BME100 s2015:Group4 9amL4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | |||||||||||||||||||||||||||||||||
|
OUR TEAM
LAB 4 WRITE-UPProtocolMaterials
PCR Reaction Sample List
OpenPCR program
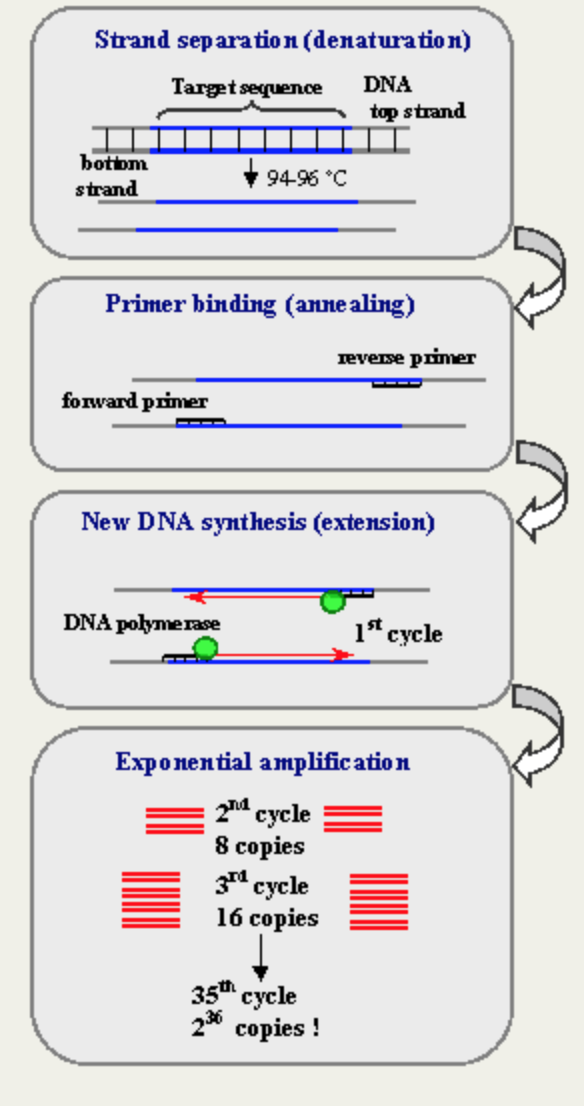
Research and DevelopmentPCR - The Underlying Technology There are many components to a PCR reaction. A full reaction requires a DNA template, a DNA polymerase, primers, and nucleotides. The DNA template is a sample of DNA that contains the sequence that needs to be analyzed. This template is then manipulated in order to separate the strands of DNA from each other. DNA polymerase is an enzyme that creates complementary strands of DNA to the target sequence of DNA. The polymerase used in this reaction is Taq DNA polymerase. Primers are small pieces of single strands of DNA. These small pieces of DNA form together to create the complimentary strand of DNA. The polymerase then begins to create the new strands starting at the end of the primer. Nucleotides are the basic building blocks of DNA. They are the bases A, T, G, and C. These building blocks are used by the polymerase and primer in order to create the complementary strand of DNA.
During the INITIAL STEP, the DNA is heated to 95*C in order to begin the denaturing process. During the DENATURE step, DNA is heated even further in order to separate the double strands of DNA. The heat causes the hydrogen bonds within DNA to break, releasing the two strands. During the ANNEAL step, the temperature is lowered in order to allow the primers to adhere to the now single strands of DNA. During the EXTEND step, the temperature is increased slightly. At this temperature, the polymerase then begins working with the primers in order to begin to create complimentary strands of DNA. The FINAL STEP is to ensure that all complementary nucleotides have been attached to the complimentary strand. The FINAL HOLD is used for an indefinite period of time in order to maintain the complimentary strands of DNA until it is time for the strands to be used. Adenine (A) bonds to Thymine (T)
| |||||||||||||||||||||||||||||||||