BME100 f2013:W900 Group6 L6
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||
OUR COMPANY
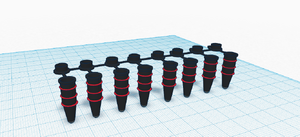
LAB 6 WRITE-UPComputer-Aided DesignTinkerCAD On November 20th, we worked with the TinkerCAD tool to design our improvements for the PCR-based Diagnostic Test. We used the TinkerCAD software to redesign the PCR Tubes, the Open PCR machine, the phone holder for the fluorimeter and the packaging for these devices. First, we worked on the PCR Tubes and made them blacked out in order to prevent the light from bleaching the SYBR green. We also added measurement markings on the sides of the PCR tubes in order to easily identify the volume of the solution in the tube. We also used the TinkerCAD tool to design the packaging for the device as well as the PCR machine (which was more of a software fix than an actual design flaw).
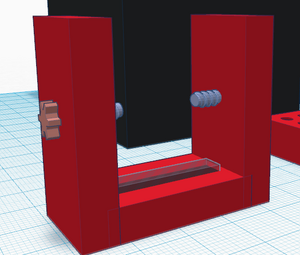
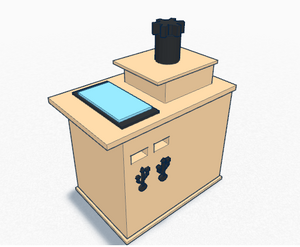
One possible way to use the TinkerCAD tool for something practical is the camera holder aspect of the fluorimeter. TinkerCAD allows us to design an object easily and is a free application, so we can design a project without the need to buy materials and supplies for the prototype. When the model is complete, TinkerCAD allows users to print off the design via 3D printer. We changed the design of the camera holder because the one used in the previous lab would not hold the phone in an upright position as it did not directly fir the phone. We developed a new design for the phone holder by making it adjustable so that it is compatible with various models of phones.
Feature 1: Cancer SNP-Specific PrimersBackground on the cancer-associated mutation Rs17879961 is a pathogenic single nucleotide polymorphism (SNP) that is found in Homo Sapiens. This single nucleotide polymorphism (SNP) is found in the 22nd chromosome of the 23 that Homo Sapiens contain. Rs17879961 affects the gene Checkpoint Kinase 2 (Chek2). This gene is a cell cycle checkpoint regulator that suppresses tumors and halts cell cycle progression when DNA is damaged.
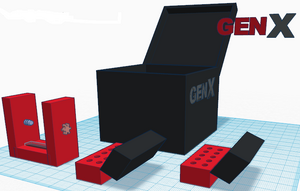
How the primers work: It beings with identifying the "target sequence," in this case the cancer-associated SNP rs17879961 . The primers specifically binds to the 3' end of the target sequence. Note that primers will only bind to the sequences that are complimentary to their own sequence. The primers then direct a DNA polymerase to synthesize the complementary DNA strands. Only DNA with the target sequence will be copied, because DNA polymerase can only copy the molecules with the primer attached; this is the reason why the non-cancer alleles will not be amplified. With each PCR cycle, partial target sequences and double stranded molecules, which contain only the target sequence, are “created” (also known as “amplified”). As the cycles continue, more target sequences of double stranded molecules (with the target sequence: rs17879961) and partial target sequences are produced. Consequently, with each PCR cycle, the number of strands with the target sequence grows exponentially (2^n). Essentially, after all cycles are complete, the end result will be a large number of pure target sequence molecules that only code for the SNP rs17879961 (the cancer-associated marker). Feature 2: Consumables KitThe GenX PCR-Based Diagnostic Test includes the PCR tubes, the PCR tube container (as well as holder), the newly designed camera/phone stand for the fluorimeter, as well as a micropipetor and the associated disposable tips.
Feature 3: PCR Machine HardwareThe PCR Machine will be used in the same way as it was set up during the previous lab, the only difference will be the logging software used during the process. The PCR machine will duplicate the DNA strands in order to detect the cancer segment of the DNA, by denaturing the DNA with high temperatures, once the DNA is separated, the annealing process begins and primers are attached to the DNA segments, thus synthesizing complementary strands of DNA and held at a constant temperature to ensure the DNA is extended completely. It was noted that one of the weakness for the PCR machine was that it didn't have a way of logging recorded data during the time that the machine was running. Our team is going to address this problem by creating a communication connection between the data of the PCR machine to a computer. We will be doing this by adding a logging program and USB connection to the PCR machine that will automatically start and record all data being given off by the PCR machine. This recording of the data will help users determine if the PCR machine is properly functioning before testing various solutions for the cancer allele. Feature 4: Fluorimeter HardwareOur lab group determined that the fluorimeter completed its job efficiently. We did, however, alter the design to the phone stand used within the fluorimeter. We designed a new stand that allows various models of phones to be used in the fluorimeter in the upright position.
Bonus Opportunity: What Bayesian Stats Imply About The BME100 Diagnostic ApproachCalculation 3 is the positive predictive value: the probability that a cancerous patient will test positive with the PCR-based diagnostic test. The percent value for calculation 3 is a small value and has low specificity, telling us that there are a number of results that came back as false-positives. This informs us that some patients are non-cancerous but came back with a positive test result from the PCR-based diagnostic test. This suggests that the test is not a good standard when testing for cancerous patients. Calculation 4 is the negative predictive value: the probability that a non-cancerous patient will test negative with the PCR-based diagnostic test. The percent value for calculation 4 is close to one and has high sensitivity, suggesting that this test is a good standard when testing for non-cancerous patients. Instruction ManualThe Open PCR machine takes a specific strand of DNA and amplifies it by making millions of copies of a specific segment of DNA. To do this process, it goes through several heating and cooling cycles, allowing the DNA to split at high temperatures (at about 95 degrees Celsius) and make copies at lower temperatures. The amount of cycles can be set and changed through the software on a computer that is connected to the Open PCR machine through a USB cable. Before beginning the diagnostic tests of the varying solutions, it is important that PCR goes through a test run. With the new USB feature on the PCR machine, users can walk away from the PCR machine while it is running through tests. Upon return, the user can plug the USB into a computer and view the log of the PCR machine ensuring that the PCR is reaching proper temperatures and cycles in the proper amount of timeframe. After going through the testing process and ensuring the PCR machine is efficient and accurate, an enzyme is added to the PCR tubes that allows a complementary strand of DNA to be paired when the temperature is at a lower temperature (in the case of our lab it was 4 degrees Celsius). The process is repeated until DNA is amplified to millions of copies. For the single drop fluorimeter we first have to place the fluorimeter in a set position and add a glass slide to which we place a drop of DNA solution that is mixed with SYBR green dye, a nucleic acid stain that usually binds to certain DNA and allows us to detect the cancer segment of DNA in our previous lab due to the fluorimeter being able to measure the intensity and wavelength of the light that passes through the DNA solution. Then the camera/phone holder is placed near the fluorimeter in order to take a photo of the DNA solution while the fluorimeter passes light through it, this is where we made modifications to the process, we redesigned the holder so that it was adjustable to any sort of phone or camera, where the original was supposed to be "Universal" but could not hold the device in an upright position. Then all of the components needed to be covered to reduce outside contamination and this includes the cover which we used a box to "blackout" the area in order to obtain the best picture possible to determine the status of the patient, whether they have cancer or not, from the information obtained by the fluorimeter. |
||||||