BME100 s2015:Group8 9amL6
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||
|
OUR COMPANY
LAB 6 WRITE-UPBayesian StatisticsOverview of the Original Diagnosis System -Patients were tested for SNP through the usage and implementation of PCR. Multiple DNA samples were taken from each of the two patients involved in the experiment to ensure a greater amount of accuracy in determining whether they exhibited the genetic mutation. The DNA samples were then placed into a primer and subsequently placed into a thermocycler, initiating the process of replicating a specific, targeted sequence of DNA, provided the specific sequence existed. The DNA was warmed to approximately 94 degrees Celsius so that the DNA double-helixes unraveled into straight strands and disconnected from each other, a step in the PCR amplification process also known as "denaturization". Next, the DNA was cooled to approximately 54 degrees Celsius to initiate the annealing process, during which the primers and Taq polymerase worked into conjunction to continuously replicate a predetermined strand of the DNA. Finally, the thermocycler heated to 72 degrees for about 120 seconds, allowing the newly formed DNA to come together. Once the DNA samples had been sufficiently amplified, a flourimeter was used to check for the presence of the SNP in droplets of the DNA sample. If the strands of DNA were present, they reacted with the SYBR Green solution to produce an observable green hue.
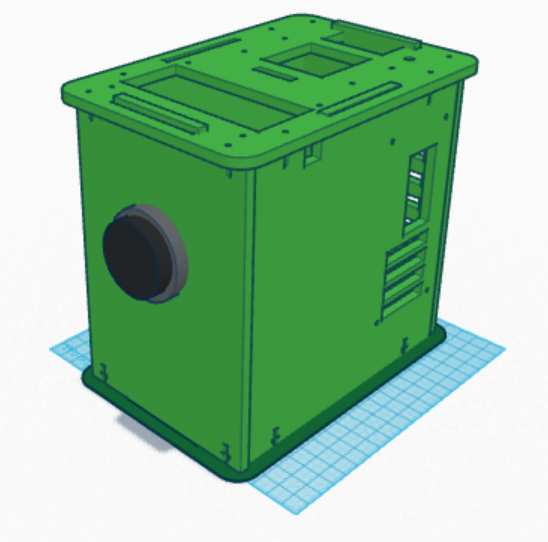
Computer-Aided DesignTinkerCAD
Feature 1: Consumables Kit-The tubes will be packaged in a very dark casing that reflects light -The reagents will also be packaged in dark casing to reflect light -The plastics will be made of a stronger polymer to ensure that they do not break in transport -Storage compartments for the tubes and tips will be added to the outside of the PCR machine. They will be stored in an air-tight and tight container so that they will not get exposed to any unnecessary light or contaminants. The problems with the consumables used in lab was that they didn't protect the reagents from light, and the plastic tubes and tips seemed a little weak. The packaging of the tubes will be very dark so that when the reagents and DNA are mixed in the tubes, the package will provide shielding from light interference. Also, the reagents will be enclosed in a dark casing so tin foil won't have to be used to prevent light from coming in. The storage compartments on the PCR machine will be the same way to prevent light interference as well. Lastly, the plastics will be made stronger to prevent cracking. Feature 2: Hardware - PCR Machine & FluorimeterHardware:PCR Machine To make the PCR process more efficient, the PCR machine could be manipulated to complete the PCR process in less than 24 hours. Also, more sample ports could be added so that more than 4 samples could be processed at once. The big change that will be made is the incorporation of real-time video analysis of the PCR process. This video will then be streamed real-time to an added spectrophotometer. The spectrophotometer will analyze the percent transmittance of light through the PCR sample. Hardware:Fluorimeter The spectrophotometer will measure percent transmittance at certain frequencies to check for the presence of a certain colored bioindicator. The spectrophotometer will measure the percent transmittance of light from the real-time video streaming from the PCR machine. This will make the process real-time, and will take only as long as the PCR machine machine takes to get through the process. This will eliminate the need of image J, which is very tedious and time-consuming. | ||||||