BME100 s2016:Group15 W1030AM L5
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | |||||||
OUR TEAM
LAB 5 WRITE-UPPCR Reaction ReportAll in all, in this experiment, Two patients submitted three samples of their DNA in order to look for a disease marker. In order to acquire an answer for either patient there was a list of clear steps that had to be followed outline in Lab 4 wiki lab report as well as the pre-lab reading. First all materials in the protocol for Lab A were acquired. After that it was necessary to cut the strip of empty PCR tubes in half to obtain to strips of four linked tubes in order to accommodate the OpenPCR machine. Now a black marker was necessary to label the sides of the empty tubes with labels specific to our group, and after that place the tubes in a tube rack We labeled all of the tubes accordingly: G151-1, G151-2, G151-3, G151-3, G152-1, G152-2, G152-3, G15P, and G15N. The labels indicate the trials for patient one (G151-1, G151-2, and G151-3), patient two (G152-1, G152-2, and G152-3), positive control (G15P), and negative control (G15N). Note: GP15 was left out of this lab report in the report following because it became redundant, twins only to signify group 15's PCR reaction tubes. At this point the experiment we began using the PCR positive control tube. Use the micropipette to transfer 50 μL of PCR reaction mix into the positive control tube. We were sure to discard the pipette tip after one use (every use) in order to avoid cross contamination. Overall, the pipetting of the samples were shaky at first but after many trials, the procedure was second nature. In order to pipette a sample using the micropipette, we had to set the sample amount (which is in microliter) press until the first stop, and place the micropipette tip within the sample, and release the stop slowly to draw in the sample. In order to release the micropipette sample, the user had to push all the way to the second stop slowly. To release the used tip press the release tab on the side of the pipette. Using a fresh pipette tip, transfer the positive control DNA/primer mix into the positive control tube. At this point the total volume in the positive control PCR reaction tube is 100 μL. The same steps will be followed for the negative control tube, patient 1 replicates 1, 2, and 3 tubes as well as patient 2 replicates 1, 2, and 3. Use the appropriate DNA/primer mix for the corresponding tubes, all of which should contain 100 μL by the end. After the process is completed for all tubes their lids should be closed tightly and they should all be taken to the assigned PCR machine. The tubes should then be placed into the slots in the heating block. Two groups will proceed with this step at the same time, so the machine can’t be started until 16 slots are filled.By the end go the lab there was still sample liquids in the test tubes, but any excess were disposed of properly and safely. Fluorimeter ProcedureWeb camera set-up Placing Samples onto the Fluorimeter
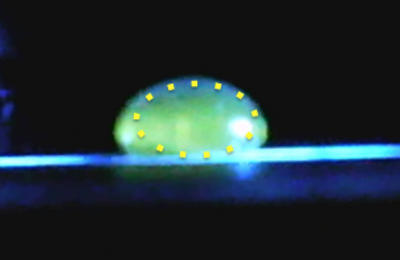
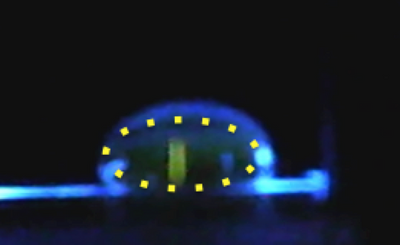
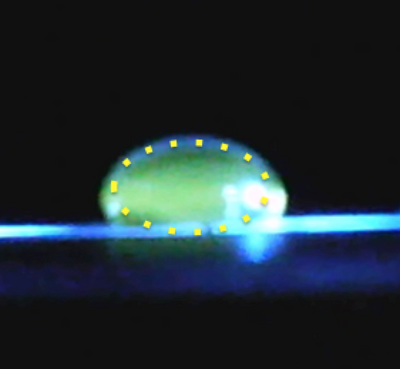
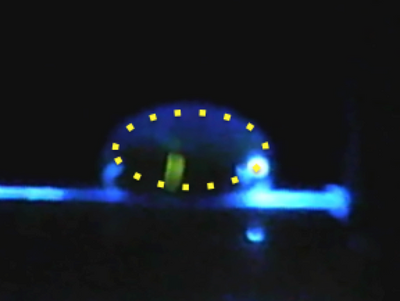
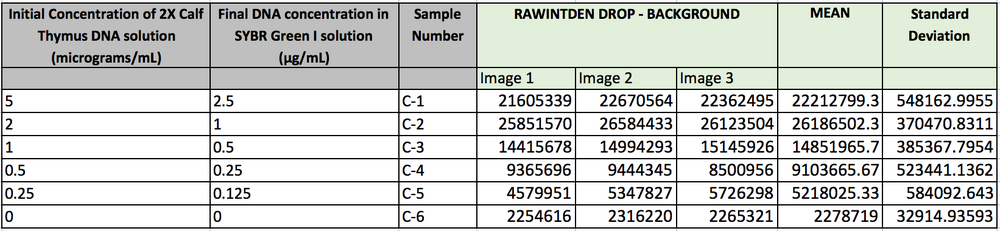
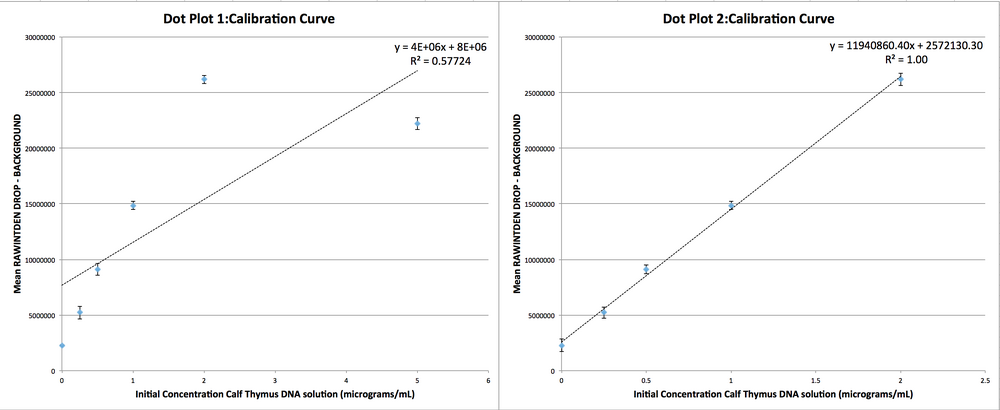
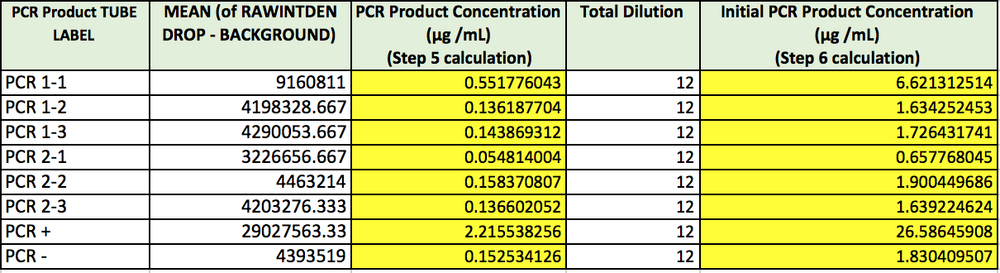
Data Collection and AnalysisImages of High, Low, and Zero Calf Thymus DNAHigh (5 μg/mL sample)Low (0.5 μg/mL sample)Zero (0.0 μg/mL sample)Calibrator Mean ValuesCalibration curvesImages of Our PCR Negative and Positive ControlsPostiveNegativePCR Results: PCR concentrations solvedPCR Results: Summary
Conclusions
| |||||||