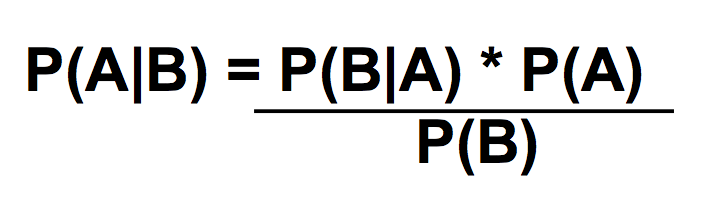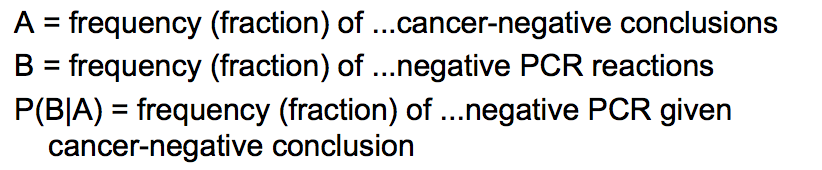BME100 s2015:Group4 9amL6
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||
|
OUR COMPANY
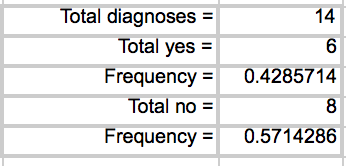
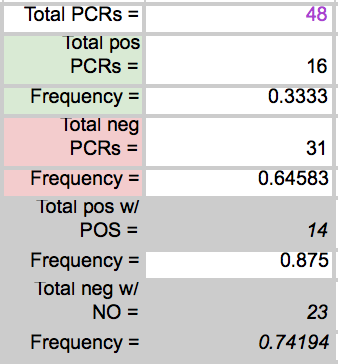
LAB 6 WRITE-UPBayesian StatisticsOverview of the Original Diagnosis System In order to test the patients for the disease associated SNP, 8 teams of 5 students each in the morning section were able to diagnose 16 patients each. Each of the patients were replicated three times in order to prevent error. A larger sample size decreases the amount of error. Images of the samples on the fluorimeter were taken and analyzed for fluorescence. In order to reduce error when the images were being taken, the same phone was used each time in order to take the photos, the settings on the camera were the same for each photo, the drop was in the same place the same distance away from the phone's camera, and all other light was excluded by enclosing the drop and camera in a dark space. For ImageJ, the image was taken from underneath the drop each time. Each PCR sample was examined using three images from each drop in ImageJ. Overall, looking at the class's final data from the spreadsheet, it can be understood that most of the tests were successful. One group's data was eliminated from overall analysis, because the data was unreliable. Overall, 14 patients' data were analyzed. Of the 14 patients, 10 were diagnosed correctly. One patient was deemed inconclusive. What Bayes Statistics Imply about This Diagnostic Approach Calculations for the first part were to determine the probability that the sample actually has the cancer DNA sequence, given a positive diagnostic signal. This value was very close to 1, so that there is a high probability that if there was a positive signal, the sample had the cancerous DNA sequence. Calculation for the second part were to determine the probability that the sample contains a non-cancer DNA sequence, given a negative diagnostic signal. This value was not very close to one, but it is significant enough to claim that there is over a fifty percent probability that if the sample gave a negative diagnostic signal, it contained the non-cancer DNA sequencel. Calculations for the third part were to determine the probability that the patient will develop cancer, given a cancer DNA sequence. The probability for this was close to 1, so there is a high chance that if a patient has the cancer DNA sequence, they will develop cancer. Calculations for the fourth part were to determine the probability that the patient will not develop cancer, given a non cancer DNA sequence. In the last part, the probability was also found to be high and close to one, meaning that there is a reasonable probability that if the patient has the non-cancer DNA sequence, the patient will not develop cancer. During the PCR detection steps, many errors could have occurred that may have affected the results of the Bayes values. One error may have been when using the micropipettor to combine the different substances. Some group members did have difficulty, so contamination may have been possible, causing more samples to show as positive rather than negative. Another error in analysis could have been in using ImageJ. Selecting the exact area each time was difficult and could have resulted in different results each time. Another error could have been in taking the photos of the water drops each time. The camera may have been a different distance away from the drop each time, resulting in different values when it was time for ImageJ analysis. Computer-Aided DesignTinkerCAD The TinkerCAD tool was used in this lab in order to recreate a design of one of the components of the PCR process. The tool was used to add extra parts to the already available design and increase some of the dimensions of the existing product. Our Design
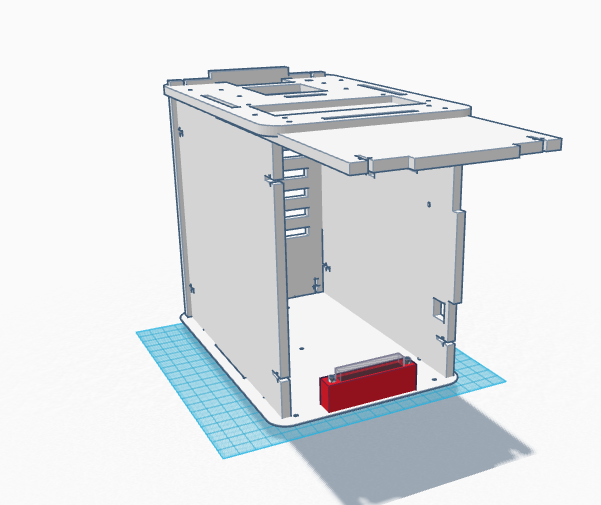
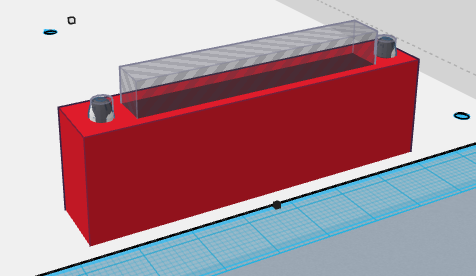
One of the issues with the original OpenPCR design was that there was a lot of difficulty with the camera on the phone and getting it to stay steady when taking the photo. Not only that, the system overall was flimsy and didn't seem to have much support. It didn't seem capable of completing what it should b able to do. We changed out design by increasing the thickness of each of the walls of the system and added a slot in the front wall of the device for the phone to rest on.
Feature 1: Consumables KitSome of the weaknesses to working with the consumables in the SNP diagnostic system was there was some confusion with what was to be added to what in order for the DNA to be properly tested and how it was difficult to use the tools for those that had no prior experience. Each consumable kit will come with a micropipettor, tips, and eight small plastic tubes. All the tools that are meant to be used together will be packaged together so that the user will have a better understanding of what items belong with which. Each of the items will also be thoroughly labeled with included instructions for the user to understand what to do. There instructions will also include a separate section directly towards the use of the micropipettor so that the user understand how to use it to ensure that no contamination will occur. The eight test tubes will be separated into three groups: one group of two tubes and two groups of three tubes. The one group of test tubes will contain the positive and negative controls and the two groups of test tubes will already have the appropriate amount of PCR reaction mix so that the user does not have to worry about pipetting the correct amount of solution into the tube. All the user will have to do is provide the DNA template to be tested.
Feature 2: Hardware - PCR Machine & FluorimeterThe PCR machine and fluorimeter are included in the overall system. Each of the walls of the system overall are easy to breakdown and separated so that there are a total of six flat sheets that can be reconstructed to form the box that is the machine. A new addition to the system is a screw in cradle in which the phone rests in so that the images of the drops can be taken all the same distance away from the drop and that the phone remains stable in the box. In packaging the overall device, the walls of the system will be broken down and stacked on top of one another. Each of the sides will be labeled with a number 1-6. On top of the six walls of the machine will be a set of instructions on how to reassemble the machine using the corresponding numbers 1-6. Another set of instructions will be included for the DNA template samples, PCR reaction mix, and micropipettor. On those instructions will be the individually packed micropipettors and tubes. What is different about this device is that the PCR machine and the fluorimeter are combined in that they are all one machine. One side of the machine is responsible for the DNA replication while the other side is responsible for capturing the image of the drop and analyzing it so that the procedure can be completed much faster and efficiently.
| ||||||