BME100 f2013:W900 Group16 L4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||||||||||
|
OUR TEAM
LAB 1 WRITE-UPInitial Machine TestingThe machine in the picture is known as an OpenPCR machine. Although this machine seems to be nothing more than a wooden box, it is actually a mechanism that is used to amplify target sequences of DNA by continuously heating and cooling the DNA to extreme temperatures. The process that occurs in this machine is known as PCR which is short for Polymerase Chain Reaction. In this process, the extreme temperatures that the DNA is exposed to causes the DNA to unwind, separate into two different strands and are used to make multiple copies of the target sequence of DNA.
When the wire which connected the LCD screen (part 3) to the Circuit Board/Mother Board (part 6) was disconnected, the machine did turn on, however no information was displayed on the LCD screen because none of the information from the Circuit Board was sent to the LCD screen to be displayed. When the white wire which connected the Circuit Board/Mother board (part 6) to the Sample Compartment (part 2) was disconnected, although the samples were being heated, the information that was being displayed on the LCD screen was inaccurate because the disconnection of the white wire stopped the correct heating temperature to be sent to the LCD screen.
This experiment began on 10-23-13 at 9:56 am and ran until 11:39 am.
The test was completed fairly quickly and the machine seemed to run smoothly.
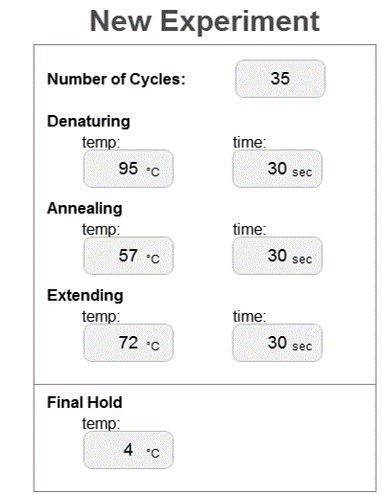
ProtocolsThermal Cycler Program DNA Sample Set-up
The PCR reaction mix consists of 8 tubes with 50 µL each of Taq DNA polymerase, MgCl2, and dNTP’s.
DNA/ primer mix The DNA/ primer mix consists of 8 tubes with 50 µL each of a different template DNA. All the tubes have the same forward primer and reverse primer.
Research and DevelopmentPCR - The Underlying Technology Function of PCR Components The Polymerase Chain Reaction requires many components to amplify the desired DNA sequence. A DNA template is necessary to begin the entire PCR process because it has a section of interest. The billions of DNA copies that result at the end of PCR are all derived from this original DNA template. Primers are needed to replicate DNA and are used in the normal DNA replication process. The primers used in PCR recognize a desired segment of DNA and bind to the DNA strand after denaturation has finished. By having a specified base sequence segment to bind to, the primers begin the process of targeting and isolating the DNA portion of interest. The job of a primer in DNA replication and PCR is to tell DNA polymerase where to start adding the complementary base pairs to the DNA strand. In PCR, primers are unique because they can be synthetically made to target a specific DNA segment. However, regular human DNA polymerase cannot be used in PCR because it cannot withstand high heats; instead, Taq polymerase, a polymerase taken from a thermophilic bacteria, is used to add the complementary base pairs to the DNA strands. Taq polymerase is used because it can withstand the heat cycles required in PCR. Magnesium chloride is used to help stabilize Taq polymerase which depends on a magnesium concentration to function properly. dNTPs are the single A, T, C, G bases and are the building blocks for the new DNA strands. For creating the new strands of DNA, the base pairing is very important. The base pairings go as follows: Adenine (A) pairs with Thymine (T), while Cytosine (C) pairs with Guanine (G). The pairings are also true vice-versa. So in other words, Thymine (T) pairs with Adenine (A) while Guanine (G) pairs with Cytosine (C). Steps of PCR The first step of PCR is a 3 minute cycle that heats to 95 degrees Celsius. During this step, the DNA unravels from its double helix formation and lines the two strands of DNA parallel to each other. This step of the PCR aligns DNA properly for the other steps to occur. Denaturation is the next step and lasts for 30 seconds and is still held at the 95 degrees Celsius. In this step, the hydrogen bonds holding the complementary base pairs break and the two DNA strands separate from one another. Annealing lasts for 30 seconds and the temperature drops to 57 degrees Celsius. This is the point in PCR where the primers bind to the DNA strands. After annealing, extension occurs. Extension is a 30 second process held at 72 degrees Celsius in which the Taq polymerase adds the complementary base pairs to the DNA strands starting at the primer. The final step is also at 72 degrees Celsius but lasts for 3 minutes. This step allows Taq polymerase to find all of the primers in the DNA sequences. The last step is the hold. The final hold is at 4 degrees Celsius and is meant for investigating how much DNA is in the sample after PCR has cycled.
The following diagram displays how primers bind to the cancer DNA template, and how Taq polymerases amplifies the DNA
| ||||||||||||||