BME100 f2013:W900 Group13 L6
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||
OUR COMPANY
PCR: Who Do you THINK you are?
LAB 6 WRITE-UPComputer-Aided DesignTinkerCAD Implications of Using TinkerCAD for Design
Feature 1: Cancer SNP-Specific PrimersBackground on the cancer-associated mutation rs17879961 is the title of the gene present in DNA sequences in humans. A nucleotide is a compound consisting of a nucleotide linked to a phosphate group and is the basic structural unit of nucleic acids. A polymorphism is where there is one or more variants of a particular DNA sequence. The species in this variation is rs17879961 is homo sapiens (humans). The clinical significance of this SNP is that is so pathogenic. This SNP (rs17879961) occurs on chromosome 22 out of the typical 23 pairs of chromosomes that humans have. CHEK 2 (Checkpoint Kinase 2) when activated inhibits CDC25C phosphotase, preventing entry into mitosis, and may stabilize the tumor suppressor protein p53, leading to cell cycle arrest in G1. This minuscule change altering CHEK2 has catastrophic affects, it brings a person one step closer to cancer.
How The Primers Work: In a PCR reaction, primers are one of the most important components in amplifying DNA. Both of the primers are designed to bind only to a specific sequence of the DNA that contains the SNP. When the primers bind to the DNA, it will take nucleotides that are complementary to the sequence of the DNA from the reaction mix and bind them together. This happens in both the forward and reverse directions in order to incorporate both strands of the DNA.If the sequence does not correlate 100% to template of the forward or reverse primer, then the complementary DNA strand will not be formed. This way, the primers will exclude DNA with non-cancer alleles and amplify DNA with the cancer-associated SNP rs17879961. Feature 2: Consumables KitIn our kit, the consumables will be clearly labeled to differentiate between the primers, PCR Mix, and the sample DNA. The tubes in which these materials are packaged will be color-coded and labeled to avoid any confusion that could lead to mistakes in the lab. We will maintain the current micropipettor design and will not be changing the PCR tubes themselves outside of the aforementioned labeling. This ids due to our belief that the shape of the current PCR tube allows for the substance to be pipetted easily, any changes to that aspect wouldn't improve the device much. We will also be making the packaging tubes have a hydrophobic surface on the inside so that the materials do not stick to the inside of the tube. These changes will help prevent human error and will also help prevent excess waste from being left over. One of the weakness's addressed in the Virtual Comment Board Powerpoint was the self-assembly associated with OpenPCR. Our consumables kit does address this problem, since self-assembly leads to much human error, our labels would minimize human error since it would make identifying PCR mix versus primers easier which would cause the experiment to be more accurate.
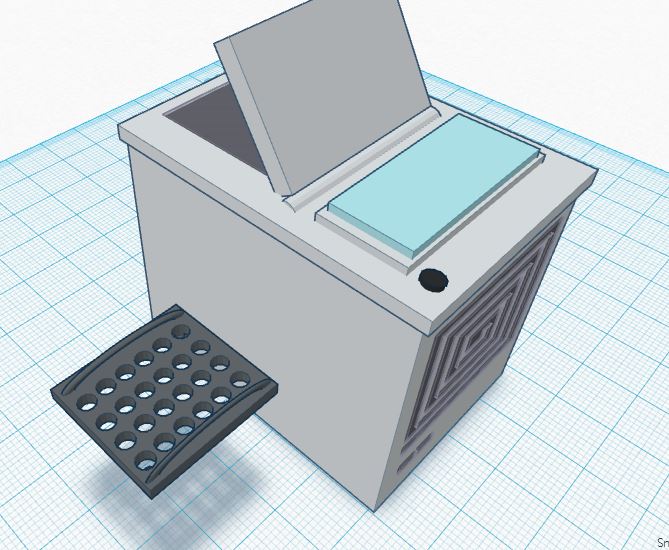
Feature 3: PCR Machine HardwareThis kit will include a redesigned PCR machine that features a latch to seal the PCR lid tight, a durable and heat-proof plastic design (as opposed to the wood-built material), and a removable plate for easy transport of PCR tubes.
The redesigned PCR machine incorporates a latch in order to keep the heat inside the PCR machine. This will keep the temperature constant to facilitate the PCR reaction in the three processes of denaturing, annealing, and extending. This prevents susceptibility for error, allowing consistent performance. A durable, heat-proof plastic design will minimize cost to allow maximum profit. The addition of the removable plate will allow for easy transport as well as facilitate loading/unloading of the samples. Also, the redesigned PCR machine includes additional holes in the sample tube racks to allow for more DNA samples to be amplified and replicated.
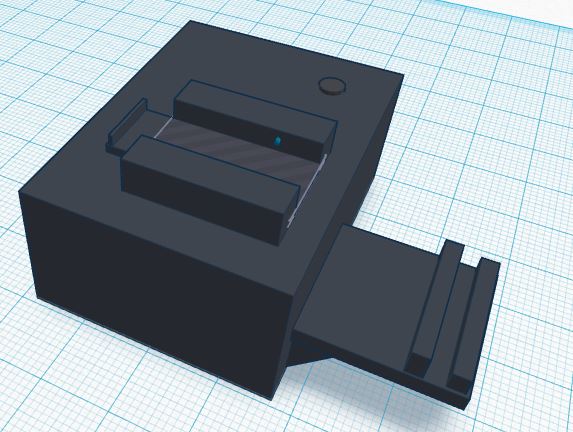
Feature 4: Fluorimeter HardwareA customized fluorimeter will be included with this kit that increases the efficiency of the fluorimeter. This redesigned device will include a stand that is attached to the box, keeping the fluorimeter and the camera stand at a set distance. Included with this kit is a user-friendly phone application that instantly sends photos taken on the fluorimeter to a computer with Image J for processing. The redesigned fluorimeter includes a stand attached to the box to minimize variables of operation. This will keep the flourimeter and the camera stand at a set distance that minimizes error from any displacement from the stand or box. The phone application included with this kit facilitates the process of calculating concentrations by cutting the step of the user sending all of the photos to the computer. This application will utilize the camera of the user's phone and--using wifi--will send photos taken on the fluorimeter to a designated computer with Image J for processing. In addition, an adjustable platform for the glass slides will be incorporated into the fluorimeter design to facilitate the movement of the slide from one drop to the next as well as both the disposal and the replacement of the sides. Our group, as well as other groups, struggled with the tediousness of placing and replacing the glass slides. The adjustable slide platform will help solve that, making calculations easier and faster to retrieve.
Bonus Opportunity: What Bayesian Stats Imply About The BME100 Diagnostic ApproachFor calculation 3, the probability that a patient will develop cancer, given a cancer DNA sequence [P(A|B)] was determined. The calculated probability was small, just under 40%. However, there is a flaw with what this probability infers. In the lab, PCR was used to determine whether or not patients had cancer. PCR is not the most efficient way to accurately determine if a patient has a SNP; sending the DNA to a sequencing lab is a more accurate method of determining if a patient has the given SNP. This probability makes the assumption that a patient has the SNP based off of three PCR reactions susceptible to false conclusions. Also in the lab, the cancer was diagnosed in the patients through a biopsy. Where this probability lacks again is that it is uncertain that the SNP causes cancer; another gene besides CHEK2 could cause this same cancer and would be undetected by the PCR reaction. Another factor is that a patient in this experiment could have been developing cancer, but it went unnoticed during the biopsy. Through these examples, the limitations of this probability are revealed which demonstrate its reliability. For calculation 4, the probability that a patient will not develop cancer, given a non-cancer DNA sequence [P(A|B)] was determined. The calculated probability was around 130%. Just like the limitations of calculation 3, this probability makes the assumption that a patient does not have the SNP based off of three PCR reactions susceptible to false conclusions. Where this probability lacks is that another gene could code for the same cancer that CHEK2 codes for, and the patient would develop cancer while not having the CHEK2 gene. These examples express the limitations of this probability which demonstrate its reliability. |
||||||