BME100 f2013:W900 Group11 L4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | |||||||||||||||
|
OUR TEAM
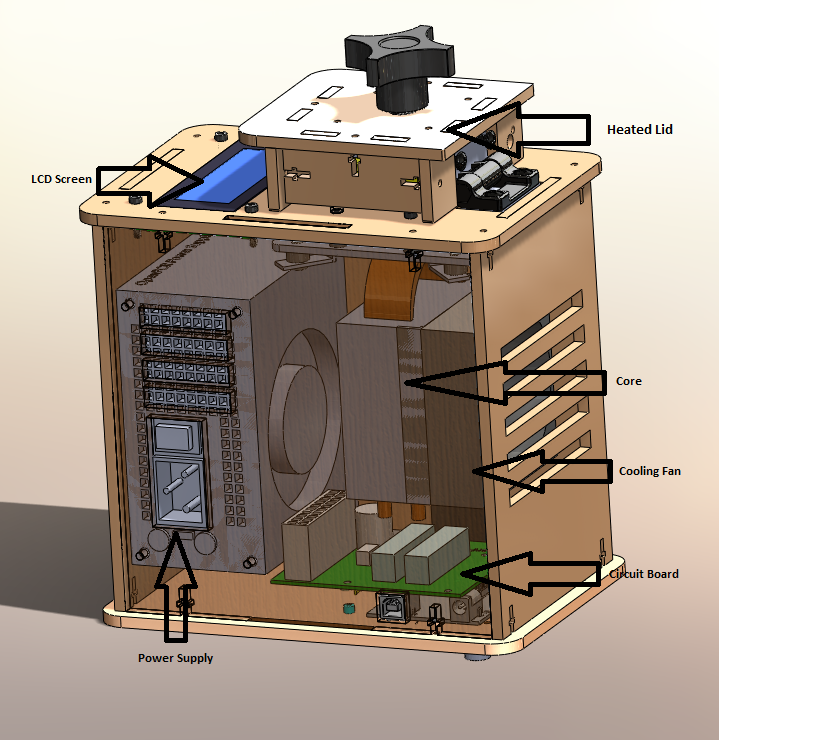
LAB 1 WRITE-UPInitial Machine TestingThe Original Design
When we unplugged the white wire that connects (part 6) to (part 2), the temperature reading that were displayed changed. the white wire controlled the accuracy of the temperature reading and when it was disconnected it did not read the temperatures right. Test Run The PCR machine was tested on October 23, 2013. unfortunately the machine did not operate as expected. when the test started with the test run it was supposed to take 7 minutes to warm up but it stayed at that station for about 30-45 min. this clearly showed that the machine was broken at some parts.
ProtocolsThermal Cycler Program When the OpenPCR machine runs, it begins with the warm-up process. The process that takes 3 min as it warms up to 95 C to denature DNA, and let run for 25 cycles. Repeat, adding PCR reaction mix to each sample, and letting run for 25 cycles. Each cycle is comprised of 30 seconds at 95 C, which separates DNA into separate strands, 30 seconds 57 C for primer attachment, and 72 C to begin action of DNA polymerase. DNA Sample Set-up
DNA/ primer mix
Research and DevelopmentPCR - The Underlying Technology The purpose of Polymerase Chain Reaction is to make a large number of copies of a DNA sequence using primers, which are short pieces of DNA created in a lab to match the template DNA. To complete this process, you need template DNA, which is the sample DNA that contains the target sequence. You will need a Taq polymerase (a DNA polymerase) that is stable in many temperatures. The polymerase is used to synthesize a new DNA strand from the template. It attaches itself near the end of the primer and begins adding nucleotides, which are the building blocks that DNA is made of (A’s, C’s, G’s, and T’s). You will need magnesium chloride, acts as a cofactor or a helper to the activity of the enzymes. The first step of PCR is denaturation. In this step, the solution is heated to 95 degrees for 30 seconds. During this time the hydrogen bonds in the DNA strand are broken, which allows it to be separated into two strands so that copying may occur. The next step is annealing, in which the solution is cooled down to 57 degrees for 30 seconds. In this step the primers form ionic bonds with the DNA so that the polymerase can start copying. This is where the magnesium chloride comes into play. Because the DNA strands are negatively charged, they need a salt to decrease the repulsion between them. Magnesium does this and allows the two strands to come together and anneal. After this is extenstion. The solution is heated to 72 degrees for 30 seconds because 72 degrees is the ideal temperature for the polymerase. During the 30 seconds, the Taq polymerase adds the deoxyribonucleotides (A’s pair with T’s and C’s pair with G’s), creating two double-stranded DNA strands from the original template DNA. This whole process repeats itself for 30-40 cycles until there are a lot of copies of the original DNA.
| |||||||||||||||