BME100 s2016:Group6 W1030AM L4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||||||||||||||||||||||||||||||
|
OUR TEAM
LAB 4 WRITE-UPProtocolMaterials
PCR Reaction Sample List
HEATED LID: 100°C INITIAL STEP: 95°C for 2 minutes NUMBER OF CYCLES: 25 Denature at 95°C for 30 seconds, Anneal at 57°C for 30 seconds, and Extend at 72°C for seconds FINAL STEP: 72°C for 2 minutes FINAL HOLD: 4°C
Research and DevelopmentPCR - The Underlying Technology PCR Procedure: The initial step is when the template DNA is heated up to the point where it is denatured. This process takes about 3 minutes to heat up and 30 seconds for the DNA to separate. Annealing the DNA includes the newly separated DNA to be bound with a primer that attaches to specific nucleotides. The DNA is then extended at 72 C for 30 seconds in which Taq Polymerase binds to both the primer and template strand and translates dNTP’s. The final step is separating the DNA from the copied DNA. Lastly, the DNA is stored at 4 C in order to prevent degradation. 3. Adenine (A) pairs with Thymine (T), Thymine (T) pairs with Adenine (A), Cytosine (C) pairs with Guanine (G), and Guanine (G) pairs with Cytosine (C). In other words, AT and CG. 4. After the DNA is denatured, the primer is then added to the DNA strands. DNA polymerase is then added to the primer to match the complementary nucleotides together.
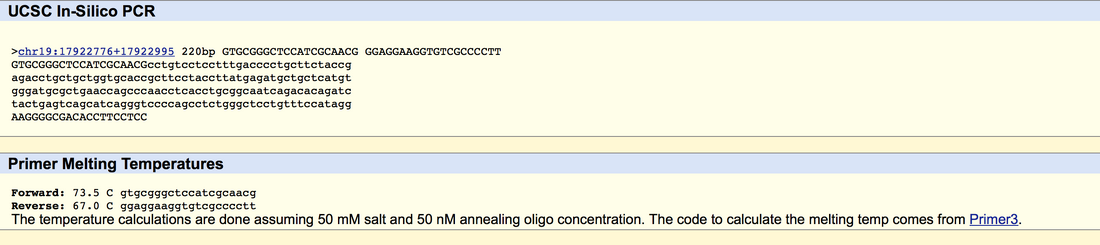
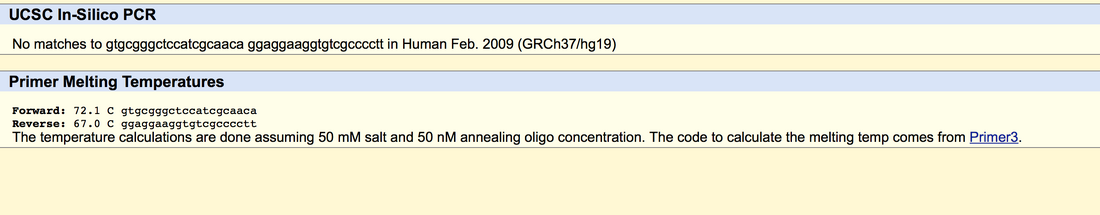
SNP Information & Primer DesignBackground: About the Disease SNP SNP stands for single nucleotide polymorphism found in Homo Sapiens. rs36686 is a reference SNP. This affects the 19th chromosome in the human genome and rs36686 changes the allele guanine paired cytosine. There is no clinical significance to rs36686 and it is linked to the disease Non-Hodgkin Lymphoma. B3GNT3 stands for betaGal-1, 3-N-acetylglucoseaminylatransferase 3 found in Homo sapiens.This is an enzyme that acts in part in regards to “ L-selectin ligand biosynthesis, lymphocyte homing and trafficking.”rs36686 changes the DNA code from CGC to CAC. This is an non disease allele that has minimal effect to the system. This allele change is located at 17811986. Primer Design and Testing The Non-disease forward primer GTGCGGGCTCCATCGCAACG. The Non disease reverse primer is located at 17812186. the sequence from left to right is GGAGGAAGGTGTCGCCCCTT. The disease forward primer is GTGCGGGCTCCATCGCAACA. The disease reverse primer is GGAGGAAGGTGTCGCCCCTT.
| ||||||||||||||||||||||||||||||||||