BME100 f2013:W900 Group12 L6
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||
OUR COMPANY
by BioPro Incorporated
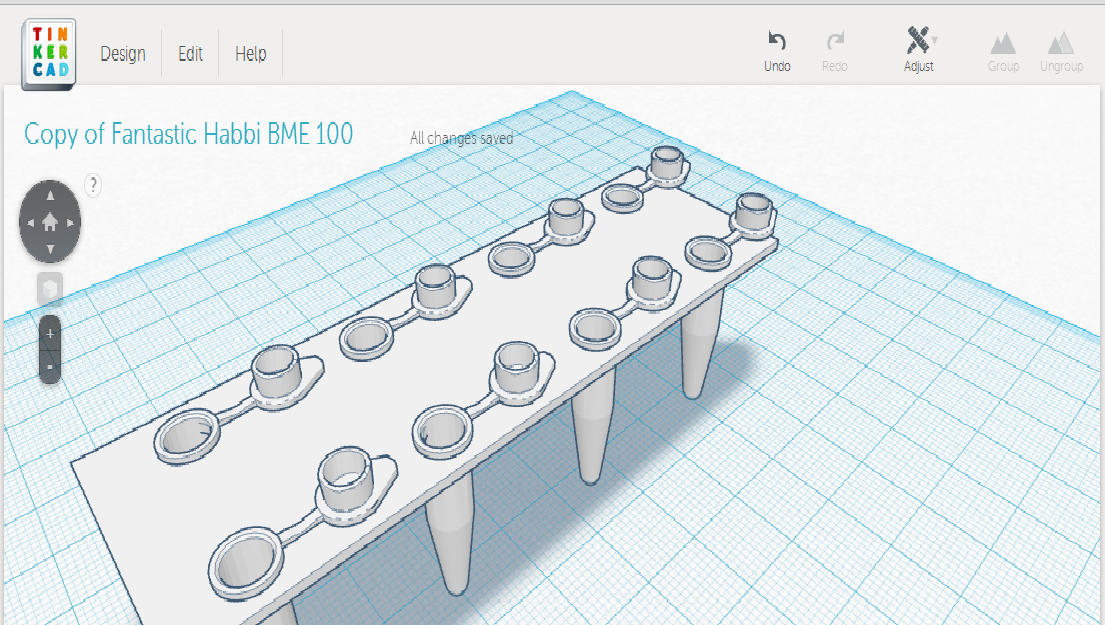
LAB 6 WRITE-UPComputer-Aided DesignTinkerCAD TinkerCAD is a web-based 3-D application. It allows for easy and clear creation of items in 3-D form in order for prototyping and production by companies. This tool is beginning to be used in the biomedical field for advanced 3-D printing. In the lab, a new and improved design of PCR tubes is created for a new PCR application kit produced by our company. Many of weaknesses that were experienced from the original PCR tubes was corrected and the strengths were kept. TinkerCAD allowed our company to design the item with precision and improve certain parts without wasting time and resources producing the actual 3-D product until it can be improved no more.
TinkerCAD can be used to redesign many items, such as the camera holder for the fluorimeter. The design of the fluorimeter has many weakness. The lack of a camera holder affects the quality of the pictures taken by the device. A camera holder can be designed using the TinkerCAD application. This device would be large enough to accommodate a smartphone with stability. It should be able to balance the phone so that the camera is visible and the subject is centered on the screen. The new design can be created in a short time using the TinkerCAD because of it's user friendly options. The design could then be sent to a 3-D printer and printed out on demand. There are many possibilities for using TinkerCAD to improve the designs of the machines used in the experiment. Due to the simple and fast process, TinkerCAD is a effective and efficient application to be used in future device design.
Feature 1: Cancer SNP-Specific PrimersBackground on the cancer-associated mutation A nucleotide is composed of a phosphate group linked by a phosphoester bond (deoxyribose), which also known as an amino acid. A polymorphism is a common variation in the sequence of DNA among individuals. SNP stands for single nucleotide polymorphism. We want to consider SNP rs17879961, known as a cancer-associated mutation that can be found in homo sapiens. The clinical significance of SNP rs17879961 is the pathogenic allele. Moreover, this SNP is found on the 22nd chromosome in the 23 pairs of human chromosomes. The affected gene is called CHEK2, which stands for the checkpoint kinase 2. CHEK2 inhibits CDC25C phosphate, preventing mitosis and leading to cell cycle arrest in G1. This links to Li-fraumeni syndrome, as well as can cause sarcomas, breast cancer, brain cancer, and tumors. Primer design
How the primers work: ' In order to amplify the DNA, each PCR reaction needs to have two primers based on the forward primer that we design. This primer contributes to the functioning of the PCR process. The numerical position 200 bases to the left of the cancer single nucleotide polymorphism (SNP) is 29120887, which means the forward primer releases from the top strand of the DNA. It is known that the non-cancer allele includes the sequence ATT, while the cancer allele contains the sequence ACT. In the PCR process, the primers will work with the forward primer and the reverse primer. When the cancer specific primer is formed, the reverse primer will bind to complete a cancer SNP that contains the sample of the patients. The forward primer will then completely bind to the sample of the patient. This is the whole process of amplifying the DNA. Unless the primer can bind 100% to the template(sample of patient), the PCR reaction cannot happen, and the template will consist of the non-cancer allele. As a result, the strand will not amplify.
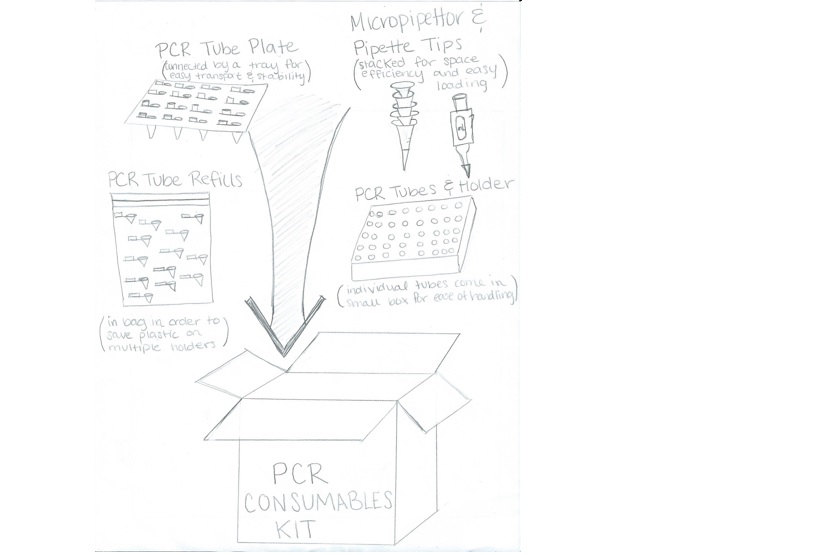
Feature 2: Consumables KitOur consumable kit we would include a PCR tube plate, micropipettor, pipette tips, PCR tubes and holder, and PCR tube refills. Our pipette tips would be packaged stacked one on top of the other to conserve space and allow for easy loading. The tube plate would resemble a tray that gives easy transport and stability. The individual PCR tubes would come in a small plastic box for ease of handling. We would also include a bag of PCR tube refills, rather than box refills, in order to save plastic and be more environmentally efficient.
Another problem with the consumables kit is the amount of waste you produce from the amount of plastic used in having to discard so many pipette tips and PCR tubes. Because these materials are considered "Hazardous Waste" you cannot recycle the actual tips or reuse the materials. For this reason, the focus should be on being economical in the use of the pipette tip box. The majority of the plastic used comes from the box, rather than the tips. By buying bags of individual pipette tips, and refilling the box, you avoid having to throw away the plastic-consuming box after you use up all its tips.
Feature 3: PCR Machine HardwareThe PCR machine will function in much the same way as it was used in lab. The steps and mechanics of copying the DNA will remain the same--denaturing, annealing, and extending in order to replicate and form the target sequence. The simplicity and compatibility will be maintained. The one difference will be in the number of PCR tubes the machine can fit and process in one run. The machines used in lab fit 16 tubes at once. Our design will double this number and have the capability to process up to 32 solutions in one run.
One of the primary weaknesses of the PCR machine is its terribly slow processing speed and low through-put. Only 16 solutions can be copied in one run, and one run takes around two hours. To account for this problem, we will double the through-put to 32, so that more PCR product is copied in one run of the machine.
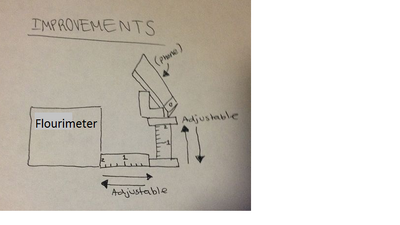
Feature 4: Fluorimeter HardwareWe will include the fluorimeter just as we used it in lab, following the same procedure of using the dark box and the smartphone. One significant change will be in the smartphone holder. We will have an adjustable stand that allows the user to easily secure the phone in place. and adjust the height and distance. In this way, the user can raise or lower the phone camera to be directly even with the drop on the slide. One of the problems with the fluorimeter was the stand used to hold up the smartphone when taking pictures. The stand was not a good height for the phone to take pictures, so the lab group had to stack books under the device in order to get a clear picture. The stand was also not stable and did not stay in a fixed spot, which allowed for inconsistent data to be taken. In order to improve the device, we will create a standardized system for the phone holder and a method of adjusting the height and distance, which would enable users to obtain better results.
Bonus Opportunity: What Bayesian Stats Imply About The BME100 Diagnostic ApproachThe Bayesian Statistic method helps identify and determine the reliability of the PCR to detect and predict the sensitivity or specificity of the cancer SNP and cancer. Calculation Three describes the sensitivity of the system to predict cancer, meaning it relays the probability that a person with cancer will test positive using the CHEK2 PCR system. As this percentage was quite small and significantly less than one, the PCR often gave "false positive" results. This means that positive predictions from the PCR are largely unreliable as many of the positive results correlated to non-cancerous patients. Calculation Four describes the specificity of the the system to predict cancer, which means it gives the probability that a person without cancer will test negatively according to the PCR. Contrary to calculation three, this percentage was quite large and only slightly less than one. This means that most of the time the PCR gave reliable non-cancerous predictions, as a negative result highly correlated to non-cancerous patients. In conclusion, the CHEK2 PCR system is fairly unreliable and has low sensitivity in its ability to predict cancer, while it has high specificity and is a good standard for predicting no cancer. |
||||||