BME100 s2015:Group1 12pmL4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||||||||||||||||||||||||||||||
|
OUR TEAM
LAB 4 WRITE-UPProtocolMaterials
HEATED LID: 100°C INITIAL STEP: 95°C for 2 minutes NUMBER OF CYCLES: 35 Denature at 95°C for 30 seconds, Anneal at 57°C for 30 seconds, and Extend at 72°C for 30 seconds FINAL STEP: 72°C for 2 minutes FINAL HOLD: 4°C
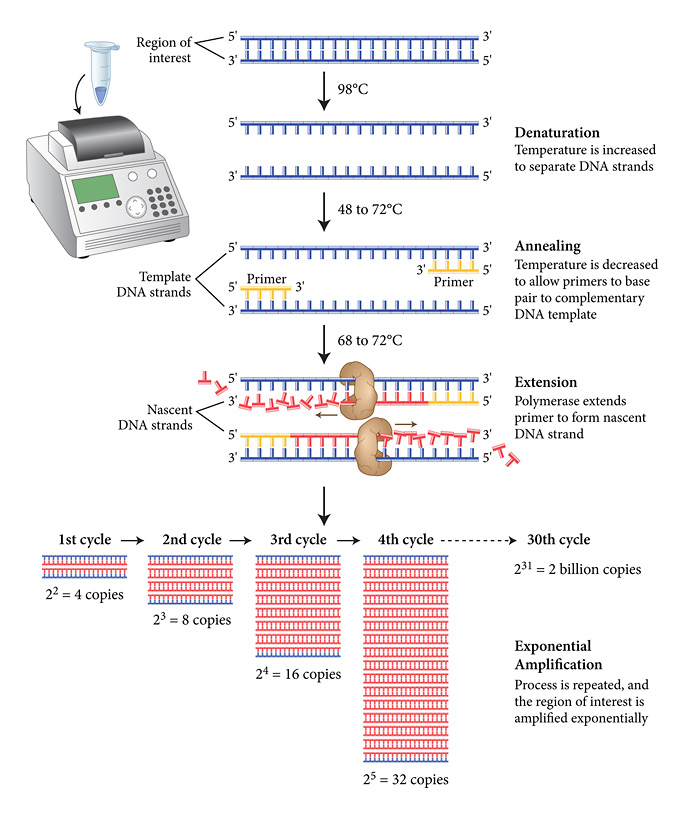
Research and DevelopmentPCR - The Underlying Technology Function of components in a PCR reaction. The template DNA is the original copy of base pairs and their specific order. The primers are both the beginning and endpoints of the specific section of the DNA to be copied. Taq polymerase reads single DNA strands and allocates a strand of its base pairs to create a new DNA strand. Deoxyribonucleotides are the base the pairs that taq polymerase attaches to a single strand of DNA to create a new strand. Thermocycling process. The initial step of 95 degrees celsius for 3 minutes of the thermal cycle is to ensure the DNA is fully "melted" and that it is no longer in its helix shape. The denaturing step in the cycle is the first step in the reoccurring process. The template DNA is split into two separate strands. During the annealing step dNTP's naturally try to bind themselves to the single strand. During the extending step the taq plymerase reads the single stranded DNA and attaches the appropriate dNTP's. The final step allows the DNA to solidify and is the last extension of the cycle. The final hold significantly slows down the reaction and allows for storing. Base pairs. Adenine and Thymine bind to each other while Cytosine binds to Guanine. Base pairing thermal cycling. Base pairing occurs during the annealing and extending steps. During the annealing step the base pairs are naturally formed due to 57 degree celsius temperature. During the 72 degree celsius temperature the taq polymerase works at its optimal level attaching base pairs to the single DNA strand.
| ||||||||||||||||||||||||||||||||||