BME100 s2015:Group4 9amL5
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||||||||||||||||||||||||||||||
|
OUR TEAM
LAB 5 WRITE-UPProcedureSmart Phone Camera Settings
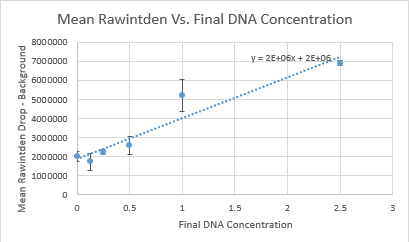
Calibration
Data AnalysisRepresentative Images of Negative and Positive Samples
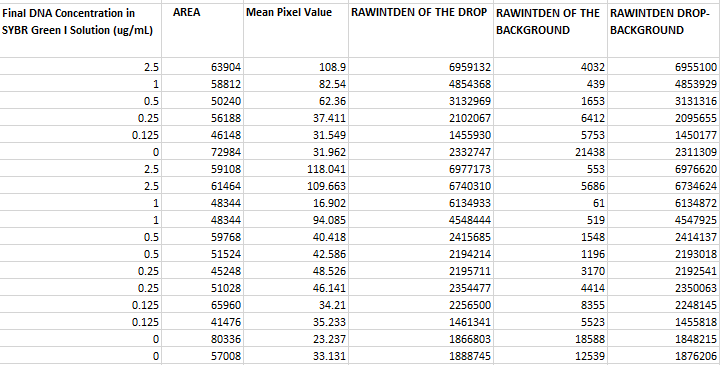
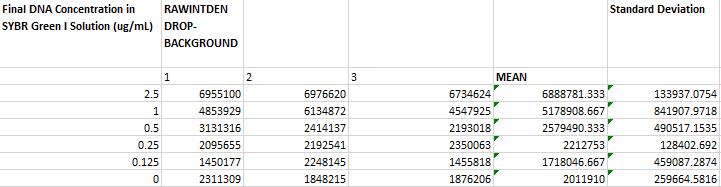
Image J Values for All Calibrator Samples
Observed results
Conclusions
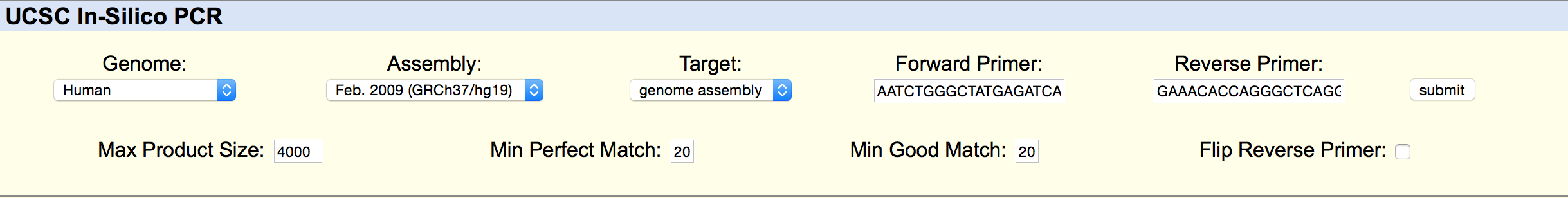
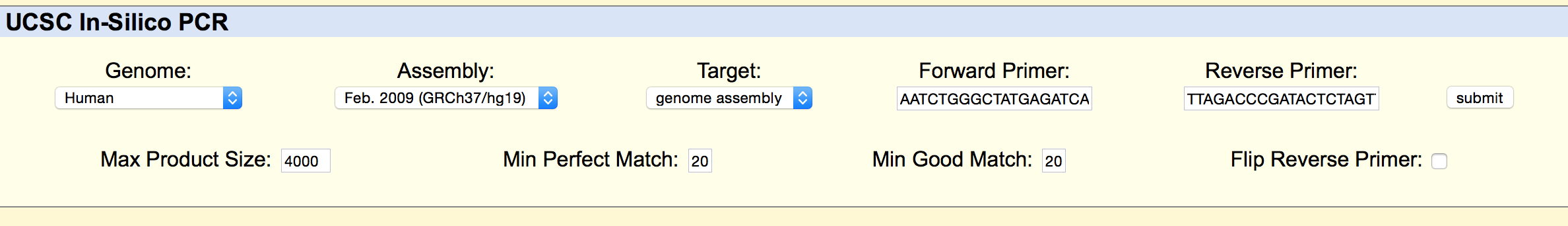
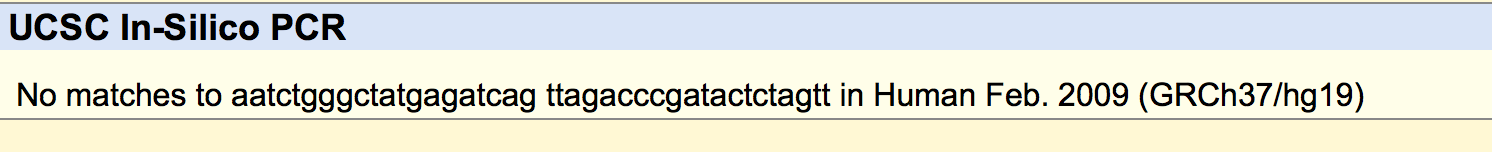
SNP Information & Primer DesignBackground: About the Disease SNP A SNP is a single-nucleotide polymorphism, a DNA sequence variation occurring within a population. In this case, the SNP being examine is rs268, found on chromosome 8 in homo sapiens. The position of the SNP is 19956018. Typically the allele is AAT, but is changed in disease to AGT. The significance of this allele is pathogenic, meaning that it is capable of causing disease. The disease associated with this SNP is coronary heart disease. Is is associated with the lipoprotein lipase gene, which is responsible for the bondings of many different lipids. Many studies have been completed confirming the existence of a close relationship between high levels of lipids such as cholesterol and triglycerides and high candidacy for coronary heart disease. Primer Design and Testing In the first part of designing the primers, it was necessary to validate the two non-disease primers that were designed using the instructions from the lab workbook. The non-disease forward primer was composed of 20 nucleotides that ended with the SNP from the top strand of DNA. These nucleotides were recorded reading left to right. The last nucleotide of the sequence was at position 19956018, which is the position of the position of the SNP. In order to find the non-disease reverse primer, the position 200 bases to the right of the disease SNP was used. The bases were taken from the second, bottom strand of DNA, read left to right, starting at the 19956218 position, 200 bases to the right from the disease SNP base. The bases were recorded until there was a total of 20 bases. The primers tested were as follows. This is the information inputted into the UCSC In-Silico PCR website These are the results of the input. From the results, it can be seen that there is a 220 bp sequence from chromosome 8, the same chromosome the SNP, rs268, is found on. This confirms that these are the forward and reverse primers found on this chromosome in a non-disease genome. Then in the second part of designing, disease forward and reverse primers were created. The final base of the non-disease forward primer was changed from A to G, which is the disease SNP base. The disease reverse primer was found using the second, bottom strand of DNA, beginning with the base in the same position as the SNP, and then completed recording the bases left to right until there were a total of 20 bases to compose the primer. The primers tested were as follows: This is the information inputted into the website These are the results of the input From the results, it can be seen that there are no matches to these primers. This is because they are being tested using a non-disease human genome sequence. The disease primers are being compared against a genome that do not have disease. | ||||||||||||||||||||||||||||||||||