Banta:BFCs

Banta Lab
Protein and Metabolic Engineering
| Home | Lab Members | Publications | Research Interests | Courses | Pictures | Positions Available |
Engineering of Enzymatic Biofuel Cells
It is becoming increasingly clear that our current global reliance on petroleum for transportation and other energy needs is not sustainable. As we explore other energy sources to meet ever rising energy demands, we will also need to explore new energy carriers and energy distribution systems. Fuel cells offer significant advantages in that they have the potential to convert chemical energy directly to electrical energy with minimal environmental impact. A standard fuel cell consists of an anode and a cathode, such that electrons are extracted from the fuel using a catalyst on the anodic side, and after passing through the external electrical circuit, they are combined with oxygen using a catalyst on the cathodic side, resulting in the production of water. Ideally, the catalysts should be robust, stable, specific, and highly active. Precious-metal based catalysts are very stable and active, but their use can be hampered by specificity issues. On the other end of the spectrum, the most specific and active catalysts known are enzymes, but these biological molecules do not have the stability of metal catalysts. Enzymes have already been naturally evolved to promote the transfer of electrons between substrates with an exquisite level of specificity. Unfortunately, they have not been evolved to operate in biofuel cells, and thus they will need to be further engineered in order to perform in this artificial environment.
We are working on engineering improved enzymes for use in biofuel cells. For example, we have improved the glucose oxidase (GOx) enzyme by site-specifically attaching gold nanoparticles to improve direct electron transfer. We have used computational design to make novel laccase enzymes. We have engineered laccase enzymes that bind DNA as well as carbon nanotubes (CNTs) so they can be immobilized on electrodes. We have created self-assembling enzymatic hydrogels that can be used on biofuel cell electrodes. And we are exploring multi-step enzymatic cascades for biofuel anodes, specifically enzymes involved in the citric acid cycle. Some of these enzymes naturally self-assemble into structures called "metabolons" and we are working to better understand the native metabolons and to create new metabolon-like complexes.
We have also developed kinetics-based models of biofuel cells. These models can be used to assess the performance of new enzymes in biofuel cells under different operating conditions, and they can provide insight into the rate limiting steps and processes in the operation of the biofuel cell.
Related Publications
- Liu F, Banta S, and Chen W. Functional assembly of a multi-enzyme methanol oxidation cascade on a surface-displayed trifunctional scaffold for enhanced NADH production. Chem Commun (Camb). 2013 May 8;49(36):3766-8. DOI:10.1039/c3cc40454d |
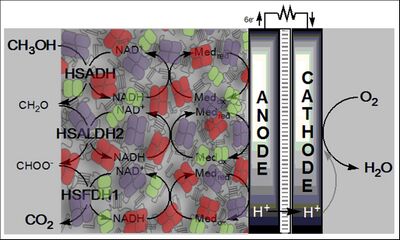
- Kim YH, Campbell E, Yu J, Minteer SD, and Banta S. Complete oxidation of methanol in biobattery devices using a hydrogel created from three modified dehydrogenases. Angew Chem Int Ed Engl. 2013 Jan 28;52(5):1437-40. DOI:10.1002/anie.201207423 |
- Glykys DJ, Szilvay GR, Tortosa P, Suárez Diez M, Jaramillo A, and Banta S. Pushing the limits of automatic computational protein design: design, expression, and characterization of a large synthetic protein based on a fungal laccase scaffold. Syst Synth Biol. 2011 Jun;5(1-2):45-58. DOI:10.1007/s11693-011-9080-9 |
- Campbell E, Meredith M, Minteer SD, and Banta S. Enzymatic biofuel cells utilizing a biomimetic cofactor. Chem Commun (Camb). 2012 Feb 11;48(13):1898-900. DOI:10.1039/c2cc16156g |
- Holland JT, Lau C, Brozik S, Atanassov P, and Banta S. Engineering of glucose oxidase for direct electron transfer via site-specific gold nanoparticle conjugation. J Am Chem Soc. 2011 Dec 7;133(48):19262-5. DOI:10.1021/ja2071237 |
- Szilvay GR, Brocato S, Ivnitski D, Li C, De La Iglesia P, Lau C, Chi E, Werner-Washburne M, Banta S, and Atanassov P. Engineering of a redox protein for DNA-directed assembly. Chem Commun (Camb). 2011 Jul 14;47(26):7464-6. DOI:10.1039/c1cc11951f |
- Glykys DJ and Banta S. Metabolic control analysis of an enzymatic biofuel cell. Biotechnol Bioeng. 2009 Apr 15;102(6):1624-35. DOI:10.1002/bit.22199 |
- Wheeldon IR, Gallaway JW, Barton SC, and Banta S. Bioelectrocatalytic hydrogels from electron-conducting metallopolypeptides coassembled with bifunctional enzymatic building blocks. Proc Natl Acad Sci U S A. 2008 Oct 7;105(40):15275-80. DOI:10.1073/pnas.0805249105 |
- Gallaway J, Wheeldon I, Rincon R, Atanassov P, Banta S, and Barton SC. Oxygen-reducing enzyme cathodes produced from SLAC, a small laccase from Streptomyces coelicolor. Biosens Bioelectron. 2008 Mar 14;23(8):1229-35. DOI:10.1016/j.bios.2007.11.004 |