Biomod/2013/Aarhus/Materials And Methods/Methods
<html> <style> /* ul.menu li.</html>Materials And Methods/Methods<html> a {
color: cyan;
}
- /
- toc {
display: none; }
- mytoc {
background: none; width: 200px; }
.toc { border: 0px solid; }
- toc ul ul,.toc ul ul {
margin: 0 0 0 1em; }
table.toc { background-color: #f0f4f4; }
- mytoc a,#mytoc a:visited {
font-size: normal; color: #222222; }
- mytoc a:hover {
font-color: #009ee0; /* text-decoration: underline; */ }
- wiki-toc {
width: 200px; margin-top: 6px; float: left; }
- wiki-body {
margin-left: 200px; padding-left: 12px; padding-right: 35px; }
- toc #toctitle,.toc #toctitle,#toc .toctitle,.toc .toctitle {
text-align: left; }
- toc h2,.toc h2 {
font-weight: normal; font-size: 17px; }
/*
- wiki-contents A {
color: #00aeef; }
- wiki-contents A:HOVER {
color: #00aeef; }
- /
- toctitle span {
display: none; }
/*
- wiki-body p,#wiki-body li,#wiki-body dd,div.thumbcaption {
font-size: medium; }
- /
/* required to avoid jumping */
- tocScrolWrapper {
/* left: 450px; */ position: absolute; /* margin-left: 35px; width: 280px; */ }
- tocScrol {
position: absolute; top: 0; /* just used to show how to include the margin in the effect */ /*margin-top: 20px; */ /* border-top: 1px solid purple; */ /*padding-top: 19px;*/ }
- tocScrol.fixed {
position: fixed; top: 0; }
- editPageTxt {
text-align: left; padding-left: 15px; }
- editPageTxt P {
clear: both; }
- toPageTop {
float: left; position: relative; top: 18px; left: 13px; color: #d13f31; } </style>
<script type="text/javascript"> $(document).ready(function() { var parentTable = $("#toc").parent(); $('#mytoc').append($("#toc").first());
$('#mytoc').find("#toc").attr("id", ""); parentTable.closest('table').remove(); });
$(document).ready( function() { var top = $('#tocScrol').offset().top - parseFloat($('#tocScrol').css('marginTop').replace( /auto/, 0)); var nav = $('#tocScrol'); var max = $('#indexing').offset().top - nav.height();
$(window).scroll(function(event) { // what the y position of the scroll is var y = $(this).scrollTop();
if (y > top) { // && signs are html decoded thus this construction if (y >= max) { nav.removeClass('fixed'); nav.css({ position : 'absolute', top : max - top }); } else { nav.addClass('fixed'); nav.removeAttr('style'); } } else { nav.removeClass('fixed'); nav.removeAttr('style'); } }); }); </script> <html> <html>
<a id="toPageTop" href="#">▲</a>
[<a href="http://openwetware.org/index.php?title=Biomod/2013/Aarhus/</html>Materials And Methods/Methods<html>&action=edit">edit this page</a>]
</html>
Methods
This section describes the methods that were used in the project.
Atomic force microscopy
Atomic force microscopy (AFM) is a method, used to visualize nanometer size samples with high resolution. The AFM consists of a small tip placed on a cantilever that can be moved in a highly precise motion by a piezoelectric element. The tip is placed closely to the sample and dragged across the sample. This can be done different ways that represent the several different modes of AFM. For biological samples, which are typically soft and easily destroyed, the less abrasive tapping mode is used. In this mode the cantilever oscillates close to its resonant frequency and taps into the sample. A signal can be recorded in two different manners, which is either by constant current and constant height. In constant height, the tip is oscillating a constant distance from the surface. Elements protruding from the surface will thus cause a decrease of the amplitude, due to atomic forces as Van Der Waals forces, electrostatic repulsion, etc. which can be measured and give a contour like depiction of the sample. When the tip is kept at a constant height, to maintain the set oscillation of the cantilever, it is possible to image the force between sample and tip and yet again produce a contour like depiction of the sample. [1]
Automated oligonucleotide synthesis
The setup is bought at BioAutomation Texas [1]. Automated oligonucleotide synthesis occurs in the 3' to 5' direction, which is opposite to the biosynthesis of DNA in vivo. Each synthesis cycle contains multiple steps, which result in the addition of one nucleotide per synthesis cycle. The synthesis can be divided into four different steps, which are detritylation, coupling, capping and oxidation. The detritylation step is performed under mild acidic conditions (triflouro acetic acid 3%), in which a 5’ alchohol is liberated for the synthesis to continue. Hereafter the phosphoramidite activated nucleotide is added, together with the nucleophillic catalyst 5-(Ethylthio)-1H-tetrazole to enhance the reactivity. A capping step is performed with acetic anhydride and N-methylimidazole in THF, to render non-coupled DNA strands inert in the subsequent steps of the cycle. Subsequntly oxidation is performed with I2 in a mixture of water, pyridine and THF. This is the end of the synthetic cycle. The cleavage of solid support is performed with AMA (Ammonium hydroxide/aqueous methylamine 1:1 v/v) or ammonium hydroxide, in which any protection groups and solid support is cleaved off, yielding the desired DNA sequence. [2]
Cell viability (MTT) assay
In order to determine if the decrease in measured luminescence was due to gene knockdown or was caused by a decrease in the number of living cells, the viability can be measured by using an MTT assay. As mitochondrial activity is relatively constant between living cells, a direct linear relationship exists between mitochondrial activity and the number of living cells in a sample. This activity can be quantified by adding the tetrazolium salt, 3-(4, 5-dimethylthiazolyl-2)-2, 5-diphenyltetrazolium bromide (MTT) to a sample of living cells. MTT is a yellow compound, that when reduced turns into purple formazan crystals. Metabollically active cells reduce the salt in their cytoplasm by i.a. dehydrogenase enzymes generating NADH and NADPH, and after solubilizing these crystals in a solvent such as DMSO, the absorbance can be measured and normalized to an untreated control.[3]
Click reactions
Click chemistry is a collective term for a group of modular chemical reactions that take place in high yields, are easy to perform and can take place in aqueous solutions. A widely used click reaction is the copper catalyzed azide-alkyne cycloaddition (CuAAC), also known as Huisgen 1,3-Dipolar Cycloaddition, in this context simply referred to as click reaction. This reaction utilizes the very specific reaction that occurs between alkynes and azides with a high yield, and is well suited for the conjugation of biomolecules. The reaction is catalyzed by copper which is added as Cu(II) and then reduced to Cu(I) by the action of a reducing agent, such as ascorbic acid which is added to the reaction buffer in an excess. A chelator is also included, e.g. TBTA, to stabilize Cu(I).[4]
Confocal microscopy
Confocal microscopy is an imaging technique widely used in biological fields. In a conventional wide-field microscope, thicker specimens, like cells, will produce a blurred image because not the whole sample can be in focus at once. It is not prominent at low magnification (max 10x) but at higher magnefication where a high numerical aperture lenses are used there is a limited depth producing a blurry image. In confocal microscopy this problem is circumvented by introducing a confocal aperture, such as a pinhole, in the path of the beam that forms the image, so that only the layer in focus will be detected. In practice, the sample is imaged with a laser, either by moving the laser in a raster mode or by moving the sample. To visualize the cellular localization of a given sample, this can be flourescently labeled and visualized on the image. To obtain a three-dimensional image, the focal plane is changed by moving the sample stage up and down, typically by a piezoelectric element. Recorded images from all focal planes are stacked in a computer program that produces a three-dimensional image. [5]
Dynamic light scattering
Dynamic Light Scattering measures the scattering of light emitted from a laser through a solution of aggregates as a function of time. As large aggregates diffuse at a slower rate than smaller molecules, scattering of light over time and size can be related. [6]
Ethanol precipitation
Charged molecules, such as nucleic acids, can be precipitated and concentrated by ethanol precipitation. The principle behind this method is that when adding ethanol to an aquous solution of nucleic acids, the ethanol, which is much less polar than water, will cause the dielectric constant of the solution to decrease, and thereby achieve a less efficient shielding of charges by water. A salt, such as sodium acetate is added, and once less shielded by the water, the cations can interact with the negative backbone of the nucleic acid and precipitate them. By centrifugation, this precipitate can be collected and re-dissolved in buffer. [7]
Fluorescence microscopy
Fluorescence microscopy is an imaging technique that is useful for visualizing cellular structures and intracellular physiological events. The method utilizes that fluorescence emission is dependent on the specific excitation wavelength of a certain compound and that the energy of excitation during single-photon absorption is greater than the energy of emission. This technique has a very high signal to noise ratio, which makes it possible to distinguish spatial distributions even in low concentration species. It is possible to use an organism’s autofluorescence, or a specimen can be labeled with a fluorophor. A typical fluorescence microscope employs filters and a dichroic beam splitter. The object is illuminated with the excitation wavelength through the same lens that collects the emission beam. The beam splitter transmits or reflects light depending on wavelength, thereby separating the excitation light from the fluorescence light. [5]
Freeze n’ squeeze
Freeze n' sqeeze is a simple method to purify double-stranded DNA fragments, or in our case DNA origamis from TBE agarose gels. The purification process happens through filtration in a spin column that consist of a filter cup (with 0.45 µm cellose acetate filter) contained within a special 2.0 mL “dolphin tube” for collection of purified sample. This method allows for recovery of DNA ranging from 50 bp to 23 kbases in size. The band of interest is cut out from the agarose gel and chopped into smaller pieces which are then transferred to the filter cup. The cup containing the DNA origamis is placed in a -20˚C freezer for 5 minutes. Following the freezing, the “squeezing” is done by centrifugating the filter at 13,000 × g for 15 minutes at room temperature, thereby releasing the liquid from the gel. Agarose debris is preserved inside the filter cup while the liquid at the bottom of the tube contains the recovered DNA origamis. [8]
Gel electrophoresis
A very commonly used method to separate and analyze macromolecules is by gel electrophosresis. The physical principle behind this technique is that a molecule with a certain electrical charge will move towards the electrode of the opposite charge, when placed in an electrical field. Molecules such as DNA and RNA have an overall negative charge, due to their sugar-phosphate backbone, and will move towards the positive pole of the field. The nucleic acids can be separated according to their size, as a larger nucleic acid will have a larger hydrodynamic size, and will be retained to a higher degree by the pores in the gel, and thereby move a shorter distance. In order to be able to compare and identify the bands of the gel, a ladder containing a number of bands of known sizes is included in the gel. The bands of the gels can be visualized by staining the gel with compounds such as ethidium bromide or less toxic dye, such as the SYBR gold or SYBR safe (Invitrogen), that was used in this project. Both compounds interact with the nucleic acids in the gel by intercalating the bases, and when irradiated with a laser of the correct wavelength, emit fluorescence, that can be detected by a scanner[REFERENCE: Weaver, R.F. “Molecular Biology” McGraw-Hill, 5. Ed. (2012): 76-80]. In this project we use polyacryl amide gel electrophoresis (PAGE) and agarose gels. Denaturing PAGE is used to check the click reactions with peptides and sisiRNA, where it is important to avoid secondary structure and other non-covalent interactions. For these reasons a denaturing loading buffer is used, and the gel is run at a high wattage with a metal plate attached to the glass plate containing the gel, to distribute and retain the heat. For origamis and annealed sisiRNA, agarose gels are employed. These are run at 4°C with non-denaturing loading buffers to avoid denaturing the samples.
LCMS
Liquid chromatography mass spectrometry (LCMS) is a chromatographic method, which just like HPLC, seperates molecules based on their polarity. This is done by using a stationary phase and a mobile phase. The mobile phase is then lead into a mass spectrometer and polarized by ESI (electro spray ionization), APCI (atmospheric pressure chemical ionization), etc. The now ionized molecules are then analyzed by mass spectry, as TOF (time of flight), quadropole, etc. This setup results in seperated molecules with the possibility of viewing the mass of each molecule in a complex mixture. [9]
Lipofection
As a transfection agent, Lipofectamine2000 (Invitrogen) was used. Lipofectamine is a cationic liposome based reagent used for introducing various sized nucleic acids into the cell. Lipofection formulations in general contains lipids with one or more positively charged nitrogen atoms. These allow the liposome to interact and complex with the negatively charged backbone of the nucleic acids, and thereby shielding the electrostatic repulsion from the membrane. The solution often also contains neutral co-lipids, that mediate the fusion of the liposome and the cell membrane, and thereby allow the nucleic acids to pass through to the interior of the cell. [10]
Luciferase assay
The luciferase gene, that was targeted for knockdown in these experiments, encodes a protein naturally found in the lanterns of male fireflies and catalyses the conversion of luciferin with the accompanying production of luminescence. The luminescence signal can be detected and used as an expression of enzyme activity and thereby gene expression. The reaction catalyzed by the luciferase enzyme occurs in two steps. First MgATP and luciferin (LH2) binds to the enzyme (LUC):
LH2 + MgATP + LUC →LUC-LH2-AMP + MgPPi
The next step is an oxidative decorboxylation of luciferin to oxyluciferin (OL) along with the production of light:
LUC-LH2-AMP + MgPPi → LUC-OL + CO2 + AMP + H2O + Light
Light is initially produced as a flash, followed by a slow decay, and the luminescence is therefore measured for over 10 seconds after addition of the substrate. As seen from the reaction schemes above, aside from the substrate, D-luciferin, Mg2+ and ATP is required for the reaction to take place, and are therefore included in the substrate buffer. Other stabilizing compounds are also added, such as Dithiothreitol (DTT) and EDTA that prevent inhibition of the reaction. As the optimal pH for the enzyme is 7.8, tris-glycine buffer is used, and the reaction is kept at room temperature which ensures optimal light emission. Coenzyme A (CoA) is also included in the buffer as this stimulates the light production by removing the product oxyluciferin from the enzyme, leading to a more constant emission of light. [11]
MALDI-TOF
Matrix-assisted laser desorption/ionization time-of-flight (MALDI-TOF) is a spectrometric mass analyzis method. In this method larger biomolecular compounds are ionized. To induce a charge on the molecules, a matrix loading buffer consisting of HPA/DPA (3-hydroxypicolinic Acid/Dipicolinic acid (1:9)) is used. The laser vaporizes the compound together with the matrix, which transfers a charge to the compound. By exposing them to an electric field the compounds can be moved. The field can be either of positive or negative charge, thus forcing the now charged molecules to fly towards the opposite pole. As the mass and velocity are inversely related, the flight time though the flight tube can be correlated to the mass.[12]
NMR
Nuclear Magnetic Ressonance is a method used to analyse an ensemble of molecules, and assign relative atomic correlation, thus making it possible to distingish molecules unambiguously. The methods exploit the so-called spin of atoms. A hydrogen atom has a spin of ½ which causes the spin to align to an outer magnetic field. By treating the aligned spin with another magnetic frequence, the rotation towards the outer magnetic field can be measured. As each atom experience a different magnetic environment, their specific rotation is unique, thereby enabling the identification of the specific atoms in a molecule.[13]
Reverse phase high performance liquid chromatography
Reverse phase high performance liquid chromatography is a chromatographic method to separate and purify molecules. The molecules are separated according to their hydrophobicity, so that the most hydrophobic molecules will interact the best with the stationary phase and elute last. In reverse phase HPLC a nonpolar stationary phase is used as opposed to a polar stationary phase in phased HPLC. The stationary phase is chosen on the properties of the molecules that need by be purified, and in the case of DNA, a reverse phased column is used. The mobile phase is typically a gradient of organic solvent of different polarity, such as acetonitrile or TEAA buffer. Both analysis and purification can be done using HPLC, as a collection tray is connected to the coloumn, allowing the collection of purified sample. [14]
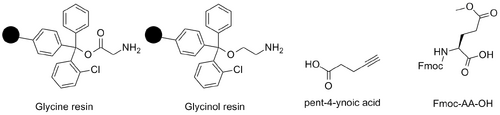
Solid phase peptide synthesis
The peptide synthesis is performed in the C-terminal to N-terminal direction. The synthesis is performed on a solid support, often called resin. Two methods are employed in the synthesis. One of these is tert-Butyloxycarbonyl (Boc) method, which uses a Boc as the protection group of the N-terminal amine. However, this requires hydrogen flouride as deprotection agent, which is not favorable due to the hazardeous nature of HF and the price of the needed apparatus for handling this chemical. An orthogonal method, namely the Fmoc (fuorenylmethyloxycarbonyl) strategy, has been developed. The Fmoc strategy resolves around the usage of fuorenylmethyloxycarbonyl as protection group, and cleavage is performed under basic conditions by piperidine. This is now a widely used method due to the overall milder conditions needed for the synthetic cycle. The resulting cleaved protection group is 9-methylene-9H-uorene which exhibits UV activity. This property is used in SPPS to determine whether or not a deprotection step has finished. When growth in UV absorbtion ceases, due to decreasing formation of 9-methylene-9H-uorene, all amines have been liberated from the Fmoc protection group. Piperidine also act as a so-called scavenger, preventing the newly formed 9-methylene-9H-uorene from reacting with the amine of the amino acid. The synthesis also requires a coupling reagent, which in most cases utilizes a diimide moiety, as for example HBTU (O-Benzotriazole-N,N,N',N'-tetramethyl-uroniumhexauoro-phosphate) or in this case HTCU (2-(6-Chloro-1H-benzotriazole-1-yl)-1,1,3,3-tetramethylaminiumhexauorophosphate). NMM (4-Methylmorpholine) is employed as a non nucleophillic base. [15]
Terminal transferase reactions
Terminal deoxynucleotidyl transferase (TdT) is a ~ 60 kDa template-independent polymerase that is able to elongate a nucleic acid with both ribonucleotides and deoxyribonucleotides as well as a number of synthetic nucleoside triphosphates, such as flourophores. The substrate for TdT activity must be at least three nucleotides long and contain a 5’ phosphorylation, but can otherwise be protruding, recessed or blunt-ended, double or single-stranded. No direct binding occurs between the protein and the primer, only electrostatic interactions with the nucleic acid backbone which causes an unspecific incorporation of bases. TdT also needs a free 3’ OH-group on the primer as the incorporation of a nucleotide without this will cause the extension to cease. This effect can be utilized to ensure that only one nucleotide is incorporated, by using dideoxy nucleotides. By adding Co2+ or other divalent ions such Mg2+ and Mn2+ as a co-factor to the reaction, the efficiency of the enzyme is improved. in the reaction makes tailing more efficient, however, it is shown that other divalent ions such Mg2+ and Mn2+, and the reaction can be stopped by adding EDTA. [16]
Transmission electron microscopy
Transmission electron microscopy (TEM) is an electron microscopy technique used to image nanoscale structures with high resolution. In TEM, an electron source is used to image a sample. An electron beam, usually with an energy of 100keV-400keV, is passed through the microscope and collimated by electromagnets before hitting the sample. The source, which is is typically either a tungsten filament or a LaB6 single crystal, is heated up enough for its electrons to escape from the Fermi level to the vacuum in the microscope. The samples are very thin, typically thinner than 100nm, although biological samples can be slightly thicker. A thin sample is required, as the image is formed by the beams that are transmitted through the sample and detected below the sample. The colors and contrasts on an image obtained by TEM, is dependent on what mode it is recorded in, either darkfield or brightfield, which was used in our project.
UV/Vis spectrophotometry
UV/vis spectrophotometry is a method that is used to determine the concentration of a certain compound in a solution. For analysis of nucleic acid concentrations, absorbances are measured at 260 nm. The absorbance is the logarithm of the relationship between intensities of incoming light and the light that passed through the cuvette or droplet:
[math]\displaystyle{ A=log\left(\frac{I_0}{I}\right) }[/math]
From the measured absorbance and the molar extinction coefficient of the molecule, the concentration of the strands can be determined using Lambert-Beer’s law:
[math]\displaystyle{ A=l\cdot C\cdot \varepsilon }[/math]
Where A is the absorbance, l is the length of the light path in the cuvette or droplet in cm and ε is the molar extinction coefficient in M-1. cm-1. [17]
References
-
Geisse, N. A. et al. AFM and Combined Optical Techniques. Mater. Today 12, 40–45 2009. [1].
-
Sinha, N. D. et al. Polymer support oligonucleotide synthesis. XVIII: use of β-cyanoethyl-N,N-dialkylamino-/N-morpholino phosphoramidite of deoxynucleosides for the synthesis of DNA fragments simplifying deprotection and isolation of the final product. Nucl. Acids Res. 12, 4539–4557 (1984).[1]
-
van Meerloo, J. et al. Cell sensitivity assays: The MTT assay. Methods Mol. Biol. 731, 237-45 (2011)[1]
-
H. C. Kolb, et al. Click Chemistry: Diverse Chemical Function from a Few Good Reactions. Angew. Chem. Int. Ed 40, 2004–2021 (2001).
-
Prasad, P. N. Bioimaging: Principles and Techniques, in Introduction to Biophotonics. (John Wiley & Sons, Inc., Hoboken, NJ, USA., 2004) [1]
-
Bloomfield, V. A. et al. Static and dynamic light scattering from aggregating particles. Biopolymers 54, 168-172 (2000).
-
Shapiro, D. J. Quantitative ethanol precipitation of nanogram quantities of DNA and RNA. Anal. Biochem. 110, 229-231 (1981). [1]
-
Thuring, R. W. J. et al. A freeze-squeeze method for recovering long DNA from agarose gels. Anal. Biochem. 66, 213-220 (1975). [1]
-
Harris, D. C. Quantitative Chemical Analysis Ch. 21. (W. H. Freeman and Company, New York, 2010).
-
Dalby, B. et al. Advanced transfection with Lipofectamine 2000 reagent: Primary neurons, siRNA, and high-throughput applications. Methods 33, 95-103 (2004). [1]
-
Ford, S. R. et al. Improvements in the application of firefly luciferase assays. Methods Mol. Biol. 102, 3-20 (1998). [1]
-
Friebolin, H. Basic One- and Two-Dimensional NMR Spectroscopy (Wiley Verlag GMBH & Ci. KGaA., Weinheim, 2005)
-
Harris, D. C. Quantitative Chemical Analysis Ch. 24. (W. H. Freeman and Company, New York, 2010).
-
Merrifield, R. B. Solid-phase peptide synthesis. Adv. Enzymol. Relat. Areas. Mol. Biol. 32 221-296 (1969).
-
Delarue, M. et al. Crystal structures of a template-independent DNA polymerase: murine terminal deoxynucleotidyltransferase. EMBO J. 21, 427-439 (2002). [1]
-
Harris, D. C. Quantitative Chemical Analysis Ch. 17. (W. H. Freeman and Company, New York, 2010).
</html>
<html> <head> <style>
- indexing {
/* float: left; position: center; */ background-color: #222; border-top: 2px solid #d13f31; color: #006e9c; margin: 0px; padding: 0px 0px 10px 0px; width: 100%; text-align: center; }
.footer-section { padding: 10px; display: table-cell; text-align: left; }
.footer-section-title { font-size: 20px; }
- footer-contents {
color: #006e9c; display: inline-table; }
.footer-section A { color: #006e9c; text-decoration: none; }
.footer-section A:HOVER { color: #00aeef; }
.footer-section ul { list-style-type: square; }
- sitemapTitle {
margin-top: 20px; font-size: 24px; }
- editFooter {
float: right; margin-top: -28px; margin-right: 5px; }
- editFooter A {
color: #006e9c; text-decoration: none; }
.cf:before,.cf:after { content: " "; /* 1 */ display: table; /* 2 */ }
.cf:after { clear: both; }
- bodyContent a[href^="mailto:"], .link-mailto {
background: url() no-repeat scroll right center transparent; padding-right: 0px; color: #006e9c;
}
</style> </head> <body>
SITEMAP | BIOMOD 2013 NANO CREATORS | Aarhus University
Copyright (C) 2013 | BIOMOD Team Nano Creators @ Aarhus University | Programming by: <a href="mailto:pvskaarup@gmail.com?Subject=BIOMOD 2013:">Peter Vium Skaarup</a>.
<img alt="Sigma - Aldrich" src="http://openwetware.org/images/3/39/Sigmaaldrich-logo%28transparant%29.png" width="300px" height="154px"> <img alt="VWR International" src="http://openwetware.org/images/2/28/Vwr_logo.png" width="300px" height="61px"> <img alt="Promega" src="http://openwetware.org/images/7/72/Promega.png" width="175px" height="105px" style="padding-right: 5px; padding-left: 5px;"> <img alt="kem-en-tec" src="http://openwetware.org/images/3/3a/Kementec.png" width="130px" height="129px"> <img alt="Centre For Dna Nanotechnology" src="http://openwetware.org/images/4/4f/CDNA_logo.png" width="420px" height="90px"> <img alt="Dansk Tennis Fond" src="http://openwetware.org/images/9/9a/Dansk_tennis.png" width="250px" height="53px">
</body> </html>