langecker, M. et al. Synthetic lipid membrane channels formed by designed DNA nanostructures. Science 338, 932–936 (2012)[1] LIGGER DOBBELT
-
Pedersen, B. W. et al. Single Cell Responses to Spatially-Controlled Photosensitized Production of Extracellular Singlet Oxygen. Photochem. Photobiol. 87, 1077-1091 (2011) [1]
-
Arian, D. et al. A nucleic acid dependent chemical photocatalysis in live human cells. Chem. Eur. J. 16, 288–295 (2010).[1] LIGGER DOBBELT
-
Bramsen, J. B. et al. Improved silencing properties using small internally segmented interfering RNAs. Nucleic Acids Res. 35, 5886-97 (2007). [1].
-
Han, D.R. et al. DNA Origami with Complex Curvatures in Three-Dimensional Space. Science 332, 342-346 (2011) [1]
-
Douglas, S.M. et al. Rapid prototyping of 3D DNA-origami shapes with caDNAno. Nucleic Acids Res. 37, 5001-5006 (2009) [1]
-
Kim, D. N. et al. Quantitative prediction of 3D solution shape and flexibility of nucleic acid nanostructures. Nucleic Acids Res. 40, 2862-2868 (2012). [1]
-
Kim, K. N. et al. Comparison of methods for orienting and aligning DNA origami. Soft Matter 7, 4636-4643 (2011). [1]
-
Zhang, Z. et al. Self-assembly of DNA origami and single-stranded tile structures at room temperature. Angew. Chem. Int. Ed. Engl. 52, 9219-23 (2013) [1]
-
Y. Ke et al. Multilayer DNA Origami Packed on a Square Lattice. J. Am. Chem. Soc. 131, 15903-15908 (2009). [1]
-
Patel, L. N. et al. Cell penetrating peptides: Intracellular pathways and pharmaceutical perspectives. Pharm. Res. 24, 1977-92 (2007). [1]
-
O. Mendes et al. MMP2 role in breast cancer brain metastasis development and its regulation by timp2 and erk1/2. Clin. Exp. Metastasis, 24, 341-351 (2007). [1]
</html>
<html> <head> <style>
- indexing {
/* float: left; position: center; */ background-color: #222; border-top: 2px solid #d13f31; color: #006e9c; margin: 0px; padding: 0px 0px 10px 0px; width: 100%; text-align: center; }
.footer-section { padding: 10px; display: table-cell; text-align: left; }
.footer-section-title { font-size: 20px; }
- footer-contents {
color: #006e9c; display: inline-table; }
.footer-section A { color: #006e9c; text-decoration: none; }
.footer-section A:HOVER { color: #00aeef; }
.footer-section ul { list-style-type: square; }
- sitemapTitle {
margin-top: 20px; font-size: 24px; }
- editFooter {
float: right; margin-top: -28px; margin-right: 5px; }
- editFooter A {
color: #006e9c; text-decoration: none; }
.cf:before,.cf:after { content: " "; /* 1 */ display: table; /* 2 */ }
.cf:after { clear: both; }
- bodyContent a[href^="mailto:"], .link-mailto {
background: url() no-repeat scroll right center transparent; padding-right: 0px; color: #006e9c;
}
</style> </head> <body>
SITEMAP | BIOMOD 2013 NANO CREATORS | Aarhus University
Copyright (C) 2013 | BIOMOD Team Nano Creators @ Aarhus University | Programming by: <a href="mailto:pvskaarup@gmail.com?Subject=BIOMOD 2013:">Peter Vium Skaarup</a>.
<img alt="Sigma - Aldrich" src="http://openwetware.org/images/3/39/Sigmaaldrich-logo%28transparant%29.png" width="300px" height="154px"> <img alt="VWR International" src="http://openwetware.org/images/2/28/Vwr_logo.png" width="300px" height="61px"> <img alt="Promega" src="http://openwetware.org/images/7/72/Promega.png" width="175px" height="105px" style="padding-right: 5px; padding-left: 5px;"> <img alt="kem-en-tec" src="http://openwetware.org/images/3/3a/Kementec.png" width="130px" height="129px"> <img alt="Centre For Dna Nanotechnology" src="http://openwetware.org/images/4/4f/CDNA_logo.png" width="420px" height="90px"> <img alt="Dansk Tennis Fond" src="http://openwetware.org/images/9/9a/Dansk_tennis.png" width="250px" height="53px">
</body> </html>