BioMicroCenter:Singular Sequencing

Singular Sequencing
The Center currently hosts a G4 platform by Singular Genomics. Singular’s G4 platform is a short-read mid-throughput sequencing platform delivering adequate sequencing data through its proprietary sequencing-by-synthesis chemistry at a significantly competitive price point. The G4 platform is unique in its flexibility to run 1 to 4 flow cells in parallel, each with 4 independent lanes.
Workflow
Library prep options for DNA or RNA are available for Singular sequencing. Once Singular-ready or Illumina libraries have been submitted, we will verify fragment size via AATI's Fragment Analyzer or Femto Pulse, adapt Singular flow cell binding sites S1/S2 if needed, and the quantify the final loading pool via qPCR. We can only guarantee minimum reads per lane if: a)the BMC has performed quality control, b)the samples are high-complexity, especially for the first 10 nt of read 1, and c)submitted libraries are at least 2 nM.
Requirements/Things to Consider
- All 4 bases (A, G, C, T) must appear within the first 10 cycles of read 1.
- Both forward and reverse read strands are present and accessible during sequencing. After the desired number of cycles have been reached for a given read, a block is introduced to prevent any continued extension of the growing read chain, but the generated read chain is NOT denatured/melted away before the priming and extension of the next read. For this reason, lane-by-lane requests must be either 50 or 150 paired-end compatible with a dual index of 8 nt each. All other requests will require a 4 lane (or 1 flow cell) minimum.
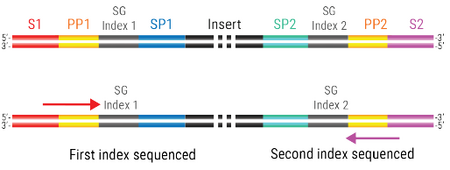
- Indexes are read off flow cell binding regions, S1 and S2. This is unlike modern Illumina platforms, which read off of sequencing priming regions of the final library (SP1 and SP2 in diagram above). For standard lane-by-lane requests, the first index will be read off the S1 region of the final library, and index 2 will be read off the S2 region. Compared to a typical Illumina library, a Singular index 1 would be the reverse compliment of the i5 index, whereas index 2 would be the reverse compliment of the i7 index.
- Each flow cell runs off a single reagent cartridge, thus each lane on that flow cell will be treated with the same primer cocktail by run. For standard lane-by-lane requests, this means all custom primers must be compatible with TruSeq, Nextera, TruSeq smaRNA, and Solexa sequencing primers.
- Singular also supports the conversion of Illumina libraries through either index replacement or anchor expansion for completed libraries that wish to be run on this platform. Index hopping rates are near zero, limiting the need for UDI's. Early quality metrics can be seen below.
Data Handling
Illumina Library to Singular Library Conversions and Custom Sequencing
The BioMicro Center can modify Illumina ready libraries by using the p5 and p7 flow cell binding sequences to attach Singular S1 and S2 flow cell binding sequences. However, this process leaves behind a 16 bp portion of the p5/p7 sequences requiring a different set of primers that will compete with the onboard S1 and S2 index primers. Like custom runs, these requests are rare and must be run on an entire flow cell.
Service Singular Sequencing INPUT Illumina or Singular libraries MIN VOLUME 12 uL MIN CONCENTRATION 4 nM INCLUDED SERVICES - Quality Control:Fragment Analyzer and qPCR
- Singular Sequencing
- Demultiplexing
ADDITIONAL SERVICES - Quality Control
- Singular library preperation
DATA FORMATS - FASTQ (stored 1 year)
- BCL (stored 30d)
QUALITY CONTROL - FASTQC
- Basic run metrics (alignment rate, complexity)
- Basic RNAseq metrics (where applicable)
- Basic paired end metrics (where applicable)
- Contamination checks
PRICING LINK SUBMISSION - MIT - ilabs
The G4 Platform
