BME100 s2015:Group6 9amL4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||
|
OUR TEAM
| ||||||
LAB 4 WRITE-UP
Protocol
Materials
- Lab coat and disposable gloves
- PCR reaction mix, 8 tubes, 50 μL each: Mix contains Taq DNA polymerase, MgCl2, and dNTP’s
- DNA/ primer mix, 8 tubes, 50 μL each: Each mix contains a different template DNA. All tubes have the same forward primer and reverse primer
- A strip of empty PCR tubes
- Disposable pipette tips: only use each only once. Never re-use disposable pipette tips or samples will be cross-contaminated
- Cup for discarded tips
- Micropipettor
- OpenPCR machine: shared by two groups
Patient IDs:
- 1. 73039
- 2. 40429
PCR Reaction Sample List
| Tube Level | PCR Reaction | Patient ID | |
| GP | positivecontrol | none | |
| GN | negativecontrol | none | |
| G1-1 | Patient1Replicate1 | 73039 | |
| G1-2 | Patient1Replicate2 | 73039 | |
| G1-3 | Patient1Replicate3 | 73039 | |
| G2-1 | Patient2Replicate1 | 40429 | |
| G2-2 | Patient2Replicate2 | 40429 | |
| G2-3 | Patient2Replicate3 | 40429 |
DNA Sample Set-up Procedure
- 1. Collect materials
- 2. Cut strip of 8 empty tubes into two sets of four to allow for easy placement into the PCR machine.
- 3. Label each of the 8 tubes with a black marker: GP, GN, G1-1, G1-2, G1-3, G2-1, G2-2, G2-3.
- 4. Place tubes back into the rack
- 5. Pipet 50 micro-liters of PCR reaction mix into empty tube labeled GP (positive control). Throw away disposable tip.
- 6. Using a new pipet tip, pipet 50 micro-liters of positive control DNA/primer mix into the same tube.
- 7. Repeat steps 5 and 6 for the rest of the tubes using the appropriate DNA/primer mix.
- 8. Close lids tightly
- 9. Place tubes into the slots of the PCR machine where DNA will be replicated. Make sure your group can identify your samples from the other group's.
OpenPCR program
Heated lid: 100°C
Initial step: 95°C for 2 minutes
Number of cycles: 35
Denature at 95°C for 30 seconds, Anneal at 57°C for 30 seconds, and Extend at 72°C for 30 seconds
Final step: 72°C for 2 minutes
Final hold: 4°C
Research and Development
PCR - The Underlying Technology
Q1. What is the function of each component of a PCR reaction?
Template DNA: sequence of DNA that contains the area interested in. This sequence undergoes extreme heat to denature so replication can take place.
Primers: single stranded DNA that is complementary to template DNA. They are synthesized from 5' to 3' to complement the template DNA (3' to 5').
Taq Polymerase: a heat-resistant enzyme that links to the fork in the template DNA and creates the primers to complement the template DNA.
Deoxyribonucleotides (dNTP's): base pairs of DNA (A,T,C,G). In RNA, U is a dNTP in place of T.
Q2. What happens to the components (listed above) during each step of thermal cycling?
Initial step: 95 C for 3 minute:
- Denature at 95 C for 30 seconds:
- Anneal at 57 C for 30 seconds:
- Extend at 72 C for 30 seconds:
Final step: 72 C for 3 minutes:
Final hold: 4 C:
Q3. DNA is made up of four types of molecules called nuclotides, designated as A,T,C and G. Base-pairing, driven by hydrogen bonding, allows base pairs to stick together. Which base anneals to each base listed below?
Adenine (A): Thymine
Thymine (T): Adenine
Cytosine (C): Guanine
Guanine (G): Cytosine
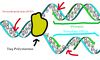 |
Picture References:
"DNA Replication." Wikipedia. Wikimedia Foundation. Web. 31 Mar. 2015. <http://en.wikipedia.org/wiki/DNA_replication>.




