BME100 f2013:W900 Group4 L6
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||
OUR COMPANY
ACUSpeed PCR
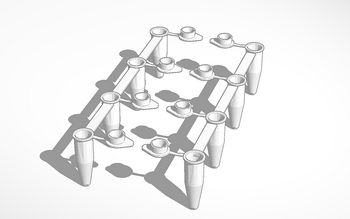
LAB 6 WRITE-UPComputer-Aided DesignTinkerCAD The TinkerCAD tool is a program that was used in lab on November 20th that allowed people to create files compatible for 3D printing. What TinkerCAD does is provides users with shape, number, and letter templates that are to be combined in a grid that makes an object in three dimensional space. Operators of TinkerCAD are exposed to a whole new world of design that that can be used as practice for a more complex and developed way of creating 3D objects that may be printed. It allows users to group objects together to make a solid figure, put a hole in another figure, or a variety of other options with the accompanied settings provided to make them possible. In the lab session on November 20, 2013, groups were given a file on TinkerCAD to search for and modify. The objects were the PCR tubes, however they were not connected or labeled. Each group did their own modification along the lines of labeling and connecting the tubes.
Practical uses for TinkerCAD range from myriad uses for different professions. Students in labs may use TinkerCAD as an introduction to the 3D printing process for the preparation for their profession. With the knowledge of the program, those students and others with knowledge of the system may use if for their own leisure use to practice their own ideas. A research lab in need of a more reliable camera stand for a fluorimeter, an openPCR machine, or even the box used to block out light for the fluorimeter may use TinkerCAD to construct those objects for their own use or for the use of engineering the object as an improvement for the disbursement into the public.
Feature 1: Cancer SNP-Specific PrimersBackground on the cancer-associated mutation The sequence "rs17879961" is found in the Homo Sapiens species and the nucleotide that is affected is clinically significant due to the fact that it contains a pathogenic allele--that for the cancer substitution. In this case, the polymorphism occurs on the 22nd chromosome out of the 23 chromosomes that humans possess. Going further into the actual gene that is affected in the "rs17879961" sequence, it is called CHEK2 which stands for "checkpoint kinase 2." Based on the NCBI database's information on checkpoint kinase 2, the damage that occurs to the DNA stops the progression of the cell cycle prior to mitosis in a cell that stabilizes the tumor supressor "protein p53" which is linked to the predisposition for the cancers: sarcomas, breast cancer and brain tumors. Primer design
How the primers work: Primers are DNA sequences, or template DNA, which provides a specific nucleotide sequence in both the forward and the reverse direction. The forward and reverse strands are complimentary to one another, meaning that their base pairs match up with their compliments--A to T and C to G. In order to amplify the DNA sequence, these complimentary nucleotides join together and add on new nucleotides to extend the strand. The primers being complimentary for the cancerous allele strand is what allows the pathogenic DNA to amplify. If the template DNA for rs17879961 contained the non-cancer allele, it would not amplify exponentially because the primers would not be complimentary to the target DNA sequence.
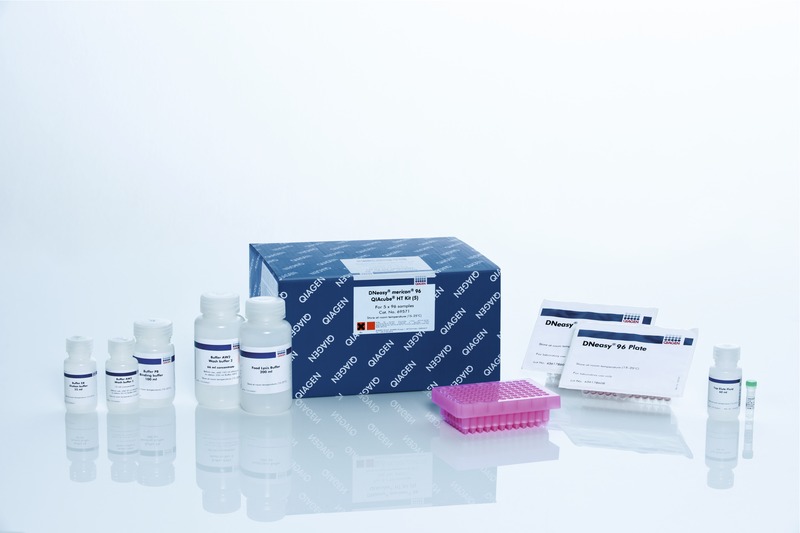
Feature 2: Consumables KitIn our consumable kit we decided to improve and consolidate our kit from the one we were given in lab. We decided that our kits could be ordered in various quantities depending on how many reactions are needed to happen. First, accurately labeled labels for PCR and primer mixtures need to be established. These mixtures will come in pre labeled, bottles that have screw off tops so that pipetting can occur easily. The kit will also include micropipette tips, standardized tubes and racks, and a micropipette. All of these items will come sealed in there own individual labeled plastic bags to ensure these items are sterile as well as organized and not mixed up. There will be a user-friendly manual that will also contain a brief tutorial on how to use a micropipette as well as the overall objective of the test being preformed by this consumable test kit. Our kit will be similar to the one promoted by QlAquick PCR. We found that the majority of this kit was similar to what our ideal kit ought to contain. A weakness in the consumables that we received was that our different mixtures needed to have clear and concise labels on them so that no mixtures would be confused. In this kit the PCR mix and primers are easily distinguished. Something that would be included in our kit would also be a large consumable waste package, such as a biohazard waste bag to help with the large clean up. A biohazard bag large enough to contain the waste products for this procedure will easily be able to be folded up and placed in the bottom of box of the kit. Ideally our kit will come in different sizes, however our most basic kit will come prepared for 50 PCR reactions so that the consumer can decide whether our style of kit is what they are looking for. Below is an image of the QlAquick PCR kit that our kit is based off of. This most accurately depicts what our improved kit would look like, however this does not include our additions and minor changes to the kit listed above:
Feature 3: PCR Machine HardwareThe PCR kit will function similarly to how it functioned in our previous experiment, in that it will be responsible for producing and amplifying new DNA strands. It contains a power supply, a cooling fan, an LED display for easy reading, a circuit board for information storage, and a heat lid to prevent condensation. The PCR machine denatures the DNA, then anneals and elongates the strands to provide more copies, which can then be used for analysis. The machine runs multiple cycles of the DNA strands found from the Consumables Kit for a exponentially increasing DNA sample, although this process does have some aspects that can be improved. Our PCR would be augmented so that the number of sample tubes that can be processed is higher, and therefore more experiments can be run during the same time interval. Also, the cooling and heating systems could be improved by making them more potent. Thus, the time required for the cycle would further increase, because the period required for specific heat up and cool down would be decreased. When we ran the PCR machines in lab, the time to prepare the specific temperature was a major factor that consumed our time. These solutions would both decrease the time it would take to run the cycles for the experiment. Feature 4: Fluorimeter HardwareThe fluorimeter works by first putting a drop of the sample being tested and a drop of the florescent dye on a glass slide that sits on the fourometer. The dye being used is usually SYBR green. When the flourometer is turned on, the dye will be exposed to the flouomiter's light. A camera under a light box is used to tell if the dye absorbed the light. If the sample appears to be glowing in the pictures taken, then the dye absorbed the light. When the dye absorbs the light, the sample contains the strand of DNA. The flourometer will come preassembled and packaged in foam. In order to make the flourometer more effective, our company made a few changes. First, the camera stand will be connected to the flouromiter. This allows the camera to be placed the correct distance from the sample every time a picture is taken. Next, the light box will be redesigned in order to shut out more light. Instead of folding the box will keep its shape at all times and have a door that latches. The new door design is more effective and easier to use. Overall these changes are cost effective and make the testing process easier.
Bonus Opportunity: Instruction ManualInstruction Manual: Materials:' Primers DNA mixture Micropipette Standardized tubes and racks Biohazard waste bag SYBR green solution Slides Labels for patient I.D.’s Instructions: 1. Make sure all of the materials listed above are fully prepared and in the diagnostic kit. 2. Download Open PCR software to your computer (a link can be found on the back of this manual). 3. Follow standard protocol for Open PCR test (can be accessed online through an additional link provided). 4. Make sure that all waste from this procedure is properly disposed of in the biohazard waste bag, and properly thrown away. There is an additional link that can be found to contact a biohazard waste removal representative near you. |
||||||