BME100 f2013:W900 Group10 L4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | |||||
|
OUR TEAM
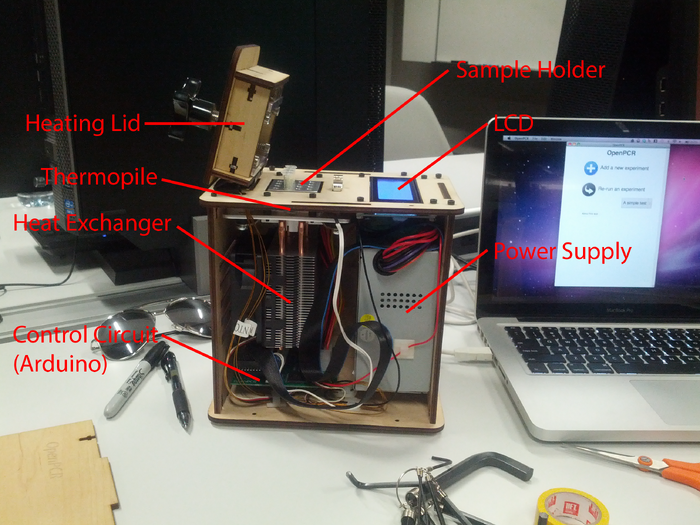
LAB 1 WRITE-UPInitial Machine TestingThe Original Design OpenPCR is a small and relatively low cost PCR (polymerase chain reaction) machine. In the past most PCR machines were prohibitively expensive, and thus not available to most schools and enthusiasts. The designers of OpenPCR changed this by creating a very small machine containing low cost and widely available electronics (Arduinos, theremometer, LCD, ect.) and other components (wooden frame, etc.). A polymerase chain reaction, or PCR, is a chemical reaction that takes advantage of biological molecules (mostly proteins) to make billions of copies of a certain segment of a template DNA strand. A PCR reaction relies on accurately timed temperature variation to work properly. This is what a PCR machine is for. Tubes containing the PCR reactants are placed into the the heat exchange block in the PCR machine. The machine then proceeds to cycle the temperature of the PCR mixture for a user defined period of time and over a user defined temperature range. The result is massive amplification of the desired DNA segment in the PCR reaction tubes.
When we unplugged the LCD screen (part 3) from Arduino and PCR shield (part 6), the LCD screen turned off because it was no longer receiving power or signal from the main circuit board. When we unplugged the white wire that connects the Arduino and PCR shield (part 6) to the thermometer (part 2), the machine began displaying incorrect temperature information.
On October 23, 2013, we ran the PCR machine through a test cycle. The machine ran smoothly throughout the entire process (10:06am - 11:24am) and ran very closely the the rate estimated by the time to completion readout. At all points during the test the information displayed on the PCR machine's LCD screen matched that displayed on the computer.
ProtocolsThermal Cycler Program - Set heating lid temperature to 100 degrees celsius. - Step 1: Heat to 95 degrees celsius for 3 minutes to fully denature all DNA. - Step 2: Hold at 95 degrees celsius for 30 seconds to denature DNA. (First step of amplification cycle). - Step 3: Hold at 57 degrees celsius for 30 seconds to allow annealing (primers attach). - Step 4: Hold at 72 degrees celsius for 30 seconds for extension (copying of the bracketed area by Taq polymerase). (Last step of amplification cycle). - Step 5: Repeat steps 2 - 4 thirty five times. - Step 6: Hold at 72 degrees celsius for 3 minutes to ensure the Taq polymerase has finished copying. - Step 7: Bring the temperature down to 4 degrees celsius to stabilize PCR mixture.
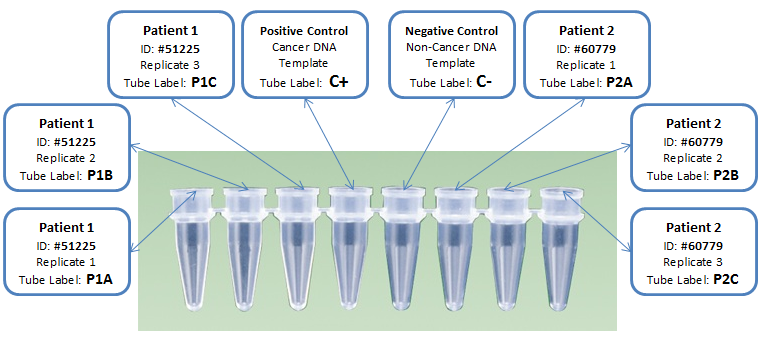
DNA Sample Set-up Procedure Step 1: Plug in and turn on OpenPCR machine. Step 2: Transfer the 8 DNA samples into their respective PCR tube, making sure to use a new pipette each time. Step 3: Transfer primer mixture into each PCR tube. Step 4: Transfer PCR reaction mixture into into each of the PCR tubes. Step 5: Open the OpenPCR and place all 8 PCR tubes into the OpenPCR machine. Step 6: Program the OpenPCR machine to follow the steps outlined in "Thermal Cycler Program" above. Step 7: Close the OpenPCR and run the program. Step 8: After the machine has finished store the PCR tubes for later use.
The PCR reaction mixture contains 50µl of each of the following: - Taq DNA polymerase - responsible for the assembly of the new DNA stands - MgCl2 - a co-factor for the Taq DNA polymerase. Is a catalyst and is therefore not consumed by the reaction. - dNTP's - the building blocks of DNA (Adenine, Thymine, Guanine and Cytosine). Required to construct the new DNA strands.
The DNA/primer mixture contains 50µl of each of the following: - DNA template - the DNA molecule containing the sequence we are trying to amplify. - Forward primer - the short single strands of DNA that is complementary to the starting sequence of the segment we are trying to copy. - Reverse primer - the short single strands of DNA that is complementary to the ending sequence of the segment we are trying to copy.
Research and DevelopmentPCR - The Underlying Technology DNA: As we would expect, everything about PCR revolves around the properties of DNA. DNA, the information storage system of life, consists of two polymer strands, each composed of four different types of nucleotides (Adenine, Thymine, Guanine and Cytosine). These two polymers interact to form a double helix structure. The body has methods of copying DNA and it is from these natural methods that PCR originated. In the body, a protein called DNA helicase "unzips" DNA to allow the molecule that actually copies the DNA (DNA polymerase) to function. For PCR to work the DNA strands must also be "unzipped", but instead of using DNA helicase to do this, PCR heats the DNA up to a temperature where it denatures (the two strands separate). PCR also takes advantage of the fact the complementary stands of DNA tend to stick together. PCR uses short strands of DNA to bracket the area of DNA that is to be copied. Proteins (Taq Polymerase): Because of the high temperature required to denature DNA, PCR cannot use human DNA polymerase which would also denature (and cease to function) at these temperatures. Therefore another type of DNA polymerase found in organisms that live in high temperature environments is used: Taq polymerase. This protein has the same function as human DNA polymerase but does not denature at high temperatures. It also has the benefit of activating at a temperature where the DNA primers are still attached to the template DNA but the template DNA strands are still separated. PCR Reaction Mixture: The solution in which the PCR reaction takes place contains molecules that facilitate the PCR reaction. For the Taq polymerase to create new strands of DNA it needs Deoxyribonucleotides (dNTP's): Adenine, Thymine, Guanine and Cytosine. Therefore these molecules are included in the reaction mixture. Also, Taq polymerase requires the presence of magnesium chloride (MgCl2) as a catalyst. Therefore this molecule is also added to the reaction mixture. Process: A thermal cycle is used to choreograph the PCR reaction. First, the PCR mixture is heated to 95 degrees celsius to denature the template DNA strands. Then the mixture is brought down to 57 degrees celsius, the temperature at which the primers will attach to the template DNA strands (known as annealing), but the DNA strands will not reattach to each other (because of the interference of the primers). Then the mixture is heated to 72 degrees celsius, the temperature at which Taq polymerase functions, causing copying to occur. Once the Taq polymerase has finished, this process is repeated. This cycle can be repeated for as long as there are dNTP's present. Usually this process is only run long enough to generate enough copies of the target strand for whatever application the copies are being used for (forensics, study, etc.).
| |||||