BME100 s2015:Group14 12pmL4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||||||||||||||||||||||||||||||||
|
OUR TEAM
LAB 4 WRITE-UPProtocolMaterials
PCR Reaction Sample List
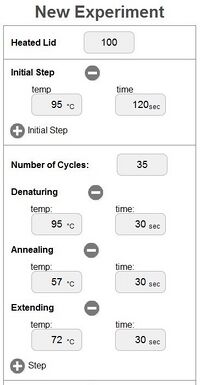
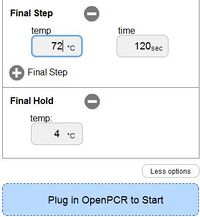
OpenPCR program
Research and DevelopmentPCR - The Underlying Technology A PCR takes a target sequence of DNA and exponentially increases the quantity of that target sequence. There are four main molecules needed for a PCR:
In a PCR, the template DNA is the DNA strand that has the target DNA sequence in it. The RNA primers in a PCR are short RNA strands that have a specific sequence. There are two primers needed to get a target DNA sequence to replicate in full. RNA primers have a specific ribonucleotide sequence that allows them to bind to a template DNA at a desired target sequence. Genetic technology allows specific RNA primer strands to be synthesized which can base pair with targeted DNA sequences. This makes sure that only target DNA can be replicated. Primers are needed to allow for DNA replication. Taq DNA polymerase cannot bind to a DNA molecule and start replication if there is not a primer for it to bind to. Once bound, Taq polymerase elongates the DNA sequence moving from the 5' end to the 3' end. Taq polymerase cannot replicate, however, without the presence of deoxyribonucleotides. These are the individual building blocks of DNA molecules. There are 4 types of deoxyribonucleotides found in DNA. These are Adanine (A), Guanine (G), Cytosine (C), and Thymine (T). In DNA, A always binds with T, while C always binds with G. These are called base-pairs. When Taq polymerase replicates DNA, it matches up the nucleotides it is adding to the corresponding base-pair. PCRs run in a cycle. In the initial step, the PCR tubes are heated at 95 degrees Celsius for 2-3 minutes. In this step, the DNA inside the test tubes begins to denature, which allows for primers to bind to the DNA and for polymerase to begin replication. The cycles then begin. During the Denature step, the PCR tubes are heated at 95 degrees Celsius for 30 seconds. Just like the initial step, this step makes sure that all DNA strands are cleaved in half so that replication can occur. During the Anneal step, the PCR tubes are cooled to 57 degrees Celsius for 30 seconds. In this step, RNA primers bind to target DNA sequences through base-pairing. The 30 seconds is a control to make sure that all of DNA has been bound by primers. For the Elongation step, the PCR tubes are heated to 72 degrees Celsius for 30 seconds. During this step, Taq polymerase binds to the DNA at the site of the primers. Deoxyribonucleotides are then bound together into a new DNA strand which is made up of the opposite bases from the template strand. Base-pairing in this step results in the replicated DNA strands. The cycle then starts over at the Denature step. After the desired number of cycles are completed, the PCR tubes are kept at 72 degrees Celsius for 2-3 minutes. This step ensures that all of the DNA has been replicated and that the process has completed. Finally, the tubes are cooled to 4 degrees Celsius. Doing so prevents the chance that the PCR reaction will continue longer than desired. When everything has been completed, the amount of DNA in the test tubes can be millions to billions of times larger than the amount initially present.
| ||||||||||||||||||||||||||||||||||||