BME100 f2013:W900 Group4 L4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||||||||||
|
OUR TEAM
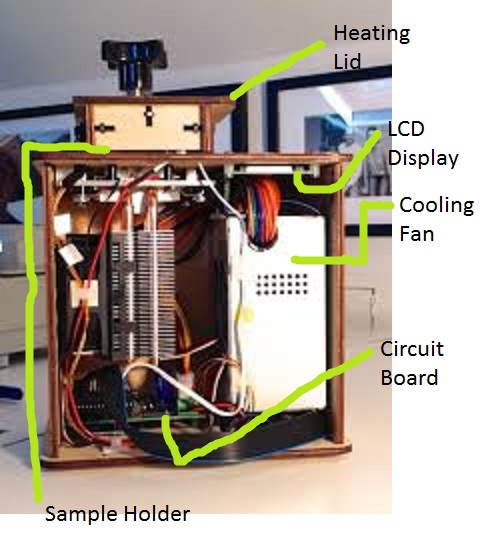
LAB 1 WRITE-UPInitial Machine TestingThe Original Design
When we unplugged LED display from the Brain Board, the machine began flashing and fluctuating several different patterns, numbers, and letters on the LED display. When we unplugged the white wire that connects Brain Board to the LED display, the machine loses the ability to take the correct temperatures and went from the starting temperature in Celsius down about sixty degrees. Test Run On October 23rd, 2013, at approximately 10:17 AM, the OpenPCR machine was started and ended at about 11:35 due to the time limits. The time remaining in the machine's cycle was about 35 minutes. When working with this machine we did experience some flaws in the construct such as loose panels and hard to reach places. Also the small screws did not have the nut to go along with most of them. This left the walls of the machine unstable and not secure when examining the machine. Other than those minor flaws, our experience with the OpenPCR machine was quite satisfying.
ProtocolsThermal Cycler Program DNA Sample Set-up
DNA Sample Set-up Procedure
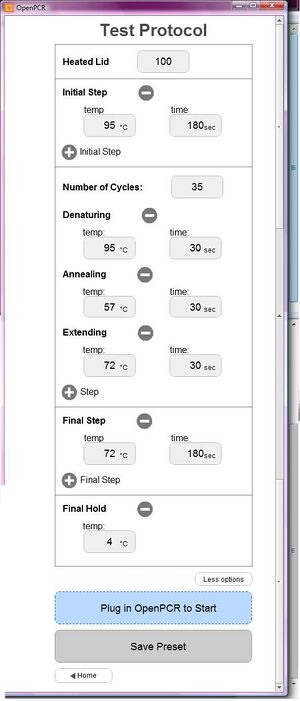
Below is a screen shot of the correct OpenPCR program set to the correct parameters and ready to run:
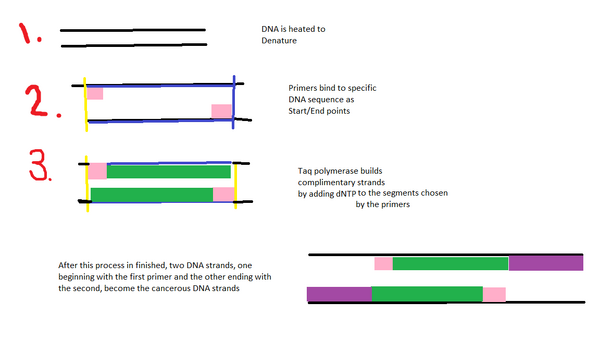
Research and DevelopmentPCR - The Underlying Technology There are five basic components of a Polymerase Chain Reaction, each with specific functions that help complete the process. The template DNA is the original strand of DNA that will then be replicated to create new strands. Primers are sequences of DNA that attach to the start and end of the specific strand trying to be replicated. Taq Polymerase reads the DNA sequence and attaches matching nucleotides to the original strand to build a copy. During the PCR reaction, Magnesium Chloride causes the reaction to go forward and determines the speed of the reaction. It is a cofactor of DNA Polymerase and must be optimized for each primer. Deoxyribonucleotides, on the other hand, are single nitrgen-base units that act as the building blocks of DNA. After the PCR reaction mix, the sample and the DNA/Primer mix have been combined, and the sample has been placed into the 16 tube PCR block, the thermal cycler will go through an initial step where it heats up to ~95 Degrees Celsius and is then held at this temperature for 3 minutes to begin the process of activating the DNA polymerase. After this step is complete, the denature step begins --this also occurs at ~95 Degrees Celsius-- and during this step, the sample DNA will separate into single strands. From here, the temperature will be decreased to ~57 Degrees Celsius for the annealing phase. During this phase, the two primers will bind to their complimentary base pair on the target DNA. The temperature will be raised once more but to ~72 degrees Celsius for the extension phase. During the extension phase, the taq polymerase will bind to the primer and will begin adding the four nucleotides described below to the second strand of DNA. According to the given hand out, the denaturation, annealing and extension phases each occur for 30 seconds. the ~72 degrees will be held for 3 more minutes after the extension step in order to ensure that all of the single strands have been fully extended. Finally the temperature will be lowered to ~4 degrees Celsius and the data collected from the thermal cycler will be stored for reference. DNA is built using four types of nucleotides, A, C, T, and G. These bases stick to corresponding bases on the second strand of the DNA via hydrogen bonding, and allow for the attachment of primers to stick to the template strand. Adenine sticks to Thymine, Thymine to Adenine, Cytosine to Guanine, and conversely Guanine to Cytosine.
| ||||||||||||||