BME494 Project Group4
Home People Course Projects Course Materials Syllabus Photos Wiki Editing Help
ABSTRACTOur construct is a light-responsive genetic toggle switch. The construct’s design was inspired by the toggle switch introduced during Unit 1. Our version improves upon that design by exchanging light for ligand molecules as a triggering mechanism for the switch between “ON” and “OFF” states. Light is advantageous over substrate-coupling mechanisms in applied systems because of enhanced control and efficiency during signal exposure. Our light-responsive toggle switch is superior to previous optogenetic methods because gene expression is self-propagated, and continues in the absence of the light signal once initiated. Our construct contains two core components: (1) a blue light responsive promoter derived from the plant Arabidopsis thaliana that was modified into BioBrick format, and (2) a green light promoter system previously used by the 2011 Freiburg iGem team. Our construct was designed using the red fluorescent protein reporter gene, which could be replaced by any gene of interest for a plug-and-play genetic engineering application.
BACKGROUND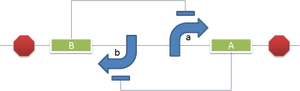 A toggle switch is an important motif in circuit design. In the context of genetic engineering, a gene circuit can be assembled that exhibits lateral inhibition between two expression states. These expression states could be unilateral, with an "ON" state and "OFF" state for a single gene or cascade. Alternatively, the expression states could yield two distinct gene products of interest. The genetic toggle switch is a fundamental design element that can be incorporated into any application. Currently, expression levels in such gene circuits are modulated based on the presence and targeted coupling of inducer ligand molecules. One limitation in the use of inducer molecules to achieve lateral gene repression is the reliance on enzyme kinetics. Achieving a switch from one expression state to the other requires that threshold concentrations of the ligand molecule be met. This need reduces the variability that users can employ and produces a lag in the rate at which the toggle switch can reset. In the discipline of brain science, researchers have developed a new method that uses light to exercise genetic control. This method is known as optogenetics. We sought to improve upon the genetic toggle switch by incorporating optogentics, and designed a toggle mechanism that responds to different wavelengths of light. One limitation to optogenetics is the requirement for continual signal exposure to sustain the genetic manipulation. Our light-responsive toggle switch self-propagates expression of the gene of interest in its "ON" state, even after the light signal is removed. With this design, we hope to achieve a more efficient toggle mechanism.
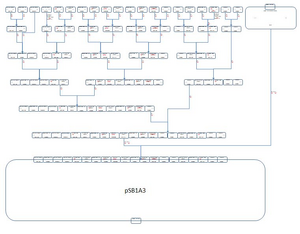
PROOF OF CONCEPT DESIGNOur construct consists of three key open reading frames (ORF 1, 2, 3). ORF 1 is regulated by a strong constitutive promoter, and contains coding regions for two light responsive proteins and the Lac I repressor, and a terminator sequence. ORF 2 contains a Lac promoter, a Tet operator, a reporter gene, BLRP, a second Tet operator, and a coding region for lac permease followed by a terminator site. ORF 3 contains a cpcG2 promoter and a coding region for the tet repressor, followed by a terminator. In the initial state, LacI inhibits the lac promoter and the reporter gene is silent. Exposure to blue light induces lac permease production mediated by BLRP activation. E. coli tranformants cultured in lactose will initiate lactose uptake, increasing intracellular lactose concentration. Intracellular lactose inactivates LacI, which de-represses the lac promoter and initiates reporter gene expression. At this time point, the blue light signal is terminated. Because there is not a terminator site separating the reporter gene and the BLRP, LacY is continually replenished while the lac promoter is active (thereby propagating the “ON” state despite lac permease half-life). The chassis will remain in the “ON” state as long as sufficient lactose is present in the culture media. To toggle the “OFF” state, the culture is exposed to green light. The receptor and response regulator proteins whose genes are located in ORF 1 respond to green light, initiating a phosphorylation mechanism that activates the cpcG2 promoter. This in turn induces TetR expression. The tet repressor targets the tet operators in ORF 2, thereby physically blocking RNA polymerases from transcribing the reporter gene and LacY. Lac permease is no longer replenished, and lactose concentration declines at a rate directly proportional to the rate of lac permease degradation. In the absence of intracellular lactose, the lac promoter is repressed and reporter gene expression ceases.


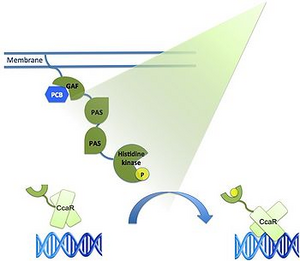 The construct's "OFF" switch relies heavily on the application of a green light responsive expression system previously reported by the Freiburg 2011 iGem team. This system contains three key parts: CcaS (a green light receptor), CcaR (a response regulator), and cpcG2 (a promoter targeted by CcaR). CcaS is a photoreceptor that autophosphorylates in the presence of green light (532 nm). CcaR contains an N-terminal receiver domain that is phosphorylated by CcaS, and a C-terminal DNA binding domain that selectively couples to the cpcG2 promoter when in a phosphorylated state.
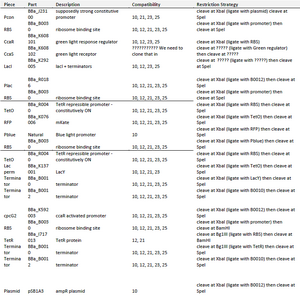
 TESTING
HUMAN PRACTICESOUR TEAM


|